* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download DNA is a double helix
United Kingdom National DNA Database wikipedia , lookup
Eukaryotic DNA replication wikipedia , lookup
DNA nanotechnology wikipedia , lookup
Microsatellite wikipedia , lookup
Homologous recombination wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Helitron (biology) wikipedia , lookup
DNA replication wikipedia , lookup
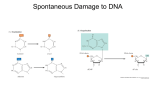
DNA is a double helix DNA damage and repair • Sources of DNA damage – Spontaneous – Environmental • Radiation • Carcinogens – Mistakes in replication Spontaneous DNA damage • Deamination of bases… 1. 2. Converts C to U etc… Altered base has different base pairing rule – 3. e.g. U pairs with A (converts CG bp to UA) Unless repaired results in transition mutation Base loss • Some times bases just fall off (more often than you might think; 10000/genome/generation) • Bases gone, but phosphodiester backbone is still intact • Purines more sensitive than pyrimidines (acid sensitive) • Causes mutation, can lead to strand breaks ROS • Reactive oxygen species; things that are or give rise to oxygen with an unpaired electron; a free radical • E.g hydroxyl radical H O • ROS produced by…. – Respiration – Radiation (ionizing, non-ionizing) • ROS removed by… – Enzymes…SOD, catalase – Reducing agents..glutathione, vitamin E, etc. Oxidative damage 8-oxoG Thymine glycol Radiation:ionizing • X-rays, gamma-rays; rarely encountered… except for medical sources – Usually damage is secondary consequence of ROS generated after radiolysis of water (DNA rarely a direct target) – Damages bases (e.g. 8-oxoG) – Strand breaks; often clustered, thus a source of double strand breaks Radiation: UV light • Non-ionizing radiation (UV light from the sun) – Bases absorb energy with peak at 260nM..this is UV – Photoactivates base, causes nasty chemistry – Result is…covalent bonds between adajacent bases, almost always adjacent pyrimidines – Distorts DNA (kink), can block transcription, replication, lead to mutation T T T C Adduct formation • Nasty chemicals(carcinogens) that glom groups on to DNA; often ring Nitrogens in bases – E.g. Alkylating agents: reactive carbon containing chemicals (ethylating agents, methylating agents) • Not always direct exposure: sometimes carcinogen is toxic product of cellular metabolism – Cigarette smoke; benzo-a-pyrene not a big deal…but the break down product is • Groups are bulky, blocks transcription, replication; can interfere with base pairing, and introduce mutation during replication Cross-linking agents • Cross-linking agents; a special case where adductformer is bifunctional (two reactive groups) – Intra-strand crosslinks: between adjacent nucleotides, like UV photoproducts – Inter-strand crosslinks: between nucleotides on opposite strands of a double helix • Interstrand cross linkers completely block transcription, replication Replication errors • Not DNA damage per se. • Substitutions – Misincorporation: <1X10-6 – Improved by proofreading, mismatch repair – Made worse by imbalanced nucleotide pools – Tautomers • Slippage – Repetitive DNA, secondary structures Tautomers • Standard tautomers…keto T, G; amino A, C. • rare tautomers (enol T, G; iminoA, C) • Results in nonwatson crick base pairs • Transition mutation Microsattelite instability • Slippage of primer or template during replication causes expansion/contraction of microsattelite DNA is a double helix DNA is a double helix Critical (if somewhat obvious) thing to remember for both replication and repair: Symmetry (base pairing) allows for easy and accurate 1. Replication: duplication of information 2. Repair: replacement of damaged information DNA REPAIR Repair Pathway Base Excision Repair (BER) Nucleotide Excision Repair (NER) Translesion synthesis (TLS) Mismatch Repair (MMR) Homologous Recombination (HR) End joining (EJ) Major functions Deaminations, Depurinations UV photoproducts, Adducts, Cross-links Bypass all of the above Replication errors Double strand breaks, Adducts, Cross-links Double strand breaks Excision repair=BER, NER, MMR DSB repair=HR, EJ Excision repair • Damage in one strand….remove it, fill in the gap (using un-damaged strand as template), ligate the remaining nick • Same process…base excision repair, nucleotide excision repair, mismatch repair • Contrast to double strand break repair (which has no template for repair) Base excision repair (BER) • Damage is recognized by glycosylase (different one for each type of damage) • Common targets are deamination products • E.g. uracil glycosylase, hypoxanthine glyosylase Glycosylase recognition Nucleotide excision repair (NER) • Recognizes distortion in DNA (more flexible than BER) • Removes most UV photoproducts, adducts • Multi-protein machine NER: recognition of damage Excision site Cyclobutane dimer Excision site NER: Transcription coupled repair • NER most efficient in transcribed regions; also template strand more efficiently repaired than nontemplate strand • NER machinery part of transcribing RNA polymerase complexes (TFIIH) • When NOT coupled to transcription, NER can still be targeted, though less efficiently, to damage in “silent” DNA…global genome repair (GGR) Mismatch repair • Supresses replication errors; substitutions, slippage (microsattelite instability) • Unlike NER/BER, not obvious which is damage, which should be used as template • Need to identify recently synthesized strand Double strand break repair and recombination • How do you get double strand breaks? – Can be intentional (developmentally programmed) • Meiosis, VDJ recombination – By accident • Ionizing radiation, replication through incompletely repaired damage – Therapeutic • Chemotherapy, radiation therapy – Malicious intent • Transposons, retro-elements How do your repair double strand breaks? 1. Homologous recombination (“HR”) • • Meiotic recombination vs. Mitotic recombination Used almost exclusively in prokaryotes, by far the more important even in yeast 2. End joining (“EJ”) • • Less accurate, but the primary pathway in vertebrates Why use end joining over homologous recombination? DNA repair BER/SSBR" DSBR" IR" VDJ" Excision HR" Synthesis" NHEJ" No template" Ligation Accurate! Cheap! Accurate! Expensive! Inaccurate? Cheap?! Vignette: synthesis with a broken template • DSBs have… – Both strands broken • Loss of chromosome continuity • No intact template – Damage in flanking nucleotides • More than just ligation NHEJ 5ʼ" AP 3’PG 5ʼ" NHEJ core factors AP sites, gaps block ligation 2. Base excision repair at broken ends? – – Need damage specific excision “creative” DNA polymerase Synthesis during BER 5ʼ" 5ʼ" NHEJ core factors • Missing at DSB: Stably annealed primer/template – At DSB, pol must work with scaffold that helps hold primer/template together Pol X in mammals Pol X Domains*" Role" BRCT pol pol b BER (NHEJ?)" TdT" NHEJ" pol l BER, NHEJ" pol ! NHEJ " lyase • TdT, Pol l, and pol m all form a complex with Ku/XRCC4-ligase IV • All three involved in NHEJ Why so many? A CG 3’ A CG • Pol m, Pol l vs. pol b – BRCT domain, and interaction with Ku/ XRCC4-ligase IV, essential GCA 3’ GCA -" b l m • In vitro NHEJ: Ku, XRCC4ligase IV, polymerase Why so many? pol" Pol m vs. Pol l With Primer/template pairing" A CG 3’ GCA A CG 3’ GCA Without Primer/template pairing" A G 3’ 3’ G A 3’ G A A G 3’ -" l m A gradient in template strand requirements 5ʼ" 5ʼ" • Pol b > Pol l > Pol m > TdT BER BER/NHEJ NHEJ/V(D)J VDJ Different substrate requirements dictate their biological roles" Structural basis for the “gradient” 1: Loop 1 b3 b4 Pol b SKG------------------ETKFM Pol l VSQEEN-------------GQQQKYL! Pol m HQHSCCESPTRLA-QQSHMDAFERSF! TdT LVESTFEKLRLPSRKVDALDHFQKCF Pol b Pol b, l, m, TdT a l p m Te nd a r t s te " nd a r t s er Prim dNTP" " Pol X polymerases: getting by with less..."and less..."and less..." H329 R175 + 5ʼ" 5ʼ" Loop NHEJ core factors • Pol b > Pol l BER > Pol m > TdT" BER/NHEJ NHEJ/VDJ VDJ Questions • Why do NER and BER excise damage differently? • Why couple NER to transcription? • NHEJ is less accurate than HR: Why do you bother with NHEJ? • NHEJ has some problems unique amongst DNA repair pathways…what are they?