* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Transition
Glass transition wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Ligand binding assay wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Ultrasensitivity wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Proteolysis wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Biochemistry wikipedia , lookup
Metalloprotein wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
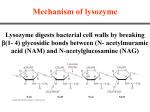
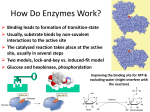
Mechanism of Enzymes Mechanisms the molecular details of catalyzed reactions Nucleophilic substitution reactions • Nucleophilic species are electron rich, and electrophilic species are electron poor • Types of nucleophilic substitution reactions include: (1) Formation of a tetrahedral intermediate by nucleophilic substitution (2) Direct displacement via a transition state Two types of nucleophilic substitution reactions • Direct displacement Two types of nucleophilic substitution reactions • Formation of a tetrahedral intermediate Cleavage reactions • Carbanion formation • Carbocation formation Cleavage reactions • Free radical formation Catalysts Stabilize Transition States Energy diagram for a single-step reaction Energy diagram for reaction with intermediate • Intermediate occurs in the trough between the two transition states • Rate determining step in the forward direction is formation of the first transition state Enzymes lower the activation energy of a reaction (1) Substrate binding • Enzymes properly position substrates for reaction (makes the formation of the transition state more frequent and lowers the energy of activation) (2) Transition state binding • Transition states are bound more tightly than substrates (this also lowers the activation energy) Enzymatic catalysis of the reaction A+B A-B Chemical Modes of Enzymatic Catalysis A. Polar Amino Acid Residues in Active Sites • Active-site cavity of an enzyme is lined with hydrophobic amino acids • Polar, ionizable residues at the active site participate in the mechanism • Anions and cations of certain amino acids are commonly involved in catalysis Acid-Base Catalysis • Reaction acceleration is achieved by catalytic transfer of a proton • A general base (B:) can act as a proton acceptor to remove protons from OH, NH, CH or other XH • This produces a stronger nucleophilic reactant (X:-) General base catalysis reactions (continued) • A general base (B:) can remove a proton from water and thereby generate the equivalent of OH- in neutral solution Proton donors can also catalyze reactions • A general acid (BH+) can donate protons • A covalent bond may break more easily if one of its atoms is protonated (below) Covalent Catalysis Step one: a glucosyl residue is transferred to enzyme *Sucrose + Enz Glucosyl-Enz + Fructose Step two: Glucose is donated to phosphate Glucosyl-Enz + Pi Glucose 1-phosphate + Enz *(Sucrose is composed of a glucose and a fructose) Triose Phosphate Isomerase (TPI) • TPI catalyzes a rapid aldehyde-ketone interconversion Proposed mechanism for TPI • General acid-base catalysis mechanism (4 slides) Proposed mechanism for TPI Proposed mechanism for TPI Proposed mechanism for TPI Energy diagram for the TPI reaction Binding Modes of Enzymatic Catalysis • Proper binding of reactants in enzyme active sites provides substrate specificity and catalytic power • Two catalytic modes based on binding properties can each increase reaction rates over 10,000-fold : (1) Proximity effect - collecting and positioning substrate molecules in the active site (2) Transition-state (TS) stabilization - transition states bind more tightly than substrates Binding forces utilized for catalysis 1. Charge-charge interactions 2. Hydrogen bonds 3. Hydrophobic interactions 4. Van der Waals forces The Proximity Effect • Correct positioning of two reacting groups (in model reactions or at enzyme active sites): (1) Reduces their degrees of freedom (2) Results in a large loss of entropy (3) The relative enhanced concentration of substrates (“effective molarity”) predicts the rate acceleration expected due to this effect Reactions of carboxylates with phenyl esters Reactions of carboxylates with phenyl esters Reactions of carboxylates with phenyl esters Weak Binding of Substrates to Enzymes • Energy is required to reach the transition state from the ES complex • Excessive ES stabilization would create a “thermodynamic pit” and mean little or no catalysis • Most Km values (substrate dissociation constants) indicate weak binding to enzymes Energy of substrate binding • If an enzyme binds the substrate too tightly (dashed profile), the activation barrier (2) could be similar to that of the uncatalyzed reaction (1) Transition-State (TS) Stabilization • An increased interaction of the enzyme and substrate occurs in the transition-state (ES‡) • The enzyme distorts the substrate, forcing it toward the transition state • An enzyme must be complementary to the transition-state in shape and chemical character • Enzymes may bind their transition states 1010 to 1015 times more tightly than their substrates Transition-state (TS) analogs • Transition-state analogs are stable compounds whose structures resemble unstable transition states • 2-Phosphoglycolate, a TS analog for the enzyme triose phosphate isomerase Induced Fit • Induced fit activates an enzyme by substrateinitiated conformation effect • Induced fit is a substrate specificity effect, not a catalytic mode • Hexokinase mechanism requires sugar-induced closure of the active site Glucose + ATP Glucose 6-phosphate + ADP Lysozyme Binds an Ionic Intermediate Tightly • Lysozyme binds polysaccharide substrates (the sugar in subsite D of lysozyme is distorted into a half-chair conformation) • Binding energy from the sugars in the other subsites provides the energy necessary to distort sugar D • Lysozyme binds the distorted transition-state type structure strongly Bacterial cell-wall polysaccharide • Lysozyme cleaves bacterial cell wall polysaccharides (a four residue portion of a bacterial cell wall with lysozyme cleavage point is shown below) Conformations of N-acetylmuramic acid (a) Chair conformation (b) D-Site sugar residue is distorted into a higher energy halfchair conformation Lysozyme reaction mechanisms 1. Proximity effects 2. Acid-base catalysis 3. TS stabilization (or substrate distortion toward the transition state) Mechanism of lysozyme Properties of Serine Proteases Zymogens Are Inactive Enzyme Precursors • Digestive serine proteases including trypsin, chymotrypsin, and elastase are synthesized and stored in the pancreas as zymogens • Storage of hydrolytic enzymes as zymogens prevents damage to cell proteins • Zymogens are activated by selective proteolysis Activation of some pancreatic zymogens Substrate Specificity of Serine Proteases • Many digestive proteases share similarities in 1o,2o and 3o structure • Chymotrypsin, trypsin and elastase have a similar backbone structure • Active site substrate specificities differ due to relatively small differences in specificity pockets Binding sites of chymotrypsin, trypsin, and elastase • Substrate specificities are due to relatively small structural differences in active-site binding cavities Identification of His at active site • The irreversible inhibitor (TosPheCH2Cl) binds to the active-site His residue in serine proteases Catalytic triad of chymotrypsin • Imidazole ring (His-57) removes H from Ser-195 hydroxyl to make it a strong nucleophile (-CH2O-) • Buried carboxylate (Asp-102) stabilizes the positivelycharged His-57 to facilitate serine ionization a-Chymotrypsin mechanism (Acyl E + H2O) (E-TI2) (E-P2) (E + P2)