* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Chem 301 Biological Chemistry I Laboratory Lab 7: Protein
Survey
Document related concepts
Magnesium transporter wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Fluorescence wikipedia , lookup
Interactome wikipedia , lookup
Metalloprotein wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Metabolomics wikipedia , lookup
Protein purification wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Protein structure prediction wikipedia , lookup
Transcript
Chem 301
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
PROTEIN SPECTROSCOPY
INTRODUCTION
A multitude of spectroscopic methods have been developed and/or optimized in
order to characterize proteins with respect to their structure, the nature of their
interactions with other molecules (e.g. DNA, other proteins, and small molecule ligands
and reaction substrates), their local solvent environment, conformational changes that
they undergo, and many other aspects of their chemistry.
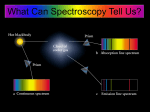
Spectroscopy explores the interaction of energy with matter. Absorption of
electromagnetic radiation results in transitions from lower energy levels to higher energy
levels. Excited states are unstable and decay back to the ground state, giving off
energy, often in the form of light. The types of spectroscopy that are commonly used to
study proteins include, but are not limited to: Electronic spectroscopies including
ultraviolet (UV)-Vis(ible) spectroscopy, fluorescence spectroscopy and circular
dichroism (CD) spectropolarimetry, Vibrational spectroscopies, including infrared (IR)
spectroscopy and particularly Raman spectroscopy, nuclear magnetic resonance (NMR)
spectroscopy, mass spectrometry (MS), and electon paramagnetic resonance (EPR)
spectroscopy also known as electron spin resonance (ESR) spectroscopy. Different
types of spectroscopy involve different regions of the electromagnetic spectrum (see
Figure 1) based on the type of energy transition being probed. Based on these
differences, the different analytical tools probe motions and/or events that occur on
different time scales (Figure 2).
Figure 1: EM spectrum (fig taken from Chemistry: The Central Science Brown, LeMay, and Bursten, 10
ed. P. 219.
th
Chem 301
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
molecular
transition:
Nuclear
10
20
10
19
Electronic
(inner e-)
10
electronic
(valence e-)
10
10
10
10
vibrational
10
Page 2 of 15
region of
spectrum:
Gamma
10
-8
10
-7
8
10
7
10
6
10
-6
10
5
10
-5
10
4
10
-4
10
3
UV
visible
10
-3
10
2
IR
10
-2
10
-1
18
X-ray
17
type of
spectroscopy:
Gamma ray emission
X-ray absorption,
emission, diffraction
16
15
14
13
10
12
10
11
10
10
10
9
10
8
10
7
10
6
10
5
10
4
10
3
10
2
Far IR
UV absorption,
emission, fluorescence
IR absorption, Raman
scattering
1
1
electron
spin
nuclear spin
-9
10
10
rotational
10
10
cm
-1
microwave
radiofrequency
10
2
10
3
10
4
10
5
10
6
10
7
10
8
microwave absorption
electron spin resonance
NMR
10
1
cm
c/s
Figure 2: Regions of the EM spectrum and the corresponding molecular transitions and types of
spectroscopy.
UV-VIS SPECTROPHOTOMETRY
Proteins have various chemical moieties that absorb light at different
Chem 301
Page 3 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
wavelengths. Some proteins, e.g. heme proteins such as hemoglobin, myoglobin, and
cytochrome c have a prosthetic group (heme) that absorbs visible light due to the high
level of conjugation (see Figure 3a). The light that is not absorbed is transmitted and
solutions containing these proteins (in their holo forms with the prosthetic groups intact)
appear that (transmitted) color, the complement of the color absorbed. For example,
oxyhemoglobin appears bright red (oxyhemoglobin is the reason oxygenated blood is
bright red) because it absorbs blue/green light (λ ~ 540-590 nm) strongly, transmitting
red light.
(a)
(b)
Figure 3 (a) heme and (b) the aromatic amino acids phenylalanine, tyrosine, tryptophan, and histidine.
There are several aromatic amino acids which absorb UV light near the visible
region of the spectrum (see Figure 3b). The more conjugated the side chain, e.g.
tryptophan, the more red-shifted the absorbance is, which is why tryptophan absorbs
light at a longer wavelength (lower energy) than phenylalanine, which absorbs longer
wavelength light than histidine. Most proteins have at least one tryptophan residue,
which is the basis for using the absorbance at 280 nm as a measure of protein
concentration and why for DNA extraction procedures, the ratio of A260 (the λmax for
DNA) to A280 (where DNA has no appreciable absorbance) is used to assess the level of
protein contamination in a DNA sample.
The spectrophotometer is widely used in biochemistry laboratories. Its basic
setup is shown below in Figure 4. The image of the filament of an incandescent lamp is
focused on the entrance slit by a lens. The divergent light emerging from this slit is
focused, after wavelength selection by the diffraction grating (or monochromator), on the
exit slit. The monochromatic light then passes through the sample in its cuvette and
falls on the electron- emitting element of a photomultipier tube (PMT). The resulting
electrical signal, which is proportional to the number of photons hitting the sensitive
element of the PMT, is amplified and used to drive a microammeter on the upper panel
Chem 301
Page 4 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
of the instrument. More modern instruments convert the electrical signal to a digital
display and some instruments use a diode array detector to simultaneously measure
light at many wavelengths.
phototube
th
Figure 4. A UV-Visible spectrophotometer (fig from Lehninger Princ of Bioch., Nelson & Cox, 4 ed, ch 3)
The instrument can be set to display either % transmittance or absorbance (optical
density or O.D. in older literature and still in use in biological applications.) An
absorbance value relates the amount of monochromatic light which is absorbed by the
sample. Percent transmittance is the percent of light that goes through the sample
relative to that which passes through a blank. (The instrument is adjusted to 100%
transmittance using the blank.) This scale is simple to understand, but is rarely used
because the % transmittance does not vary directly with the concentration of absorbing
material as does the absorbance.
The Molar Extinction Coefficient (ε) is a measure of how much light of a particular
wavelength that a molecule will absorb. ε is defined as the absorbance that a 1 molar
solution of that compound with a 1 cm path length through the solution would have at a
particular wavelength (the wavelength is often subscripted next to the symbol, e.g. ε
refers to the extinction coefficient at 600 nm… typically λmax is used as long as there is
minimal interference at that wavelength from other solution components). This is a
calculated value since the absorbance of a 1 M solution is almost always too high for
instruments to measure.
Beer's law states that the extent of absorption (A) of light by a solution of a
colored solute is a function of the concentration (c) of the solution if the path length (l)
through the solution and the wavelength of light are kept constant. It states that the
intensity of a ray of monochromatic light decreases exponentially as the concentration of
the absorbing medium increases. Absorbance, a logarithmic function, is directly
proportional to the concentration: A = ε c l
Beer's law applies to monochromatic light and holds true only when changes in
concentration are not accompanied by other changes that affect the nature of the solute.
Not all colored materials follow Beer's law and those that do may not over a large range
of concentrations… deviations typically occur at high concentrations.
600
FLUORESCENCE SPECTROSCOPY
Chem 301
Page 5 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
When matter absorbs electromagnetic radiation, the “normal” distribution of high
energy (excited) and lowest energy (ground) states is disrupted because the absorbed
energy will induce many ground state à excited state transitions. The majority of these
newly excited states will immediately decay back to their ground state electron
configuration, giving off the energy that was just absorbed. Some of the states in the
system, however will lose only some energy in non-radiative processes (those that do
not emit radiation) and will decay to a lower (but still excited) energy state. Within each
electronic energy level are all of the vibrational energy levels (see Figure 5). Nonradiative decays often result in decay back to the lowest vibrational state within a given
excited electronic state. When the resulting state eventually decays back to the ground
state, the energy emitted is lower in energy (and therefore longer in wavelength) than
that which was initially absorbed. This process of absorbing shorter wavelength light
(often in the UV region), decaying non-radiatively and then ultimately emitting longer
wavelength light (often in the visible region) is called fluorescence.
NON-RADIATIVE
DECAY
FLUORESCENCE
ABSORBANCE
Figure 5. Diagram of electronic transitions that occur in fluorescence.
Fluorescence efficiency is quantitated as a value known as quantum yield, Q and
is represented by the fraction of photons there were absorbed that are ultimately
emitted:
photonsemitted
Q=
photonsabsorbed
Q can approach efficiencies of 1 but are typically in the range of 0.3-0.7 for biological
samples. An instrument used to measure fluorescence is called a fluorimeter (see
Figure 6). It is designed so that excitation light is passed through a sample and the
resulting emission is detected at a right angle (otherwise the excitation light that is
transmitted through the sample would interfere with the light emitted). This is why
fluorescence cuvettes, which are made of quartz (which does not absorb UV light), are
not frosted on any sides.
Chem 301
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
Page 6 of 15
Figure 6. General setup of
the fluorolog-3
spectrofluorimeter
(Fluorolog-3 v. 2.2 manual)
Tryptophan is the only amino acid that naturally fluoresces (i.e. it is a
“fluorophore”) and any protein that contains this residue will exhibit fluorescence
characteristics. The amount of fluorescence depends on the number of tryptophan
residues in the protein, the concentration of the protein, and the local environment of the
tryptophan fluorophore. If exposed to a polar solvent, tryptophan will exhibit a lower
quantum yield than if in a nonpolar environment. Many amino acid residues and/or
prosthetic groups, however, e.g. methionine and heme, are natural fluorescence
“quenchers” which act as energy sinks, taking the energy of the fluorophore before it
can emit at its normal intensity. Tryptophan residues are often found on the interior of a
folded protein and protein unfolding can therefore often be monitored using
fluorescence intensity, which may increase or decrease depending on the particular
protein. Tryptophan fluorescence has also been used to monitor protein-nucleic acid
interactions. A common way in which proteins interact with DNA or RNA involves van
der Waals and π electron interactions between trp side chains and the large planar
aromatic ring systems of the nucleotide bases. Once they come in contact, the trp
fluorescence increases due to the decreased local polarity of the trp fluorophore.
CIRCULAR DICHROISM SPECTROPOLARIMETRY
Circularly polarized light can have a right-handed rotation and a left-handed
rotation. Some substances absorb right- and left-handed circularly polarized light to
different extents; this process is called circular dichroism (CD) and CD spectroscopy or
CD spectropolarimetry quantitates the circular dichroism of a given sample. CD is most
useful in protein structure analysis when used to measure secondary structure content.
Alpha helices, beta sheets, and random coil conformations have characteristic CD
spectra. Quantitation of secondary structure content (e.g. percent alpha, beta, or coil) is
only considered reliable for alpha helical structure. CD is particularly useful when
measuring changes in the structure of the same protein as a function of solvent,
temperature, or single point mutations. Comparing different proteins in a quantitative
way is difficult using this method, but CD has benefits over other methods of secondary
structure determination in that it is fast, inexpensive instrumentation-wise, and requires
no special sample handling (most buffers have negligible CD interference, proteins can
be studied at low concentration and without isotopic enrichment or modifications).
Chem 301
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
Page 7 of 15
VIBRATIONAL SPECTROSCOPY
Molecules have vibrational energy levels in addition to the electronic energy
levels that you have most likely been exposed to at this point (the energy levels probed
by UV and visible spectroscopies. These energy levels, depicted below, have
differences that are smaller in energy (~100J/mol) and therefore probed by lower
energy, longer wavelength (~1 mm) electromagnetic radiation, typically infrared (IR)
light. By convention, wavenumbers (1/λ) are used in vibrational spectra rather than
wavelength, with typical units of cm-1 (“wavenumbers”).
Fig. 7 P.E. function for two
atoms as a function of
internuclear distance, R. In
reality, there are many more
energy levels than what is
shown here.
E
R
Proteins have a large number of vibrational modes that exhibit defined bands in Raman
and IR spectra; some of these are characteristic for specific secondary structures (see
Table below; taken from Gremlick & Yan. Infrared and Raman Spectroscopy of
Biological Molecules, chapter 11 (2001)). Molecules with permanent dipolemoments
typically have strong absorption in IR spectra. Biological molecules are often studied in
aqueous solution which has significant background absorption in IR spectra and
obscures the spectral signals of the molecule being studied (proteins, nucleic acids,
etc). For this reason, the vibrations in biological molecules are often studied using
Raman spectroscopy.
When matter absorbs light, there is a large amount of light that is not absorbed
but scattered. Most of the scattered light has the same wavelength of the incident
(incoming) light (“Rayleigh scattering”) but some is at a slightly higher or lower
energy/frequency than the incident light (“Raman scattering”). Raman spectroscopy is
relatively insensitive due to the fact that you are measuring the very small amount of
Raman scattering that occurs. Because of this, the incident light must be very intense
(visible lasers are commonly used today) and samples must be relatively concentrated
(protein sample concentrations required are ~25-100 mg/ml).
Table 1. Vibrational modes of peptide bond characteristic of given 2° structures.
Assigned vibrational mode
Random coil
α-helix
β-sheet
-1
-1
Amide I
1660-1645 cm
1680-1665 cm
1670-1660 cm-1
-1
-1
Amide III
1310-1260 cm
1240-1225 cm
1260-1240 cm-1
Skeletal C-C
950-885 cm-1
1010-1000 cm-1
960-950 cm-1
Chem 301
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
Page 8 of 15
Vibrational spectroscopy is one of the few spectroscopic methods available to
study solid phase samples. These methods are therefore prominent in the study of
fibrous (structural) proteins, such as collagen and keratin, as well as amyloidogenic
proteins implicated in amyloid diseases, such as Alzheimer’s, Huntington’s, Parkinson’s,
and transmissible spongiform encephalopathies, like Creutzfeldt-Jacob and mad cow
disease.
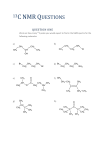
NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY
You have learned about Nuclear magnetic resonance (NMR) with regard to
organic molecule identification and chemical structure determination. But NMR is a
powerful tool for studying biological molecules including proteins, nucleic acids, lipids
and carbohydrates. The same principles apply but additional information can also be
garnered from NMR spectra of these more complex organic molecules. You learned in
general chemistry that electrons have “spin”, but many nuclei also have magnetic
moments that, when exposed to an external magnetic field, will align either with (lower
energy) or against (higher energy) the applied field. These two energy states are the
energy states that are “probed” by NMR. Nuclei that are “spin-1/2 nuclei” have NMR
signals; these include 1H, 13C, 15N, 19F, and 31P. 1H is the most naturally abundant
isotope of hydrogen, so any compound containing hydrogen atoms will have an NMR
signal. 19F NMR is a method for studying ligand-binding interactions and is also used in
imaging (MRI); 19F is 100% abundant but does not naturally occur in biological
molecules and its study therefore requires fluorination at specific sites on the molecule
under study. 31P naturally occurs in DNA and RNA but this particular nucleus has
problems in that its signal “relaxes” quickly and is therefore extremely broad and difficult
to observe or interpret. 13C and 15N are not the most abundant isotopes of carbon or
nitrogen, respectively, but their presence in biological molecules makes NMR study
feasible. They are routinely incorporated into proteins by growing bacteria that express
the protein-of-interest on media that is isotopically enriched in these isotopes (13Cglucose and 15NH4Cl are added as the sole C and N sources, respectively.
Proton chemical shifts for amino acid residues in proteins are sensitive to their
local electronic environment. Aliphatic protons in general have upfield shifted peaks
(lower ppm, more "shielded" because of a higher electron density), whereas aromatic
protons in general have downfield shifted peaks (higher ppm, "deshielded" due to the π
electon system influencing the protons on the sides of the ring plane). Nearby
substituents can influence the chemical shift through shielding or deshielding (removing
electron density). For example, serine has beta protons that are shifted downfield
relative to where they would be if this were an aliphatic chain; this deshielding occurs
because the hydroxyl oxygen is electronegative and withdraws electron density, shifting
the frequency to a lower field ("downfield"). Table 2 below is taken from Wishart et al
(1991) J. Mol. Biol., 222, 311-333 and includes the averaged proton chemical shifts
observed for protons in amino acid residues in proteins. Another NMR phenomenon is
the concept of scalar (or J) coupling. A H nucleus that has n hydrogens (equivalent to
each other but not equivalent to the H nucleus in question) attached to the same or an
Chem 301
Page 9 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
adjacent atom (usually C atoms), will have an NMR signal that is split into n+1 peaks.
You will observe this coupling in the 1D 1H spectra you will collect in lab but you will
NOT see this phenomenon in the multidimensional high resolution data you receive
from me. Multidimensional spectra often have the coupling removed (i.e. they are
"decoupled") by the experimental setup of the spectrometer.
Table 2: Averaged 1H chemical shift values for amino acids in all configurations (from Wisart, et al.)
Residue
Gly
Ala
Val
Ile
Leu
NH
8.33
8.12
8.26
8.27
8.25
α-H
4.14, 3.64
4.20
4.13
4.14
4.29
1.36
2.00
1.79
1.75, 1.50
Pro
Ser
Thr
8.32
8.27
4.41
4.52
4.50
2.21, 1.84
3.95, 3.79
4.15
Asp
8.33
4.62
2.88, 2.62
Glu
8.30
4.22
2.11, 2.00
γ-CH2 2.35, 2.28
Lys
8.19
4.23
1.87, 1.72
Arg
Asn
Gln
Met
Cys
Trp
Phe
8.22
8.38
8.23
8.21
8.35
8.22
8.36
4.24
4.68
4.31
4.32
4.74
4.63
4.58
1.86,
2.93,
2.14,
1.92,
3.16,
3.41,
3.10,
γ-CH2 1.42, 1.34
δ-CH2 1.63, 1.60 ε-CH2 2.95, 2.89 ε-NH3 7.51
γ-CH2 1.62, 1.52 δ-CH2 3.12, 3.08 NH 7.30, 6.62
γ-NH2 7.61, 6.91
γ-CH2 2.37, 2.25 δ-NH2 7.40, 6.71
γ-CH2 2.25, 2.00 ε-CH3 1.53
Tyr
8.36
4.62
3.08, 2.77
2,6
3.12, 2.86
2
His
8.26
4.57
Others
β-H
1.71
2.64
1.97
1.66
2.76
3.11
2.81
γ-CH3 0.91, 0.72
γ-CH2 1.22, 1.16 γ-CH3 0.77
γ-CH 1.54 δ-CH3 0.79, 0.67
γ-CH2 1.91, 1.77 δ-CH2 3.70, 3.46
γ-CH3 1.17
2
H 7.24 4H 7.65 5H 7.17 6H 7.24
H 7.30 3,5H 7.39 4H 7.34
2,6
H 7.15
7
H 7.50 NH 10.22
3,5
H 6.86
4
H 8.12 H 7.14
YOU WILL BE ASSIGNED TO EITHER GROUP A OR GROUP B. GROUP A WILL
CONDUCT PARTS I-III THIS WEEK AND PARTS IV-VI NEXT WEEK. GROUP B
WILL CONDUCT PARTS IV-VI THIS WEEK AND PARTS I-III NEXT WEEK.
PART I. UV-VIS SPECTROPHOTOMETRY
The purpose of this segment of the lab is to collect visible absorbance spectra
collected on fully oxidized and fully reduced cytochrome c and to use those spectra to
determine the extent of oxidation in an unknown sample of cytochrome c.
PROCEDURE:
1. You will be provided with the following solutions: 100 mM ascorbic acid,
100 mM ferricyanide, and 0.5 mg/mL cytochrome c in potassium
phosphate buffer, pH 7.0.
2. Prepare the following solutions, each with a total volume of 1.0 mL and
each with identical phosphate buffer concentrations.
Chem 301
Page 10 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
a. 0.45 mg/ml cytochrome c in 10 mM ascorbic acid.
b. 0.45 mg/ml cytochrome c in 10 mM ferricyanide.
3. Collect absorbance scans for these solutions along with the unknown
solution provided for the entire visible region of the electromagnetic
spectrum.
DATA ANALYSIS/INTERPRETATION:
Using the scans collected, estimate the fraction of oxidized cytochrome c
in the unknown provided. You can assume that the [cytochrome c] in the
unknown sample is 0.45 mg/ml. Provide all three spectra in your report along
with your rationale behind your estimation.
PART II. FLUORESCENCE SPECTROSCOPY
The purpose of this segment of the lab is to monitor changes in structure of
bovine serum albumin (BSA) and cytochrome c that occur with increasing chemical
denaturant concentration using the intrinsic fluorescence of the protein. You will use the
chaotropic agents urea and guanidinium chloride to unfold the proteins.
PROCEDURE:
1.
You will be provided with a 20.0 mg/mL solution of BSA or a 10.0 mg/mL solution
of cytochrome c. Work with the group across from you and make one set of solutions
(either BSA or cyt c); the other two groups at your bench will make the other set. Using
this stock solution, phosphate buffer, and the chemical denaturant provided (8.0 M urea
or 6.0 M guanidine hydrochloride), make the following solutions in test tubes.
Sample
Vol. protein stock soln (mL)
Vol. buffer (mL)
Vol. denaturant (mL)
1
0.10
3.70
0.20
2
0.10
3.60
0.30
3
0.10
3.30
0.60
4
0.10
2.80
1.10
5
0.10
2.30
1.60
6
0.10
2.10
1.80
7
0.10
1.90
2.00
8
0.10
1.80
2.10
9
0.10
1.30
2.60
10
0.10
0.80
3.10
11
0.10
0.30
3.60
12
0.10
0
3.90
2.
Mix each solution thoroughly and collect a fluorescence emission
spectrum for each sample with the following parameters:
Collect à Experiment and then fill in the following parameters:
Chem 301
Page 11 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
λexcitation = 295 nm, λemission = 300 nm – 500 nm
Increment = 1 nm, Integration time = 0.5 s
Bandpass slitwidths (click on "slits") = 5 nm for both excitation and emission
Signals à clear all and double click “S” and then "ok"
Datafile: (give a unique name so that you can retrieve it later to export it). Save
these files to C: in datamax/data/chem301/2009
3.
When the instrument is free you should export your data as an ASCII file
and save to C: in datamax/data/chem301/2009. I will post exported data on
Bb; you will need both your data and the data of the other group.
DATA ANALYSIS/INTERPRETATION:
1.
You should display the emission scans for all the samples in your set (so you will
have one graph, and it will have all 12 scans on it). If the figure looks better with only
some of the scans you can remove select scans but label what remains.
2.
Plot the maximum fluorescence intensity as a function of [denaturant] for
both the transitions (you will need data from the other group to complete this
section; they can give you their excel data from part 1 or part 2, or the raw
data. You should negotiate a fair trade with them.). Discuss how the
fluorescence changes as the protein unfolds and what that might mean about
the change in trp’s local environment. Do both proteins have the same emission
(do both increase or do both decrease) upon unfolding? Explain what might be
happening for each. Do the unfolding transitions appear to be cooperative? Why
or why not? Which denaturant is more effective and why?
PART III. CD SPECTROPOLARIMETRY
The purpose of this segment of the lab is to identify general patterns in CD
spectra with respect to secondary structure content in proteins.
PROCEDURE:
(You can do this portion of the lab anytime à you don't need to do this today
during lab time but if there is time, I would suggest not procrastinating.)
1.
You will be provided with CD spectra (these are on the Bb site) that have already
been collected on an Aviv Model 202 CD Spectrometer. These spectra correspond to
human serum albumin (PDB ID 1BM0), Nup153 (PDB ID 2EBV), and chitinase (2CZN).
2.
From the PDB website, look at the structure of each protein. Based solely
on visual inspection, estimate if the given protein is all alpha, all beta, mixed
alpha/beta, or mostly unstructured.
3.
Also on the PDB website, obtain a Ramachandran plot for each protein by
going to Geometry tab (blue tabs at top: “Summary, derived data, sequence…”
click geometry tab then click on “Click here to download molprobity
Ramachandran plot.” This generates a pdf file that you can print and/or save
Chem 301
Page 12 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
and print later for your lab report. Use this plot to corroborate your conclusion
in step 2 about the secondary structure identity.
DATA ANALYSIS/INTERPRETATION:
1.
Assuming that these proteins are representative of their general “family”
of secondary structure, specifically discuss the CD spectra that alpha helical,
beta sheet, and random coil proteins exhibit. Provide λ values and discuss if
these are maxima or minima. Also include the relative intensity (molar ellipticity)
of each (i.e.does the λmax of alpha have a similar intensity to the λmax of beta?).
2.
Include all CD spectra on the same graph in your lab report (that will most likely
require different line types, e.g. dashed, solid, dotted, without points for clarity purposes)
and label each one clearly with which type of secondary structure it represents. Include
both ribbon diagrams of the three proteins and Ramachandran plots for each.
PART IV. RAMAN SPECTROSCOPIC ANALYSIS OF PROTEIN SECONDARY
STRUCTURES
In the lab are samples of silk and wool; you should collaborate with the group
across from you and share spectral data in order to compare the two samples (i.e. if
they do silk then you should do wool). You will need both spectra for analysis.
PROCEDURE:
Setting up the spectrometer and microscope:
1. You will be using the Labspec 5 program. Open this and allow it time to
load before proceeding. Once it is loaded, make sure that the laser is off by
making sure that in the “Aramis” box on the far right, the laser icon looks like
this.
2. Click the “Setup” button in the “Aramis” box directly below the laser switch.
Set the “Measure Location” to “micro,” and under “Visualization” make sure
that it is set to “video on” and “trino off.” Press OK to go back to the main
screen. In the toolbar at the top, click “video” and then click “camera 1.”
3. Under “laser” make sure that it is set to “Diode IR.” Make sure that under
“filter” no filter is selected (choose the triple dash in the drop down menu). Set
the grating ( ) to 300 then set the spectrometer to 1600 cm-1 and press enter
to verify the change. Under “Acquisition” set the “exposure time” to 5 seconds
and the “accumulation number” to 1.
4. Open the microscope chamber (the big double doors) on the Raman
spectrometer. Place your sample on a slide and place that on the stage under
the microscope.
Chem 301
Page 13 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
5. To get a good spectra, you will need to move your sample so that the laser is
focused on a fiber. To do this, first press the camera ( ) button in the top
toolbar. This opens a new window allowing you to view your sample under the
microscope. You won’t see anything unless the lamp is on and the microscope
is in focus. Turn the lamp on to around 6-8.
6. Use the joystick to move the stage until the image on the microscope is very
clear. Center the crosshairs on a very clear portion of a fiber.
7. Once you have set all the parameters and have focused the microscope on a
fiber, you are ready to take your sample. First, close the doors to the
microscope chamber and turn off the lamp. You do not need to turn on the
laser yourself, it will automatically turn on when you take the spectra and turn
off when it is done.
8. Take the spectra by clicking
in the top toolbar or by pressing Ctrl+A. It
will gather the spectra for two and half minutes. After it is done collecting the
spectra save the file as a txt file so that it can be later opened as an excel file.
9.
Change the ET to 30 seconds and collect another spectrum.
10.
Change the AN to 5 and collect a final spectrum.
DATA ANALYSIS/INTERPRETATION:
1.
For the spectra that your group collected, show the three spectra and comment
on the effect of ET and AN.
2.
Use your final spectrum (AN=5 and ET=30) and identify any relevant Raman
bands and assign them to a particular vibration as provided in the introduction.
3.
Overlay the final spectra (AN=5 and ET=30) spectra of both samples and discuss
the likely secondary structures present in these proteins. Use the literature (your
textbook, the web, other books) to confirm (or not) your analysis.
PART V. 1H NMR OF AMINO ACIDS IN AQUEOUS SOLUTION
You will collect one dimensional 1H NMR spectra of an amino acid of your choice.
Each member of your group will prepare a sample as described below.
PROCEDURE:
1.
There are amino acid samples on the cart in the lab. Choose your favorite amino
acid from the ones that are available and bring it back to your bench.
2.
Prepare the samples below.
(a)
0.7 mL of 0.3 M amino acid in water (the molar mass is probably on the
bottle; if not then the chemical formula is and there’s a periodic table on the
wall).
(b)
0.7 mL of 0.3 M amino acid (the same one) in deuterium oxide (D2O).
3.
Collect an NMR spectrum of each sample your group has made.
DATA ANALYSIS/INTERPRETATION:
Chem 301
Page 14 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
1.
Discuss the NMR spectra of your sample in the two solvents used; discuss (1)
the difference in the size of the solvent peak in each spectrum and why they are not the
same, (2) the relative concentration of 1H in your sample vs. the concentration of 1H in
water, and (3) why D2O is indispensible for this experiment.
2.
Answer the questions provided on Bb regarding the spectra.
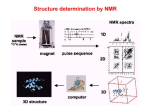
PART VI. ANALYSIS OF NMR SPECTRA OF PEPTIDES AND PROTEINS: MULTIDIMENSIONAL NMR
NMR spectra of peptides and proteins requires multiple dimensions due to the
problem of “spectral overlap”. You will be given NMR spectra of various proteins and
peptides in order to begin to understand how NMR is used to characterize larger
molecules.
DATA ANALYSIS/INTERPRETATION:
(You can do this portion of the lab anytime à you don't need to do this today
during lab time but if there is time, I would suggest not procrastinating.)
1.
You will be provided (these are on Bb) with NMR spectra that have been
collected on a 500 MHz or a 600 MHz NMR spectrometer (these are considered “high
resolution” NMR spectrometers; for comparison, ours is 60 MHz. Organic molecules are
typically studied on 60 MHz – 300 MHz spectrometers. Proteins and peptides typically
require a magnetic field strength that gives a proton frequency of at least 400-500 MHz.
The highest field spectrometers in routine use today are ~900-950 MHz.)
2.
Look at the one dimensional spectra of the peptide hormone oxytocin
(CYIQNCPLG). Count as many peaks as you can in the amide/aromatic region,
downfield (left) of the water peak. Can you find all amide proton signals that are
in the peptide? One residue will have side chain peaks down here; which one is
that?
3.
Notice the larger degree of overlap in this spectrum relative to the amino
acids. Look at the corresponding 2D spectra of this peptide. On 2D (and 3 and
4D) spectra, each “dot” is a peak. The 2D spectrum you have of this peptide is
homonuclear (both axes or “dimensions” are chemical shifts of the same
identity nucleus and that typically means both are 1H) or a 1H-1H spectrum. You
know it is homonuclear and that both axes are the same b/c there are “diagonal
peaks” that together form a diagonal line on the spectrum.
4.
Look at the protein 2D NMR spectra provided. There are three spectra; all
1
are H-15N heteronuclear 2D spectra collected on proteins in aqueous solution
(mostly H2O; otherwise you would not see the amide peaks). You can see that
the axes (“dimensions”) are different in the size of the chemical shift values.
Remember that 13C chemical shifts are very different than proton chemical
shifts in value (about 10 times larger or so for aliphatic C, though other
carbons, e.g. alkenes, alkynes can be higher) and amide 15N chemical shifts are
even larger than that. Note that there are no diagonal peaks. Each peak on this
particular spectrum corresponds to an amide proton and its attached nitrogen.
Chem 301
Page 15 of 15
Biological Chemistry I Laboratory
Lab 7: Protein Spectroscopy
Use the spectrum to estimate the number of residues in this protein. Which
residues will not have amide protons?
5.
The first two heteronuclear spectra are 1H-15N spectra collected on the
same protein in pH 7 buffer and in buffer containing 8M urea, respectively. What
do you notice about the chemical shifts in the presence of urea? What is urea
doing to the protein? Which chemical shift is more sensitive to its environment,
the amide proton or the amide nitrogen?
7.
The last spectrum is an overlay of a protein in the absence (dark peaks)
of a ligand and in the presence of the ligand (light colored peaks which are only
evident when not on top of a dark peak). 1H-15N spectra are commonly collected
to determine if ligand-binding is occurring and which residues are likely involved
in the binding interaction. Chemical shift is sensitive enough to the local
environment that ligand-binding will cause a change in signal for those residues
that are close to the ligand-binding site. Can you identify any peaks that
correspond to residues that are likely a part of the ligand-binding site? What are
the chemical shifts (provide 1H &15N shifts in ppm).