* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Zinc finger nuclease wikipedia , lookup
Primary transcript wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Transcription factor wikipedia , lookup
Protein moonlighting wikipedia , lookup
Histone acetyltransferase wikipedia , lookup
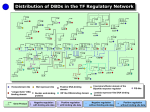
Domain Assignment to Transcription Factors 416 Proteins with at least one SCOP DBD assignment 416 proteins with DBD assignment PFAM assignments Remove 145 proteins (Transposases, Replication/Repair and Restriction Enzymes) 271 Transcription Factors 113 with regulated gene information + 158 with DBD To gain insight into the evolution and organisation of the transcription factors in E. coli, we used the information available in RegulonDB (Salgado et al., 2001) on TFs and their regulated genes. To find out about the domain architecture and family membership of E. coli TFs, we used the SUPERFAMILY (Gough et al., 2001) database of structural assignments to predicted proteins from the E. coli genome, and extracted all those with DNA-binding domains (DBDs). Using the functional annotation for the E. coli genome, we removed proteins involved in replication & repair, restriction enzymes and transposases. Repertoire of DNA Binding Domains in E. coli TFs 1efa 1jhf “Winged” helix Lambda repressor-like 1jhg Trp repressor 1ihf IHF-like DNA-binding proteins 1rnl 1etx 1bl0 C-terminal effector domain of the bipartite response regulator 1cmb Homeodomain-like FIS-like 1mjc Met repressor-like Nucleic acid binding protein 1g8e Flagellar transcriptional activator FlhD 1exi Putative DNA binding protein Domain Analysis of E. coli TFs Small Molecule Binding Domain 2% 13% 9% 1 Domain TFs Enzyme Domain 22% 2 Domain TFs Receiver Domain 44% 3 Domain TFs 76% 12% Protein Interaction Domain 7% 4 Domain TFs 11% 5% DNA Binding Domain Domain of unknown function On average each TF has two domains: one DBD and a partner domain which is generally the control domain The activity of ~90% of transcription factors is controlled by a second domain and only ~10% is controlled at the transcriptional level In E. coli, there are 271 TFs that belong to 11 DBD families and 46 different partner domains as identified in SUPERFAMILY. The domains can be broadly classified into six functional classes as mentioned above. About two thirds of the 271 proteins have the same domain architecture as at least one other protein, and have thus arisen as a consequence of duplication of a complete gene (Apic et al., 2001 show that proteins with identical domain architecture are highly likely to be direct duplicates.) On average, each TF is made up of two domains, and the neighbouring domains to the DBD are of the small molecule binding type in 120 of the 240 multidomain proteins.