* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture 2 - Pitt CPATH Project
Survey
Document related concepts
Transcript
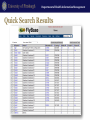
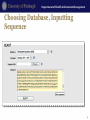
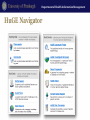
Genomics and Personalized Care in Health Systems Lecture 2. Databases Leming Zhou, PhD School of Health and Rehabilitation Sciences Department of Health Information Management Department of Health Information Management Outline • Nucleotide and protein sequence databases – NCBI • GenBank; RefSeq; dbEST; UniGene – PDB • Flybase • dbSNP • OMIM and HuGE Navigator Department of Health Information Management Molecular Biology Databases • Categories – – – – – – – Nucleotide Sequence Databases Protein Sequence Databases Structure Databases Metabolic and Signaling Pathways Human Genes and Diseases Microarray Data and other Expression Databases … • Each Database contains specific information • Each of these databases is interrelated Nucleotide and Protein Sequence Databases Department of Health Information Management NCBI • Created as a part of National Library of Medicine in 1988 – Establish public databases – Perform research in computational biology – Develop software tools for sequence analysis – Disseminate biomedical information • Databases – Sequence, such as GenBank, RefSeq, dbSNP – Literature, such as PubMed, OMIM • Tools – Entrez. Blast, Cn3D, etc. NCBI Homepage Department of Health Information Management Molecular Databases • Primary Databases – Original submissions by experimentalists – Database staff organize but don’t add additional – Information, for instance, GenBank • Derivative Databases – Human curated • Compilation and correction of data • Example: SWISS-PROT, NCBI RefSeq – Computationally Derived • Example: UniGene Department of Health Information Management GenBank • http://www.ncbi.nlm.nih.gov/Genbank/ • Nucleotide only sequence database • GenBank Data – Direct submissions individual records (BankIt, Sequin) – Batch submissions via email (EST, GSS, STS) – ftp accounts established for sequencing centers • Data shared nightly amongst three collaborating databases: – GenBank – DNA Database of Japan (DDBJ). – European Molecular Biology Laboratory Database (EMBL) 160 140 1990 1991 1992 1993 1994 1995 1996 1997 1998 1999 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 Sequences (million) Base Pairs (billion) Department of Health Information Management Growth of GenBank (1990-2011) Base Pairs Sequences 120 100 80 60 40 20 0 Year Department of Health Information Management GenBank Release 187.0 • ftp://ftp.ncbi.nih.gov/genbank/ • Full release every two months • Incremental and cumulative updates daily Release 181.0 (12/15/2011) • 146,413,798 • 135,117,731,375 Sequences Base Pairs Department of Health Information Management GenBank Record (Header) LOCUS DEFINITION ACCESSION VERSION KEYWORDS SOURCE ORGANISM REFERENCE AUTHORS JOURNAL PUBMED NM_001963 5600 bp mRNA linear PRI 15-JAN-2012 Homo sapiens epidermal growth factor (EGF), transcript variant 1, mRNA. NM_001963 NM_001963.4 GI:296011011 . Homo sapiens (human) Homo sapiens Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Primates; Haplorrhini; Catarrhini; Hominidae; Homo. 1 (bases 1 to 5600) de Diesbach,M.T., Cominelli,A., N'Kuli,F., Tyteca,D. and Courtoy,P.J. TITLE Acute ligandindependent Src activation mimics low EGF-induced EGFR surface signalling and redistribution into recycling endosomes Exp. Cell Res. 316 (19), 3239-3253 (2010) 20832399 Department of Health Information Management GenBank Record (Features) FEATURES Location/Qualifiers source 1..5600 /organism="Homo sapiens" /mol_type="mRNA" /db_xref="taxon:9606" /chromosome="4" /map="4q25" gene 1..5600 /gene="EGF" /gene_synonym="HOMG4; URG" /note="epidermal growth factor" /db_xref="GeneID:1950“ /db_xref="MIM:131530" exon 1..579 /number=1 CDS 453..4076 /codon_start=1 /protein_id="NP_001954.2" /db_xref="GI:166362728“ /db_xref="GeneID:1950“ /db_xref="MIM:131530" /translation="MLLTLIILLPVVSKFSFVSLSAPQHWSCPEGTLAGNGNSTCVGP … exon 580..779 /number=2 exon 780..961 /number=3 Department of Health Information Management GenBank Record (Sequence) ORIGIN 1 61 121 181 241 301 361 421 481 541 601 661 721 781 841 901 961 1021 aaaaagagaa caagggttgt gccttgctct cataagggtg ctgtggcttc ggacaacagc ccgcatctgg ttttcttttg cagtagtttc aaggtactct tctcccatgg tggtggatgc gggtggattt gagtatgtaa ttatttggtc cccacattct ggtttatatt tgggagtgaa actgttggga agctggaact gtcacagtga tcaggtattt ccttggcagg acaacaggag ggtcaatcat aaagttcaaa aaaatttagt cgcaggaaat aaatagtatc tggtgtctca agaaagacaa tatagagaaa aaatcaacag tttaagtgct ttggtcttca ggctctgttg gaggaatcgt ttccatcagt agtcagccag cttactggct ctgcattcag agtaaaagat actcaccttg ctcatcaaga tttgttagtc gggaattcta tttaggattg gtgatcatgg cttttgcaaa aatgtttctg gaaggaatca ttaaaatatc gaggtggctg gagacatcag atctccatat tcttcctttc agcagggctg tccaaagaaa aaggtctctc gccccagggc cccgggccat ttatgctgct tctcagcacc cttgtgtggg acacagaagg attttcatta gagtttttct gaatggcaat ttacagtaac ctgcaaatgt gaagccttta agaaaataac ttcttctttc tttttcctct ttaaactctg catagataaa agttgaagaa tgaggcctcc gctccagcaa cactcttatc gcagcactgg tcctgcaccc aaccaattat taatgagaaa gaatgggtca aaattggata agatatgaaa agcagttgat tagagcagat agctgtgtca agccccaatc ctaagccttt tgaaatttgt gaaatctttc agagcttgga gctcaggcag aatcaagctg attctgttgc agctgtcctg ttcttaattt gagcaattgg agaatctatt aggcaagaga aatgaagaag ggaaataatt ccagtagaaa ctcgatggtg ttggatgtgc Department of Health Information Management FASTA Format >gi|371502116|ref|NM_001126113.2| Homo sapiens tumor protein p53 (TP53), transcript variant 4, mRNA GATGGGATTGGGGTTTTCCCCTCCCATGTGCTCAAGACTGGCGCTAAAAG TTTTGAGCTTCTCAAAAGTCTAGAGCCACCGTCCAGGGAGCAGGTAGCTG CTGGGCTCCGGGGACACTTTGCGTTCGGGCTGGGAGCGTGCTTTCCACG ACGGTGACACGCTTCCCTGGATTGGCAGCCAGACTGCCTTCCGGGTCAC TGCCATGGAGGACCGCAGTCAGATCCTAGCGTCGAGCCCCCTCTGAGTC AGGAAACATTTTCAGACCTATGGAAACTACTTCCTGAAAACAACGTTCTGT CCCCCTTGCCGTCCCAAGCAATGGATGATTTGATGCTGTCCCCGGACGAT ATTGAACAATGGTTCACTGAAGACCCAGGTCCAGATGAAGCTCCCAGAAT Department of Health Information Management Too Many Results Department of Health Information Management Search Limits Department of Health Information Management Reduced Search Results Department of Health Information Management Gene Record Department of Health Information Management RefSeq • Database of reference sequences – http://www.ncbi.nlm.nih.gov/RefSeq/ • Curated – Many experimentally validated – Some partially validated via ESTs – Some computationally predicted • Non-redundant; one record for each gene, or each splice variant, from each organism represented • Status Codes: – Provisional (temporary) – Reviewed – Predicted Department of Health Information Management Department of Health Information Management Accession Numbers • DNA sequences and other molecular data are tagged with accession numbers that are used to identify a sequence or other record relevant to molecular data • RefSeq provides an expertly curated accession number that corresponds to the most stable, agreed-upon “reference” version of a sequence. • RefSeq identifiers include the following formats: – Complete chromosome NC_###### – Genomic contig NT_###### – mRNA (DNA format) NM_######, XM_##### – Protein NP_######, XP_#### Page 26 EST Department of Health Information Management EST • mRNA: Genomic regions actively transcribed in cell • cDNA (complementary DNA) – Copy of mRNA using mRNA as a template – Sequence is complementary to mRNA • EST: Expressed Sequence Tag (a short sub-sequence of a transcribed cDNA sequence) – – – – – Partial cDNA sequence Can be 5’ or 3’ Typical size: 200 - 500 bp Represents mRNA actively transcribed in cell Use to identify • Genes; Alternative splicing; etc. Department of Health Information Management dbEST (release 120111, Dec. 1, 2011) • http://www.ncbi.nlm.nih.gov/dbEST/dbEST_summary.html • Number of Entries: 71,276,166 – Homo sapiens (human) 8,315,294 – Mus musculus (mouse) 4,853,562 – Arabidopsis thaliana (thale cress) 1,529,700 – Danio rerio (zebrafish) 1,488,275 – Drosophila melanogaster (fruit fly) – Gallus gallus (chicken) 821,005 600,433 Department of Health Information Management Access to dbEST Data • EST sequences are included in the EST division of GenBank, available from NCBI by anonymous ftp and through Entrez • The nucleotide sequences may be searched using the BLAST server • EST sequences are also available as a flat file in the FASTA format by anonymous ftp in the /repository/dbEST directory at ftp.ncbi.nih.gov Protein Structure Department of Health Information Management Cn3D: ftp://ftp.ncbi.nih.gov/cn3d/Cn3D-4.3.msi Department of Health Information Management Crystal Structure of A Protein Department of Health Information Management Protein Databases • Proteins have structure and function • InterPro: Protein families and domains http://www.ebi.ac.uk/interpro • Protein Information Resource (PIR): http://pir.georgetown.edu/ • SWISS-PROT/TrEMBL curated protein sequences: http://www.expasy.ch/sprot • UniProt: http://www.expasy.uniprot.org/index.shtml Department of Health Information Management Protein Sequence Motifs Databases • Proteins have conserved regions (motifs, domains) which may have functional significance • Databases exist to store protein families, motifs, and structural domains • CDD: http://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml • Pfam: http://www.sanger.ac.uk/Software/Pfam/ • PROSITE: http://www.expasy.org/prosite Department of Health Information Management Protein Structure Databases • Proteins take on 3D structure • 3D data for some proteins is available due to techniques such as NMR and X-Ray crystallography – PDB http://www.pdb.org/ – SCOP http://scop.mrc-lmb.cam.ac.uk/scop – MMDB http://www.ncbi.nlm.nih.gov/Structure/ Department of Health Information Management PDB (www.pdb.org) • The Protein Data Bank (PDB) is the single worldwide depository of information about the 3D structures of large biological molecules, including proteins and nucleic acids. • Understanding the shape of a molecule helps to understand how it works. • As of January 2010, there are 62,787 searchable structures in the PDB database • PDB provides – Sequence, Atomic Coordinates, Derived geometric data, Secondary structure content, Annotations about protein literature references Department of Health Information Management PDB Statistics Yearly Growth of Total Protein Structures in PDB 90000 80000 70000 60000 50000 40000 Yearly Total 30000 20000 10000 0 http://www.rcsb.org/pdb/statistics/contentGrowthChart.do?content=total&seqid=100 FlyBase http://www.flybase.org Department of Health Information Management FlyBase Introduction Department of Health Information Management Quick Searches Department of Health Information Management Quick Search Results Department of Health Information Management Gene Report Page: gfzf Department of Health Information Management More Details: Gene Model & Product Department of Health Information Management Sequence Searches (BLAST) Department of Health Information Management Choosing Database, Inputting Sequence 41 Department of Health Information Management More BLAST Options Department of Health Information Management BLAST Results Genetic Variations Department of Health Information Management Polymorphisms • Genomic sequences from two unrelated individuals are 99.9% identical. • The 0.1% difference is due to genetic variations, and mainly (~90%) one form of variation called Single Nucleotide Polymorphisms (SNPs, single-base variations). Department of Health Information Management Importance of Genetic Variations • Genetic variations underlie phenotypic differences among different individuals • Genetic variations determine our predisposition to diseases and responses to drugs, therapies, and environmental insults such as bacteria, virus, and chemicals • Genetic variations reveal clues of ancestral human migration history Department of Health Information Management Major Types of Genetic Variations • Single nucleotide mutation – Majority of SNPs do NOT directly contribute to any phenotypes • Insertion or deletion of one or more nucleotides – Tandem repeat polymorphisms (Genomic regions consisting of variable length, usually 1-100 bases long, of sequence motifs repeating in tandem with variable copy number) • Used as genetic markers for DNA finger printing (forensic, parentage testing) • Many cause genetic diseases – Insertion/Deletion polymorphisms (Often resulted from localized rearrangements between homologous tandem repeats) • Gross chromosomal aberration – Deletions, inversions, or translocation of large DNA fragments – Often causing serious genetic diseases Department of Health Information Management SNPs and Mutations • Terminology for variation at a single nucleotide position is defined by allele frequency – A single base change, occurring in a population at a frequency of >1% is termed a single nucleotide polymorphism (SNP) – When a single base change occurs at <1% it is considered to be a mutation • A SNP is a polymorphic position where the point mutation has been fixed in the population • In practice, however, SNPs databases contains multiple types of variations, including SNPs, mutations, insertions, deletions, tandem repeats, copy number variations, etc. Department of Health Information Management SNPs • SNPs can occur anywhere on a genome, they are classified based on their locations. – Many SNPs in genomic, non-coding regions – SNPs in gene regions, including promoter region, coding region, intronic, exonic regioin, UTR, etc. • Often play an important role in differentiation and disease Department of Health Information Management The Effect of SNPs • The phenotypic consequence of a SNP is significantly affected by the location where it occurs (gene or nongene), as well as the nature of the mutation (synonymous or non-synonymous) – No consequence – Affect gene transcription quantitatively or qualitatively – Affect gene translation quantitatively or qualitatively – Change protein structure and functions – Change gene regulation at different steps Department of Health Information Management Simple/Complex Genetic Diseases and SNPs • Simple genetic diseases (Mendelian diseases) are often caused by mutations in a single gene – e.g. Huntington’s, Cystic fibrosis, etc. • Many complex diseases are the result of mutations in multiple genes, the interactions among them as well as between the environmental factors – e.g. cancers, heart diseases, Alzheimer's, diabetes, asthmas, obesity, etc. Department of Health Information Management Sickle Cell Anemia • Due to 1 swapping an A for a T, causing inserted amino acid to be valine instead of glutamine in hemoglobin 1 Normal red blood cells; 2 Sickled red blood cells http://mmcenters.discoveryhospital.com/shared/enc/img_htm/IM-56.htm Department of Health Information Management A Few Relevant Concepts • Allele: A specific “version” of a gene, or an alternative DNA sequences at the same physical locus, which may or may not result in different phenotypic traits • Genotype: the genetic constitution of a cell, an organism, or an individual • Genotyping: the process of identifying what genotype a person has for any given locus (loci) Department of Health Information Management Genetic Variations Databases • dbSNP – http://www.ncbi.nlm.nih.gov/SNP/ • Online Mendelian Inheritance in Man (OMIM) – http://www.ncbi.nlm.nih.gov/omim • International HapMap Project – http://www.hapmap.org/ • Genome Variation Server (Seattle SNPs) – http://gvs.gs.washington.edu/GVS/ Department of Health Information Management dbSNP • The Single Nucleotide Polymorphism database (dbSNP) is a publicdomain archive for a broad collection of simple genetic variations • This collection of polymorphisms includes: – Single-base nucleotide substitutions (or single nucleotide polymorphisms -SNPs) • Roughly 10 million in human population or on average 1 per 300 bps • Less than half of these SNPs are identified and stored in the database – Microsatellite repeat variations (or short tandem repeats - STRs) • In sillico estimation of potentially polymorphic variable number tandem repeats (VNTR) are over 100,000 across the human genome – Small-scale multi-base deletions or insertions • The short insertion/deletions are difficult to quantify and the number is likely to fall in between SNPs and VNTR Department of Health Information Management dbSNP Data Types • The dbSNP contains two classes of records: – Submitted record • The original observations of sequence variation; submitted SNPs (SS) records started with ss – Computationally annotated record • Generated during the dbSNP "build" cycle by computation based the original submitted data, Reference SNP Clusters (ref SNP) start with rs Department of Health Information Management A dbSNP Record >gnl|dbSNP|ss5586300|allelePos=214|len=475|ta xid=9606|alleles='A/G'|mol=Genomic ATAAACATGG ATCAAGTCAT CTTTGAGGAA TTCCAAGTAC TTTAAAGRAG R CTCAAGCAAT GTATTAATGA AGAAACAGAG ACCTGAGGTC AAATAAAAAA TTCTCTCCAT ACTTTTACAA YTGTTAAAAC CATTCAATRT AGTGAGCACA CCA AACCCATATC TAAATGTAAG CACCTGAAAG ATTAGCCGTA GTATACCACC AAAAATCTGC AGAAATGGGA ATAACATTAG ACTTTTTCCC TAGAGGAAAA AATGAGAACA AGAAAATGTT ATTAATGAAG AATAGGTTCC GGCCAAAATT TATAAACAAA GCAAGAATAT A TAGGTTCCAG AGTGATGAAA GAATGCTATG GTCTTCCTGG GAAGAAGTAG TACTAATGAA ACATTCAAGC CTTAGATTAG AAGTAATTGT TTCAGACTGT GTGGGCTCCA AGAACTAGGT GGGTTTTGCA AAGCATCCTG TAATACAGAT Department of Health Information Management International Union of Pure and Applied Chemistry (IUPAC) Code and Meaning IUPAC code A C G T M R W S Y K V H D B N Meaning A C G T A or C A or G A or T C or G C or T G or T A or C or G A or C or T A or G or T C or G or T G or A or T or C Department of Health Information Management Different Ways to Search SNPs in dbSNP • dbSNP web site – Direct search of SS record; batch search; allow SNP record submission; No search limit • Entrez SNP – http://www.ncbi.nlm.nih.gov/sites/entrez?db=Snp – Search limits options allows precise retrieval Department of Health Information Management Search SNPs from dbSNP Web Page • http://www.ncbi.nlm.nih.gov/SNP/index.html Department of Health Information Management dbSNP Search Examples Search using wild-card(*), ranging(:), AND, OR, and NOT operators Example Description BRC*[Gene Name] Search SNPs on all genes with names starting with the letter 'BRC' (ie. BRCA1 and BRCA2) 1[CHR] AND Search SNPs located on chromosome 1 with (frameshift[Function_ function class 'frame-shift' Class]) 1[CHR] OR 2[CHR] Search all SNPs on chromosome 1 or 2 1[CHR] OR 2[CHR] Search all SNPs on chromosome 1 or 2 NOT detected by all methods except 'unknown'. unknown[METHOD] Department of Health Information Management Legend in Results Department of Health Information Management Search dbSNP: Example • Some mutations on human BRCA1 gene have been reported to be involved in the early onset of breast cancer • Retrieve all validated non-synonymous coding reference SNPs for BRCA1 from dbSNP • Starting from the Entrez SNP: http://www.ncbi.nlm.nih.gov/sites/entrez?db=Snp Department of Health Information Management Entrez SNP Search Results Department of Health Information Management dbSNP Ref http://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?rs=799920 Department of Health Information Management SNP Location >gnl|dbSNP|rs799916|allelePos=301|totalLen=601|taxid=9606|sn pclass=1|alleles='A/C'|mol=Genomic|build=130 AAAATAATCA AGAAGAGCAA AGCATGGATT CAAACTTAGG TATTGGAACC AGGTTTTTGT GTTTGCCCCA GTCTATTTAT AGAAGTGAGC TAAATGTTTA TGCTTTTGGG GAGCACATTT TACAAATTTC CAAGTATAGT TAAAGGAACT GCTTCTTAAA CTTGAAACAT GTTCCTCCTA AGGTGCTTTT CATAGAAAAA AGTCCTTCAC ACAGCTAGGA CGTCATCTTT GACTGAATGA GCTTTAACAT CCTAATTACT GGTGGACTTA CTTCTGGTTT CATTTTATAA AAGCAAATCC M GGTGTCCCAA AGCAAGGAAT TTAATCATTT TGTGTGACAT GAAAGTAAAT CCAGTCCTGC CAATGAGAAG AAAAAGACAC AGCAAGTTGC AGCGTTTATA GTCTGCTTTT ACATCTGAAC CTCTGTTTTT GTTATTTAAG GTGAAGCAGC ATCTGGGTGT GAGAGTGAAA CAAGCGTCTC TGAAGACTGC TCAGGGCTAT CCTCTCAGAG TGACATTTTA ACCACTCAGG TAAAAAGCGT GTGTGTGTGT GCACATGCGT GTGTGTGGTG TCCTTTGCAT TCAGTAGTAT GTATCCCACA Department of Health Information Management SNP Fasta Header Format Fasta header line starts with '>' and has fields separated by '|'. Each field is explained below: Gnl Internal use. dbSNP Database name. dbSNP accession for the snp: ss# refers to submitted snp accession. rs# refers ss or rs number to the accession of refSNP cluster of one or more submitted snp. allelePos Variation allele position(1 based) on the fasta. It is always the 5' length plus 1. Total number of bases of the fasta sequence, a sum of length of 5', 3' and len,totalLen variation. Variation is expressed in one IUPack code and has a length of 1 in the totalLen calculation. Only for submitted snp. The two fields after "totalLen" are the submitter handle|submitted_snp_id handle and submitter snp id. Taxid NCBI taxonomy id Molecular source of the sequence. Valid values are: genomic, cDNA or Mol mitochondria. Variation class of the snp, most common value is 1 - single nucleotide snpclass: polymorphism. Click on "snpclass" for details. Alleles Lists alleles of the snp separated by '/'. Sequence in lower case is used for sequence identified by RepeatMasker as Lower or upper case low-complexity or repetitive elements. ATCG Green color is used for assay sequence (observed by the submitter). ATCG Black color is used for flank sequence (extracted from sequence databases ). Header Department of Health Information Management GeneView of a SNP Department of Health Information Management Links to Various Gene Records Gene and Disease Department of Health Information Management Disease Causing Genes Disease centric databases: • OMIM: http://www.ncbi.nlm.nih.gov/omim/ • CDC HugeNavigator: http://hugenavigator.net/ • HGMD: https://portal.biobaseinternational.com/hgmd/pro/start.php • A Catalog of Published Genome-Wide Association Studies: http://www.genome.gov/26525384 Department of Health Information Management NCBI—OMIM Department of Health Information Management Online Mendelian Inheritance in Man (OMIM) • http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=OMIM • OMIM is a human genetic disorders database built and curated using results from published studies • Each OMIM record provides a summary of the current state of knowledge of the genetic basis of a disorder, which contains the following information: – description and clinical features of a disorder or a gene involved in genetic disorders; biochemical and other features; cytogenetics and mapping; molecular and population genetics; diagnosis and clinical management; animal models for the disorder; allelic variants. • OMIM is searchable via NCBI Entrez, and its records are cross-linked to other NCBI resources. Department of Health Information Management OMIM: Variant • The OMIM database includes genetic disorders caused by various mutation/variation, from SNPs to large-scale chromosomal abnormalities • Variants are represented by a 10-digit OMIM number, and can be searched in two ways – Search for a gene or a disease, when retrieved, view its variants Department of Health Information Management Variants in OMIM Records • For most genes, only selected mutations are included – Criteria for inclusion include: the first mutation to be discovered, high population frequency, distinctive phenotype, historic significance, unusual mechanism of mutation, unusual pathogenetic mechanism, and distinctive inheritance. • Most of the variants represent disease-producing mutations, NOT polymorphisms • A few polymorphisms are included, many of which show a positive statistical correlation with particular common disorders • Few neutral polymorphisms are included in OMIM • Some SNPs in the dbSNP records are not linked to the corresponding OMIM records Department of Health Information Management Office of Public Health Genomics, CDC • The CDC established the Office of Public Health Genomics (OPHG) in 1997. • OPHG aims to integrate genomics into public health research, policy, and programs. Doing so could improve interventions designed to prevent and control the country’s leading chronic, infectious, environmental, and occupational diseases. • OPHG's efforts focus on • conducting population-based genomic research • assessing the role of family health history in disease risk and prevention • supporting a systematic process for evaluating genetic tests • translating genomics into public health research and programs • strengthening capacity for public health genomics in disease prevention programs Department of Health Information Management HuGENet • The Human Genome Epidemiology Network (HuGENet™) : – Established to help translate genetic research findings into opportunities for preventive medicine and public health by advancing the synthesis, interpretation, and dissemination of population-based data on human genetic variation in health and disease. • HuGENetTM resources: – HuGE Navigator, Coordinating centers, Collaborators, Workshops, Reviews, Case studies, Book • HuGE Navigator provides access to a continuously updated knowledge base in human genome epidemiology – information on population prevalence of genetic variants – gene-disease associations – gene-gene and gene- environment interactions Department of Health Information Management HuGE Navigator Department of Health Information Management Finding Disease Causing Genes Department of Health Information Management Finding Gene’s Associated Diseases Department of Health Information Management Disease Databases • Genes are involved in disease • Many diseases are well studied • Description of diseases and what is known about them is stored – OMIM: http://www.ncbi.nlm.nih.gov/Omim/ – Tumor Gene Family Databases: http://www.tumorgene.org/tgdf.html Department of Health Information Management Homework 1 • Using PubMed, search for a recent paper related to genetic disease (or disease with known genetic basis) and with a research method named “Genome-Wide Association Study or GWAS”. The title of the journals could be “PLoS genetics”, “Nature Genetics”, etc. The genetic disease could be obesity, type II diabetes, etc. • Choose one or a few SNPs record or genes mentioned in the paper which may highly relevant to the disease; search for the corresponding dbSNP records in dbSNP. Summarize the genetic variation, relevant genes, and location of the variation. • Search for the Genbank record of the relevant genes, extract and save their sequences in FASTA format. Identify the corresponding protein sequences and use Cn3D to display the structure of the protein.