* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture2-2010
Magnesium transporter wikipedia , lookup
Interactome wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Western blot wikipedia , lookup
Peptide synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Point mutation wikipedia , lookup
Genetic code wikipedia , lookup
Metalloprotein wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
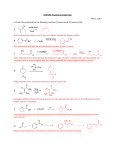
A one-dimensional (1D) NMR spectrum of a protein H HN H H 2O H CO N Methyl C CO Backbone HN H Aromatics (H N ) (H ) ref 600134800 600132400 600130000 600.130000 (Hz) (MHz) (ppm) (ref ) / ref 9 8 1H 7 6 5 4 chemical shift (ppm) 3 2 1 0 Upfield shifted methyls Chemical shifts in parts per million (ppm) Are independent of the field strength of the Static magnetic Bo field. See the supplementary lecture material and Rattle, ‘NMR Primer for Life Scientists Pages 19-21, 26. 1 Y215Q 2 H +H3N C COO- H C H + 4 C H N 1 C C2 H N3 H H 8.0-8.8 ppm A lot of work done with histidine since the C2 proton appears at higher frequency than most other protons. 6.8-7.2 ppm H1 = His105 H2 = His119 H3 = His12 H4 = His48 Measure pKa of each histidine pKa His105 6.7 His119 6.2 His12 5.8 His48 is more complex, sudden discontinuity in the curve. Repeat titrations in the presence of an inhibitor. His105 in this case, cytidine-3’monophosphate (3’-CMP) His48 O N O OPO3- His12 His119 HOCH2 NH2 OH His48 and His105 are unchanged His12 and His119 curved are shifted downfield. His119 changes from 6.2 to 8.0 His 12 changes from 5.8 to 7.4 Why downfield?? Both His12 and His119 are protonated in the enzymeinhibitor complex. The proton is protected from exchange by the presence of the inhibitor. Need to go to higher pH to remove it. NH2 HOCH2 O N O OPO3- OH A simple 1D NMR Spectrum - this is ethanol but the spectrum has the sort of simplicity we might get with a one amino acid protein (there is no such thing!) A 36 amino acid protein A successful NMR experiment comes in 3 stages, 1.) Resolve the resonances - that is, obtain a spectrum where individual signals are clearly resolved from one another. 2.) Assign the resonances. Each peak comes from one atom in the protein - but which one? Our 36 amino acid protein is a mess! The record to date is 723 amino acids With full assignment of the spectrum - how did they do this? 3.) Interpret the data. Effect of increasing spectrometer frequency 1 GHz soon?? COSY Spectrum of a small protein Areas of Spectrum Typical Amino Acid spin-system patterns on COSY spectra 1.) Just see 3J coupling 2.) Do not see couplings across the peptide bond.