* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Arraying
Survey
Document related concepts
Transcript
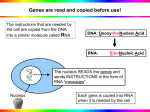
Genomic Arrays – an overview Dr. Colin Campbell The Central Dogma Genome Regulation Transcription DNA Transcription Translation mRNA Protein Genomics in perspective Post Genomic Challenges Sequences available for hundreds of genomes viruses/plasmids >> mammalian genomes Genome sequence only the start Need to understand: genomic structure, replication, expression Problem of scale, complexity and diversity Advent of HTS functional genomic technologies: microarray, Si RNA, mutagenesis, proteomics, imaging Post genomic approaches Functional genomics toolbox Sequence Classify Identify ascribe function Monitor expression all genes used to assemble an organism Microarrays – a post genomic technology Mammalian Genome Database Gene Expression/Genotyping Proteomics Fundamental and applied biomedical research Supporting Technologies Statistics/Bioinformatics HTS Technology Developments: Arraying/ Scanning/ Lab-on-a-chip Computing/ Databases Evolution of array technology Traditional method: taking gene by gene approach Insufficient to meet magnitude of problem Array technology Developed to provide a systematic way of studying RNA expression, genotyping, DNA/ RNA interactions and numerous other applications Array = A regular or uniform arrangement e.g. of DNA probes or other elements such as proteins or tissue sections arranged on glass slides or nylon membranes The Central Dogma Genome Regulation Transcription DNA Transcription Translation mRNA Protein RNA transcription analysis Expression of RNA assessed by Northern blotting, RNAase protection, RT-PCR methods Low to medium throughput approaches. Do not easily accommodate scale, complexity and diversity challenges e.g. Northern Blot Filters exposed to labelled DNA probe and subject to radiography Cell DNA mRNA Denature Gel electrophoresis, RNA separated by Size and blotted on filter proteins RNA transcripts anlysed singly. Definiton of transcriptome would take thousands of blots The microarray solution cDNA(s) or oligonucleotide(s) representative of genes spotted on slide 1 genes 2 3 4 Intensity value 1 Intensity value 2 Array 3 = Relative Value +ve = upreg 4 DNA GENOME Hybridise to array Test cDNA control cDNA DNA mRNA proteins Reverse transcribe RNA Using Cy3 (test RNA) or Cy5 (control) dCTP Relative expression of RNA defined at whole genome level Microarray options First attempts at exploiting array approaches involved filter based screening of clone libraries Basic genomic and RNA expression studies Two key innovations have enhanced the utility of genomic microarrays 1. Use of glass substrates to construct miniaturised arrays DIRECT DEPOSITION: Using automated printers: ~30-40K DNA probe elements deposited on a glass slide IN SITU SYNTHESIS: several million individual DNA probe elements defined by photolithography on silicon wafers 2. The use of fluorescence for detection Method 1. Array of 5,000 mouse genes - direct deposition method The microarray solution cDNA(s) or oligonucleotide(s) representative of genes spotted on slide 1 genes 2 3 4 Intensity value 1 Intensity value 2 Array 3 = Relative Value +ve = upreg 4 DNA GENOME Hybridise to array Test cDNA control cDNA DNA mRNA proteins Reverse transcribe RNA Using Cy3 (test RNA) or Cy5 (control) dCTP Relative expression of RNA defined at whole genome level Direct deposition DNA microarray scanner image Method 2. In situ synthesised oligo array - Affymetrix GeneChip® system Gene Sequence representative DNA sequences derived from 3’ end of gene L 25 mer T G C A T G C A T G C A T G A T G C A T G C A T G Many million fold bound in specific feature 20 features used to represent one gene 400,000 features per array representing ~ 12,000 genes 3’ Affymetrix target labelling Cell/ Tissue of interest 1st strand cDNA synthesis DNA AAA AAA AAA AAA Isolation of total RNA 2nd strand cDNA synthesis AAA TTT TTT AAA TTT TTT AAA TTT TTT AAA TTT TTT T7 Promoter incorporated in first strand synthesis ds cDNA Affymetrix labelling and hybridisation In vitro transcription using Biotinylated dNTPs Biotinylated cRNA Hybridise to Array L TTT b SA b b L TTT b b SA b TTT b L b SA b TTT b L b SA b Affymetrix Gene Chip results Expression of 10K genes – but what is the result ? Statistics and Bioinformatics essential Microarray technology - pros and cons Scale - true global analyses possible Semi-quantitative advantages High throughput Sensitivity Precision Scale demands stringent QC and analytical routines disadvantages Emerging standards for analysis Relative cost/logistics Context independent Microarrays in cancer biology RNA Expression profiling arrays: Targets > pathways Genotyping arrays: HTS SNP analysis > gene association studies Protein arrays: marker sets Expression based classification to detect dominant patterns of expression in heterogeneous tumours Can identify: •Tumour markers •Origin of tumour •Developmental stage •Metastatic potential •Therapeutic response profile •Fundamental insights >> definition of cancer pathways and control •Contribute to diagnosis, prognosis and therapy. Clustered gene sets Interferon related Breast luminal cell profile Basal epithelial cell profile Lung adenocarcinoma enriched profile Proliferation gene set DNA microarrays – a platform technology DNA microarrays now extensively employed for RNA expression profiling studies in biomedical research. Crucial role for statistics, bioinfomatics and computational science to turn HTS data into useful information (gene targets and pathway definition) for the biologist to interpret Provides a critical approach to a thorough understanding of fundamental biological processes. Also contributing to applied areas such as disease diagnosis and definition. DNA microarrays providing a HTS and global platform technology for numerous biomedical and genomic research applications - splicing - sequencing and SNP analysis (v. high density oligo arrays under development) - CGH, BAC clones - epigenetic studies e.g. DNA methylation - Also, platforms developing for: proteins, cells and tissues DNA microarray approaches will ultimately replace many of the standard methods genetic analysis. Biological context Full definition of biological processes requires additional contextual inforrmation (e.g. spatial, temporal, modification) Methods for precise micro sampling of complex cell populations and tissues can be combined with microarray readouts. Initial step involves precise sampling via cell sorting/enrichment or micro-dissection techniques Combine with target sample (micro RNA sample) amplification methods to enable readout on standard DNA microarray platforms Increases power of analysis and biological interpretation Future potential in biology and medicine Array technology will continue to develop for DNA, RNA, protein and various other physiological measurements. Developments will require increasing interface of biology with physical sciences and technology. Allow new questions to be asked at the whole genome/proteome level. Integration of HTS genomic, proteomic and cellular readouts will be required to define biological complexity and approach systems level understanding Key to this is input from bioinformatics and computational science to analyse, store and visualise data