* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download ppt
United Kingdom National DNA Database wikipedia , lookup
Neocentromere wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Public health genomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
History of genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Population genetics wikipedia , lookup
Microevolution wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Linkage stuff
Vibhav Gogate
A Review of the Genetic Model
Our
View
All except
yellow
nodes
2
1
3
4
E A Thompson
et al’s view
X
S1
X2
S2
X3
S3
Xi-1
Si-1
Xi
Si
Xi+1
Si+1
X
Y1
X2
Y2
X3
Y3
Xi-1
Yi-1
Xi
Yi
Xi+1
Yi+1
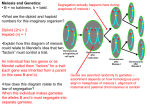
A few other notation
Divide the S variables
Si,j denotes the indicator in meiosis i at location j.
Si,j = 0 if DNA at meiosis i locus j is parent’s maternal
DNA
Si,j=1 if DNA at meiosis i locus j is parent’s paternal
DNA
S.,j = {Si,j| i=1,..,m}
Si,. = {Si,j| j=1,..,l}
Assuming that there are m meiosis and l locations.
More on S
L21m
L22m
L21f
L22f
S23m
S23f
L23f
L23m
X23
X21
The variables in the circle are S.,2
X22
l
P( S ) P( S ,1) P( S , j | S , j 1)
j2
i.e. the set of variables indicating meiosis at locus 2.
Gibbs sampling: Review
X1
X3
X6
X2
X5
X8
X4
X7
X9
Generate T samples {xt} from
P(X|e):
t=1 x1 = {x11,x12,…x1k}
t=2 x2 = {x21,x22,…x2k}
…
After sampling, average:
1
t
P( xi | e) P( x | x \ xi , e)
T t
Gibbs Properties: Review
Good:
Gibbs sampling is guaranteed to converge to P(X|e) as
long as P(X|e) is ergodic:
Shortcoming:
Hard to estimate how many samples is enough
Variance is too big in high- dimensions
Not guaranteed to converge to P(X|e) with
deterministic information
Rao-Blackwellisation (RB)
(Casella & Robert, 1996)
Rao-Blackwellisation provides salvation in
some cases:
Partition X into C and Z, such that we can
compute P(c|e) and P(Z|c,e) efficiently.
Sample from C and sum out Z (RaoBlackwellisation).
Rao-Blackwellised estimate:
1
P( xi | e) P( xi | c t ,e)
T t
Gibbs sampling with RB on
Linkage (E A Thompson et al.)
Two versions
L-sampler
Locus Sampler
RB set is chosen from S.,j
M-sampler
Meiosis sampler
RB set is chosen from Si,.
L-sampler
P(Y ) P(Y | S ) P( S )
S
P( S , j | {S , j ' , j ' j}, Y ) P( S , j | S , j 1, S , j 1, Y , j )
P(Y , j | S , j 1, S , j 1) P(Y , j | S ,j ) P( S , j | S , j , S , j 1)
S , j
l
P( S ) P( S , j ) P( S , j | S , j 1)
j2
A single locus is selected and inheritance indicators at the
locus are updated based on the genotype data at all loci and
on the current realization of inheritance indicators at all loci
other than j.
L-sampler
L-sampler can be implemented on any
pedigree on which single-locus peeling is
feasible
Provided each inter-locus recombination
fraction is strictly positive, the sampler is
clearly irreducible.
However, if the loci are tightly linked, mixing
performance will be poor.
M-sampler
At each iteration a single meiosis is selected and
inheritance indicators for that meiosis are updated
conditional on the genotype data at all loci and the
current realization of inheritance indicators for all
other meioses
P({Si ,; i M *} | Y ,{Si ' ,; i ' M *})
Peel along the chromosome using the Baum Algorithm
with a state space of size 2|M* |
Other Advances
Use of Metropolis Hastings step
Restart
Sequential Imputation
The Actual Bayesnet output by
Superlink
The Variables output by
Superlink
Genetic Loci. For each individual i and locus j, we denote two random
variables Gi,jp, Gi,jm whose values are the specific alleles at locus j in
individual i's paternal and maternal haplotypes respectively.
Marker Phenotypes. For each individual i and marker locus j, a
random variable Pi,j whose value is the specific unordered pair of alleles
measured at locus j of individual i.
Disease Phenotypes. For each individual i, a binary random variable
Pi whose values are affected or unaffected.
Selector Variables. For each individual i and marker locus j, two
binary random variables Si,jp and Si,jm, the values of which are
determined as follows.
If a denotes i's father and b denotes i's mother, then
Si,jp = 0 if Gi,jp = Ga,jp
Si,jp =1 if Gi,jp = Ga,jm
Si,jm is dened in a similar way, with b replacing a.
LOD SCORE
1. Pr(Gi , jp | Ga , jp, Ga , jm, Si , jp), Pr(Gi , jm | Gb , jp, Gb , jm, Si , jm)
2. Pr( Pi , j | Gi , jp, Gi , jm),
3. Pr(Gi , jp)and Pr(Gi , jm)
4. Pr( Pi | Gi , jp, Gi , jm)
5. Pr( Si ,1 p ) Pr( Si ,1m) 0.5, Pr( Si , jp | Si , j 1 p,j 1)and
Pr( Si , jm | Si , j 1m, j 1)
If either j or j - 1 is a disease locus, then the paramter j - 1 is
unknown and is varied to maximize the probabilit y of .
Pedigree data is an assignment e of values to a
subset E of marker and disease phenotype variables
P(e | 1)
LOD ( 1) log 10
P(e | 0.5)
SampleSearch to compute
LOD-SCORE
P(e | 1)
LOD ( 1) log 10
P(e | 0.5)
(1) Compute P(e|θ1) using SampleSearch+Importance Sampling
(2) Compute P(e|θ=0.5) using SampleSearch+Importance Sampling
Computing (1) and (2) is same as computing the probability of
evidence.
SampleSearch-LB
LOD ( 1) log 10
P(e | 1)
P(e | 0.5)
SampleSearch-LB computes a lower bound on the probability of
evidence (i.e. both the numerator and denominator)
Use of Bounding the LOD score.
•If LOD score > 3 then the location is significant
•If we know that the lower bound on the LOD score is 3, then we
have our location
•Unfortunately SampleSearch-LB is not enough as we need an
upper bound on the denominator
•Use SampleSearch in conjunction with Bozhena’s bounding
techniques to upper bound the denominator.