
Rab Proteins and the Organization of Organelle Membrane Domains
... For example, cargo from the extracellular environment is internalized into early endosomes where it is sorted for recycling to the plasma membrane or degradation in lysosomes. Clearly, such steps need to be coordinated in time and space. The work on Rab GTPases has revealed molecular features and fu ...
... For example, cargo from the extracellular environment is internalized into early endosomes where it is sorted for recycling to the plasma membrane or degradation in lysosomes. Clearly, such steps need to be coordinated in time and space. The work on Rab GTPases has revealed molecular features and fu ...
Analysis of the bipartite networks of domain compositions and
... Isolated. A domain subunit is isolated if it does not cocatalyze with other domain subunits in any effect reaction. In other words, an isolated domain subunit is not connected to any other domain subunit in Gh . In the lower right graph of Figure 3, the fatty acid hydroxylase superfamily (PF04116) a ...
... Isolated. A domain subunit is isolated if it does not cocatalyze with other domain subunits in any effect reaction. In other words, an isolated domain subunit is not connected to any other domain subunit in Gh . In the lower right graph of Figure 3, the fatty acid hydroxylase superfamily (PF04116) a ...
Functional and structural studies of a C
... ribosomes. Ribosomes are nanomachines which translate the genetic DNA-code into proteins. Prokaryotic ribosomes are constituted of two unequal subunits, the small ribosomal subunit 30S and the large 50S subunit, which assemble during the initiation step of protein biosynthesis to form the active 70S ...
... ribosomes. Ribosomes are nanomachines which translate the genetic DNA-code into proteins. Prokaryotic ribosomes are constituted of two unequal subunits, the small ribosomal subunit 30S and the large 50S subunit, which assemble during the initiation step of protein biosynthesis to form the active 70S ...
N o v e l s ite s o... Johan Ohlson
... (figure 2). The mammalian ADAR1 is expressed from at least three different promoters, one interferon inducible promoter that is giving rise to a 150 kDa protein (ADAR1p150) (George & Samuel, 1999a) and two other promoters giving rise to shorter 110 kDa proteins (ADAR1p110) (George & Samuel, 1999b; ...
... (figure 2). The mammalian ADAR1 is expressed from at least three different promoters, one interferon inducible promoter that is giving rise to a 150 kDa protein (ADAR1p150) (George & Samuel, 1999a) and two other promoters giving rise to shorter 110 kDa proteins (ADAR1p110) (George & Samuel, 1999b; ...
On the Nucleotide Sequence of Yeast Tyrosine Transfer RNA
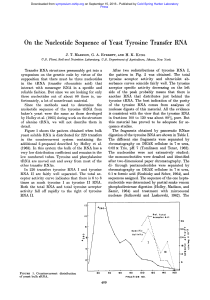
... Transfer RNA structures presumably get into a symposium on the genetic code by virtue of the supposition that there must be three nucleotides in the tRNA (transfer ribonucleic acid) that interact with messenger RNA in a specific and reliable fashion. But since we are looking for only three nucleotid ...
... Transfer RNA structures presumably get into a symposium on the genetic code by virtue of the supposition that there must be three nucleotides in the tRNA (transfer ribonucleic acid) that interact with messenger RNA in a specific and reliable fashion. But since we are looking for only three nucleotid ...
A C-terminus Mitochondrial-localization Region and BH3 Domain of
... Merwin, Liz A., "A C-terminus Mitochondrial-localization Region and BH3 Domain of Puma are Required for Apoptotic Function" (2013). Electronic Thesis and Dissertation Repository. Paper 1283. ...
... Merwin, Liz A., "A C-terminus Mitochondrial-localization Region and BH3 Domain of Puma are Required for Apoptotic Function" (2013). Electronic Thesis and Dissertation Repository. Paper 1283. ...
A Calcium-Regulated Gatekeeper in Phloem Sieve Tubes
... concluded that the most likely function of P-protein is to seal the sieve plate pores of injured sieve elements as a rapid first line of defense against the loss of assimilates. It is possible to obtain electron micrographs from well-preserved and largely uninjured sieve tubes using gentle tissue pr ...
... concluded that the most likely function of P-protein is to seal the sieve plate pores of injured sieve elements as a rapid first line of defense against the loss of assimilates. It is possible to obtain electron micrographs from well-preserved and largely uninjured sieve tubes using gentle tissue pr ...
by 3AB and 3ABC trans Complemented in Formation, and the
... P3 region and the sequence was verified by the dideoxynucleotide method. To prepare P3 plasmids mutated at two cleavage sites, the cDNA containing a single nucleotide change was mutated at the second site by the same procedure. To construct mutated full-length HAV genomes, the P3 region of pT7-18f ( ...
... P3 region and the sequence was verified by the dideoxynucleotide method. To prepare P3 plasmids mutated at two cleavage sites, the cDNA containing a single nucleotide change was mutated at the second site by the same procedure. To construct mutated full-length HAV genomes, the P3 region of pT7-18f ( ...
Electrophoresis Basi..
... neutral pH’s are either basic or acidic depending upon their AA composition. Most proteins placed into basic conditions become negatively charged. Acidic conditions cause most proteins to develop a positive charge. ...
... neutral pH’s are either basic or acidic depending upon their AA composition. Most proteins placed into basic conditions become negatively charged. Acidic conditions cause most proteins to develop a positive charge. ...
Mechanisms and applications of disulfide bond formation
... and mRNAs translated into the amino acid sequence of proteins. The sequence of a protein determines its native structure and function (Anfinsen CB 1973). In nature, proteins are synthesized as linear polypeptides, the so called primary structure, and then fold into their final secondary, tertiary, a ...
... and mRNAs translated into the amino acid sequence of proteins. The sequence of a protein determines its native structure and function (Anfinsen CB 1973). In nature, proteins are synthesized as linear polypeptides, the so called primary structure, and then fold into their final secondary, tertiary, a ...
Biological significance of structural differences between two highly
... Mms2 and Uev1 (Fig. 1B). As seen in Fig. 3, Uev1D30-INNSS (a.a. 116–120) caused a complete loss in binding by LN2A, but only partially reduced binding by LN2 or LN2B. In sharp contrast, the other substitution construct, Uev1D30-ARSIP (a.a. 125–129) completely abolished binding by LN2 and LN2B, but d ...
... Mms2 and Uev1 (Fig. 1B). As seen in Fig. 3, Uev1D30-INNSS (a.a. 116–120) caused a complete loss in binding by LN2A, but only partially reduced binding by LN2 or LN2B. In sharp contrast, the other substitution construct, Uev1D30-ARSIP (a.a. 125–129) completely abolished binding by LN2 and LN2B, but d ...
Rabbit Reticulocyte Lysate Technical Manual
... from added mRNA templates at approximately the same rate as treated reticulocyte lysate for up to 60 minutes. The primary disadvantage of such a system for in vitro translation is that the template mRNA is translated in competition with endogenous globin mRNA, making quantitation of the activity of ...
... from added mRNA templates at approximately the same rate as treated reticulocyte lysate for up to 60 minutes. The primary disadvantage of such a system for in vitro translation is that the template mRNA is translated in competition with endogenous globin mRNA, making quantitation of the activity of ...
INTEINS: Structure, Function, and Evolution
... contacts using residues from the endonuclease domain (93). However, a part of the other domain that is distant from the PI-SceI cleavage site also contributes to the recognition of the target sequence (24, 43). These additional interactions were determined using photo-crosslinking and affinity cleav ...
... contacts using residues from the endonuclease domain (93). However, a part of the other domain that is distant from the PI-SceI cleavage site also contributes to the recognition of the target sequence (24, 43). These additional interactions were determined using photo-crosslinking and affinity cleav ...
Basic region of residues 228-231 of protein kinase CK1[alpha] is
... in these assays was produced by 35S labeling through an in vitro transcription-translation system or by allowing CK1a to autophosphorylate with 32P. Alternatively, the presence of active CK1a bound to axin on the sepharose beads can be assayed by determining its capacity to phosphorylate a specific ...
... in these assays was produced by 35S labeling through an in vitro transcription-translation system or by allowing CK1a to autophosphorylate with 32P. Alternatively, the presence of active CK1a bound to axin on the sepharose beads can be assayed by determining its capacity to phosphorylate a specific ...
Studies on the structure and function of 16S ribosomal RNA using
... (iii) An unexpectedly stable structure has been identified in the region between positions 109 and 279, where many residues remain unreactive even at 90°C in EDTAcontaining buffer. This region may correspond to a structural 'core' that is important for early events in ribosome assembly (Garrett et a ...
... (iii) An unexpectedly stable structure has been identified in the region between positions 109 and 279, where many residues remain unreactive even at 90°C in EDTAcontaining buffer. This region may correspond to a structural 'core' that is important for early events in ribosome assembly (Garrett et a ...
tRNA
... - How many unique 3-digit numerical codes can be obtained using all 10 numerical characters (0-9)? 103! How many 3-letter words are possible using all 26 alpha characters? 263! - Collectively, the 64 codons constitute what has come to be known as the “genetic code”—it is essentially “Genglish” made ...
... - How many unique 3-digit numerical codes can be obtained using all 10 numerical characters (0-9)? 103! How many 3-letter words are possible using all 26 alpha characters? 263! - Collectively, the 64 codons constitute what has come to be known as the “genetic code”—it is essentially “Genglish” made ...
COMPLEX FORMATION AND PROTEIN INTERACTION IN THE
... paralogs, but homology between orthologs (Vergara & Carpita 2001). Other features of possible importance, illustrated in Figure 1, are a cysteine-rich RING-type zinc finger domain located in the N-terminus, the D, D, D, QxxRW highly-conserved catalytic motif characteristic of processive glycosyltran ...
... paralogs, but homology between orthologs (Vergara & Carpita 2001). Other features of possible importance, illustrated in Figure 1, are a cysteine-rich RING-type zinc finger domain located in the N-terminus, the D, D, D, QxxRW highly-conserved catalytic motif characteristic of processive glycosyltran ...
1 Causality, Transfer Entropy and Allosteric
... performs its activity. Protein-protein and protein-DNA interactions, drug action and all processes that depend on signal transduction involve allosteric activity for the system to carry out its normal function. Most known cancer causing mutations lead to the disruption of normal allosteric communica ...
... performs its activity. Protein-protein and protein-DNA interactions, drug action and all processes that depend on signal transduction involve allosteric activity for the system to carry out its normal function. Most known cancer causing mutations lead to the disruption of normal allosteric communica ...
NTPase/helicase of Flaviviridae: inhibitors and inhibition of the
... (Gorbalenya et al., 1989; Lain et al., 1989). The SF2 NTPase/helicases are further divided, according to the sequence surrounding the conserved D-E residues (Walker motif B) in three different subgroups of proteins. The first is formed by the classic D-E-A-D box proteins, and the other are named D-E ...
... (Gorbalenya et al., 1989; Lain et al., 1989). The SF2 NTPase/helicases are further divided, according to the sequence surrounding the conserved D-E residues (Walker motif B) in three different subgroups of proteins. The first is formed by the classic D-E-A-D box proteins, and the other are named D-E ...
Developmentally regulated, alternative splicing of the Rpn10 gene
... amino acid residues, respectively. Thus, each Rpn10 form has a unique structural feature that distinguishes one from another. To understand how the multiple Rpn10 cDNAs are produced, we isolated genomic clones for Rpn10 from a mouse l phage library. Only one Rpn10 gene, designated Psmd4, was identi® ...
... amino acid residues, respectively. Thus, each Rpn10 form has a unique structural feature that distinguishes one from another. To understand how the multiple Rpn10 cDNAs are produced, we isolated genomic clones for Rpn10 from a mouse l phage library. Only one Rpn10 gene, designated Psmd4, was identi® ...
Turnover of protein phosphorylation evolving under
... – with a few spectacular exceptions (Lynch et al., 2011) – the estimates of the relative rate of phosphorylation site evolution based on large samples led to some challenging conclusions. Some studies concluded that phosphorylation sites are generally under strong evolutionary constraints, i.e., tha ...
... – with a few spectacular exceptions (Lynch et al., 2011) – the estimates of the relative rate of phosphorylation site evolution based on large samples led to some challenging conclusions. Some studies concluded that phosphorylation sites are generally under strong evolutionary constraints, i.e., tha ...
The presence of monoglucosylated N196
... no feeding [15,16]. In addition to their role as a storage protein, ...
... no feeding [15,16]. In addition to their role as a storage protein, ...
calculating the structure-based phylogenetic relationship
... Chair and dissertation advisor, Gerald Wyckoff, Ph.D., for his continuous supervision, guidance, time, and patience over the course of my tenure at the School of Biological Sciences. ...
... Chair and dissertation advisor, Gerald Wyckoff, Ph.D., for his continuous supervision, guidance, time, and patience over the course of my tenure at the School of Biological Sciences. ...
SR protein

SR proteins are a conserved family of proteins involved in RNA splicing. SR proteins are named because they contain a protein domain with long repeats of serine and arginine amino acid residues, whose standard abbreviations are ""S"" and ""R"" respectively. SR proteins are 50-300 amino acids in length and composed of two domains, the RNA recognition motif (RRM) region and the RS binding domain. SR proteins are more commonly found in the nucleus than the cytoplasm, but several SR proteins are known to shuttle between the nucleus and the cytoplasm.SR proteins were discovered in the 1990s in Drosophila and in amphibian oocytes, and later in humans. In general, metazoans appear to have SR proteins and unicellular organisms lack SR proteins.SR proteins are important in constitutive and alternative pre-mRNA splicing, mRNA export, genome stabilization, nonsense-mediated decay, and translation. SR proteins alternatively splice pre-mRNA by preferentially selecting different splice sites on the pre-mRNA strands to create multiple mRNA transcripts from one pre-mRNA transcript. Once splicing is complete the SR protein may or may not remain attached to help shuttle the mRNA strand out of the nucleus. As RNA Polymerase II is transcribing DNA into RNA, SR proteins attach to newly made pre-mRNA to prevent the pre-mRNA from binding to the coding DNA strand to increase genome stabilization. Topoisomerase I and SR proteins also interact to increase genome stabilization. SR proteins can control the concentrations of specific mRNA that is successfully translated into protein by selecting for nonsense-mediated decay codons during alternative splicing. SR proteins can alternatively splice NMD codons into its own mRNA transcript to auto-regulate the concentration of SR proteins. Through the mTOR pathway and interactions with polyribosomes, SR proteins can increase translation of mRNA.Ataxia telangiectasia, neurofibromatosis type 1, several cancers, HIV-1, and spinal muscular atrophy have all been linked to alternative splicing by SR proteins.











![Basic region of residues 228-231 of protein kinase CK1[alpha] is](http://s1.studyres.com/store/data/015745345_1-81da768c37e720623b4b1898472e2f2e-300x300.png)










