* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Test Info Sheet
Survey
Document related concepts
Transcript
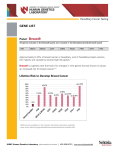
GeneDx 207 Perry Parkway Gaithersburg, MD 20877 Phone: 888-729-1206 Fax: 301-710-6594 E-mail: [email protected] www.genedx.com/oncology Test Information Sheet OncoGeneDx: Breast High/Moderate Risk panel Gene List for the OncoGeneDx Breast High/Moderate Risk panel: ATM BRCA1 BRCA2 CDH1 CHEK2 PALB2 PTEN TP53 Clinical Features: In the general population, approximately 1 in 8 women (12%) will develop breast cancer in their lifetime (SEER). Most cases of breast cancer develop sporadically with no family history of the cancer; however, 5‐10% of cases are thought to be due to a hereditary predisposition. The features suggestive of a hereditary cancer predisposition include: young age at diagnosis (before age 50), multiple primary cancers in a single individual, diagnosis of a cancer type that is not common in the general population (such as ovarian cancer, male breast cancer, or pancreatic cancer), and several relatives affected with related cancers spanning multiple generations. The OncoGeneDx Breast Cancer High/Moderate Risk panel includes genes associated with an increased lifetime risk for breast cancer of 20% or higher. The genes included are: ATM, BRCA1, BRCA2, CDH1, CHEK2, PALB2, PTEN and TP53. It is estimated that 20‐25% of familial breast cancer risk can be attributed to pathogenic variants in the BRCA1 or BRCA2 genes (Easton 1999, Pharoah 2002, van der Groep 2011). The contribution of pathogenic variants in the ATM, CDH1, CHEK2, PALB2, PTEN and TP53 genes to familial breast cancer risk overall is less well‐characterized but is considerably lower than the contribution of BRCA1 and BRCA2 pathogenic variants. ATM: Women with a pathogenic variant in ATM have approximately a two‐fold increase risk for breast cancer (RR = 2.2‐2.4) (Thompson 2005, Renwick 2006, Tavtigian 2009). Thompson et al. (2005) studied 1160 ATM pathogenic variant carriers and concluded that female heterozygous ATM pathogenic variant carriers who are less than 50 years of age had a significantly increased risk for breast cancer (RR = 4.9) compared to women over 50 years of age where a statistically significant risk could not be identified. This same study suggests an increased risk for colon cancer, but the confidence intervals are wide (Thompson 2005). Roberts et al. (2012) reported an association with pancreatic cancer showing that 2.4% of familial pancreatic cancer patients were found to carry a pathogenic variant in ATM, and 4.6% of families with 3 or more cases of pancreatic cancer carried a pathogenic variant in ATM. Of note, certain missense pathogenic variants in the ATM gene may confer a higher breast cancer risk (Tavtigian 2009). BRCA1 and BRCA2 (Hereditary Breast and Ovarian Cancer syndrome): Women with pathogenic variants in BRCA1 or BRCA2 have a 41‐87% lifetime risk to develop breast cancer and an up to 63% risk for contralateral breast cancer (Antoniou 2003, Chen 2007, Claus 1996, Ford 1998, King 2003, Graeser 2009, Risch 2006). Studies have shown that the lifetime risk to develop ovarian cancer is between 24‐54% for carriers of pathogenic variants in BRCA1 and 11‐ 27% for BRCA2 pathogenic variant carriers (Antoniou 2003, Chen 2007, Ford 1998, King 2003, Risch 2006). Other cancers associated with pathogenic variants in BRCA1 and BRCA2 in women include fallopian tube carcinoma, primary peritoneal carcinoma, and uterine serous carcinoma (Levine 2003, Biron‐Shental 2006, Pennington 2013). The lifetime risk for breast cancer in male carriers of a BRCA1/2 pathogenic variant is approximately 7% with a BRCA2 pathogenic variant and slightly increased with a BRCA1 pathogenic variant (Liede 2004, Tai 2007). Other malignancies reported in families with pathogenic variants in BRCA1 or BRCA2 include prostate cancer in men, as well as pancreatic cancer and melanoma in both men and women. Information Sheet on Breast High/Moderate Risk panel Page 1 of 7 © GeneDx Revision Date: 07/2016 CDH1 (Hereditary Diffuse Gastric Cancer syndrome): Women with a pathogenic variant in CDH1 have a 39‐52% lifetime risk for lobular breast cancer. The lifetime risk of diffuse gastric cancer has been estimated to be 40‐67% for men and 63‐83% for women (Kaurah 2007, Pharoah 2001). Diffuse gastric cancer generally occurs before age 50 in CDH1 pathogenic variant carriers, and even cases under the age of 18 have been reported in families with hereditary diffuse gastric cancer (Guilford 1998). Signet ring cell cancer of the colon has also been reported in individuals with a pathogenic variant in CDH1 (Brooks‐Wilson 2004). More recently, CDH1 pathogenic variants have also been identified in families with lobular carcinoma in situ or invasive lobular carcinoma of the breast (LCIS/ILC) but no history of gastric cancer, suggesting the spectrum may include breast‐only families (Petridis 2014, Schrader 2011). CHEK2: Pathogenic variants in CHEK2 are known to confer an increased risk of breast cancer and have been suggested to also increase the risk of colon and others cancers. While studies have shown that the risk for breast cancer may vary depending on the type of pathogenic variant and family history, in general those found to harbor a pathogenic variant in CHEK2 are estimated to have a 2‐fold increased risk. (Cybulski 2004, The CHEK2 Breast Cancer Consortium 2004, Liu 2012, Han 2013). Additionally, pathogenic variants in CHEK2 have been reported to be associated with an increased risk for colon, prostate, endometrial, and ovarian cancer; however, these risks are not well defined (Seppala 2003, Dong 2003, Cybulski 2004, Kilpivaara 2006, Einarsdóttir 2007, Suchy 2010, Walsh 2011, Liu 2012, Han 2013, Pennington 2013). PTEN (PTEN Hamartoma Tumor syndrome): Cowden Syndrome (CS) and Bannayan‐Riley‐Ruvalcaba (BRRS) are two conditions belonging to the spectrum of PTEN hamartoma tumor syndrome (PHTS) and are associated with an increased risk of developing cancer. Cowden Syndrome is specifically is associated with an increased risk for malignant tumors of the breast, thyroid, and endometrium. The lifetime risk for these cancers are estimated to be 25‐45% for female breast cancer, 5‐10% risk for endometrial cancer, and 10% risk for non‐medullary thyroid cancer in both men and women and men (Hobert 2009). More recent reports have suggested that cancer risks for individuals with Cowden syndrome may actually be higher than previously reported; the most striking of these risks being the cumulative risk for female breast cancer of 77 to 85% (Bubien 2014, Tan 2012, Riegert‐Johnson 2010). Additional clinical features in CS include macrocephaly, trichilemmomas, papillomatous papules, glycogenic acanthosis and Lhermitte‐Duclos disease (dysplastic gangliocytoma of the cerebellum which is considered to be pathognomonic). CS may be also associated with benign conditions including thyroid goiters, benign breast disease, mixed gastrointestinal polyposis and uterine fibroids. BRRS is associated with macrocephaly, intestinal hamartomas, pigmented macules of the glans penis, and can be associated with developmental delay or autism. Vascular abnormalities, such as hemangiomas and arterio‐venous malformations, have also been reported in individuals with pathogenic variants in PTEN. PALB2: Pathogenic variants in the PALB2 gene have been estimated to confer a 2 to 3‐fold increased risk of breast cancer over the general population resulting in a lifetime risk of approximately 25% to 40% (Erkko 2008, Rahman 2007). More recent data has suggested a lifetime risk (up to age 70) ranging from 33% to 58% depending on the individual’s family history of breast cancer (Antoniou 2014). Women with a pathogenic variant in PALB2 who have a family history of early‐onset breast cancer may have a lifetime risk up to 58% (Byrnes 2008, Antoniou 2014). Casadei et al. (2011) found that PALB2 pathogenic variant carriers are 6‐times more likely to have a family history of pancreatic cancer, 1.3‐ times more likely to have a family history of ovarian cancer and 4 times more likely to have a family history of male breast cancer. Although the association of pathogenic variants in PALB2 and pancreatic cancer has been established, the exact risks are not yet well‐understood (Jones 2009, Slater 2010). TP53 (Li‐Fraumeni syndrome): The following core cancers account for 70‐77% of LFS‐associated tumors (in order of frequency): breast cancer, soft tissue sarcoma, brain tumors, osteosarcoma, and adrenocortical carcinoma (Gonzalez 2009, Olivier 2003, Ruijs 2010). Other types of cancer that may be associated with LFS include ovarian, gastrointestinal, pancreatic, genitourinary, skin, thyroid and lung cancers as well as leukemia, lymphoma, and neuroblastomas. Age‐related and sex‐specific cancer risks have been reported. Individuals with LFS who have been diagnosed with cancer have up to a 57% risk of developing a second primary cancer within 30 years of the first Information Sheet on Breast High/Moderate Risk panel Page 2 of 7 © GeneDx Revision Date: 07/2016 diagnosis and up to a 38% risk of a third primary diagnosis (Hisada 1998). Several studies have demonstrated that subsequent tumors often develop in the radiation field of the previously treated cancer (Chompret 2000, Hisada 1998). Specific cancer screening and prevention recommendations for individuals with pathogenic variants in all of the genes on this panel can be found in the NCCN Clinical Practice Guidelines in Oncology Genetic/Familial High‐Risk Assessment: Breast and Ovarian. In addition, cancer screening and prevention recommendations for individuals with pathogenic variants in CDH1 can be found in the NCCN Clinical Practice Guidelines in Oncology: Gastric Cancer; additionally, the International Gastric Cancer Linkage Consortium (IGCLC) has published updated consensus guidelines for the management of individuals with hereditary diffuse gastric cancer (Fitzgerald 2010). The available medical management guidelines pertinent to each gene are listed in the attached table. Inheritance Pattern: Hereditary Breast and Ovarian Cancer syndrome (BRCA1 and BRCA2), hereditary diffuse gastric cancer syndrome (CDH1), Cowden syndrome/PTEN hamartoma tumor syndrome (PTEN), Li‐Fraumeni syndrome (TP53), and pathogenic variants in the ATM, CHEK2 and PALB2 genes are inherited in an autosomal dominant manner. While de novo pathogenic variants in the BRCA1, BRCA2, and CDH1 genes are uncommon, PTEN and TP53 pathogenic variants have been shown to occur de novo in a subset of cases. At this time, the de novo pathogenic variant rate of ATM, CHEK2 and PALB2 is unknown. Biallelic pathogenic variants in the BRCA2 and the PALB2 genes (i.e. one pathogenic variant in each copy of the gene) are associated with an extremely rare autosomal recessive syndrome called Fanconi anemia. This condition is characterized by an increased risk for malignancy in children, including leukemia and certain solid tumors, bone marrow failure, and distinctive clinical features including radial abnormalities. Additionally, biallelic pathogenic variants in the ATM gene are associated with another rate autosomal recessive syndrome called ataxia telangiectasia (AT). This multisystem disorder is characterized by progressive neurodegeneration, telangiectasias, immunodeficiency, and increased cancer risk. Individuals found to carry a pathogenic variant in BRCA2, PALB2 or ATM who are considering pregnancy should be offered reproductive counseling. Reason for referral: 1) Identify the genetic basis of breast cancer for individuals who have features and/or a family history consistent with one of the hereditary cancer syndromes described above. 2) To possibly help determine appropriate clinical management recommendations based on a molecular diagnosis. 3) Identify family members at‐risk to develop features associated with a specific hereditary cancer syndrome. Methods: Genomic DNA from the submitted specimen was enriched for the complete coding region and splice site junctions of the genes on the panel using a proprietary targeted capture system developed by GeneDx. For PTEN, nucleotides c.‐ 700 through c.‐1300 in the promoter region are also sequenced. The products were sequenced on either an Illumina MiSeq or HiSeq instrument with 2x150 or 2x100 paired‐end reads, respectively. The sequence was aligned to reference sequences based on human genome build GRCh37/UCSC hg19. Capillary sequencing was used to confirm all variants with clinical or uncertain significance and to analyze regions with inadequate coverage by Next Generation sequencing. If present, apparently homozygous variants were confirmed using alternate primer pairs to significantly reduce the possibility of allele drop‐out. Concurrent deletion/duplication testing was performed for all of the genes on the panel using either exon‐level array CGH or MLPA. Confirmation of copy number changes was performed by MLPA, qPCR, or repeat aCGH analysis. Data analysis was performed using gene‐specific filtering. The array was designed to detect most single‐exon deletions and duplications. All sequence alterations are described according to the Human Genome Variation Society (HGVS) nomenclature guidelines. Benign and likely benign variants, if present, are not reported but are available upon request. The genes evaluated by this test are listed on the first page of the report. Information Sheet on Breast High/Moderate Risk panel Page 3 of 7 © GeneDx Revision Date: 07/2016 Test Performance: DNA sequencing will detect nucleotide substitutions and small insertions and deletions, while array CGH will detect exon‐level deletions and duplications. These methods are expected to be greater than 99% sensitive in detecting variants identifiable by sequencing or array CGH. The likelihood of a false positive result is expected to be <1%. Technical Limitations: Neither sequencing nor exon‐level aCGH can reliably detect mosaicism, and cannot detect chromosomal aberrations. Deletions involving more than 20bp and insertions involving more than 10bp are not reliably detected by the sequencing methodology, and deletions or duplications of less than 250bp are not reliably detected by array CGH. Regions of certain genes have inherent sequence properties that yield suboptimal data, potentially impairing accuracy of the results. For instance, sequence and deletion/duplication analysis of CHEK2, among others, is complicated by the presence of pseudogenes or homologous sequences that involve multiple exons of this gene. In the absence of mRNA/cDNA studies, we cannot completely exclude the possibility of undetectable clinically significant variants in certain regions of these genes. False negatives may also occur in the setting of bone marrow transplantation, recent blood transfusion, or suboptimal DNA quality. In individuals with active leukemia or lymphoma or with known chronic myeloid or lymphoid neoplasms (such as low grade MDS, CML, ET, P. vera, PMF, CLL), there is a possibility that testing of specimens containing leukocytes may detect an acquired somatic variant, resulting in a false positive result. In this situation, please contact one of our genetic counselors to discuss the utility of submitting an alternate specimen. Additionally, rare false negatives may occur when testing for a specific variant identified at a laboratory other than GeneDx if a positive control is not provided. Based on the specific array design and technology used, the reported coordinates of duplications and deletions at the exon or gene level can slightly differ among family members tested but, in general, relatives are expected to have the same copy number variant. The ability to detect genetic variants and naming conventions can differ among laboratories. Reporting of Results: Results will be interpreted and reported following recommendations of the American College of Medical Genetics as a guideline (www.acmg.net). Variations detected by sequencing or deletion/duplication analysis will be analyzed and classified into the following categories based on current scientific knowledge. Our analysis includes a comprehensive assessment of the variation on a molecular and clinical level in order to determine its clinical significance and classification. Trained PhD analysts perform a detailed review of the variation on the molecular level, including exhaustive searches of gene– and locus‐specific databases and the Human Gene Mutation Database (HGMD), and genetic counselors and clinical molecular geneticists carefully review literature reports. Pathogenic Variant Examples of variations that may be reported as pathogenic include frameshift variants and nonsense variants that are predicted to result in premature protein truncation or mRNA decay, canonical splice site variants, and previously reported missense variants that are recognized as disease‐causing. Likely Pathogenic Variant Variations for which there is significant, but not conclusive, evidence supporting pathogenicity will be classified as Likely Pathogenic. Variant of Uncertain Significance Variations for which there is not sufficient evidence for classification will be classified as Variant of Uncertain Significance. Negative No variation of clinical or uncertain significance was detected. Any variation detected and classified as a likely benign or benign variant based on population data, review of the literature, Human Gene Mutation Database (HGMD), and appropriate locus‐specific databases will not be reported. Information Sheet on Breast High/Moderate Risk panel Page 4 of 7 © GeneDx Revision Date: 07/2016 Specimen Requirements and Shipping/Handling: Blood: Two EDTA (lavender top) tubes containing 4 ml each whole sterile blood Oral Rinse: Saliva collected in at least 30mL of mouthwash using our GeneDx collection kit Extracted DNA: >20 µg Buccal Swab: For family member testing only (excluding deletion/duplication family testing) Test Codes and Turnaround Times – Please contact us for price information: Test Code Description Turnaround Time J055 Breast High/Moderate Risk panel 2 weeks References Antoniou A et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet. 2003 May;72(5):1117‐30. (PMID 12677558) Antoniou A et al. Breast‐cancer risk in families with mutations in PALB2. N Engl J Med. 2014 Aug 7;371(6):497‐506. (PMID 25099575) Biron‐Shental T et al. High incidence of BRCA1‐2 germline mutations, previous breast cancer and familial cancer history in Jewish patients with uterine serous papillary carcinoma. Eur J Surg Oncol. 2006 Dec;32(10):1097‐ 100. (PMID 16650962) Brooks‐Wilson AR. Germline E‐cadherin mutations in hereditary diffuse gastric cancer: assessment of 42 new families and review of genetic screening criteria. J Med Genet. 2004 Jul;41(7):508‐17. (PMID 15235021) Bubien V et al. High cumulative risks of cancer in patients with PTEN hamartoma tumour syndrome. J Med Genet. 2013 Apr;50(4):255‐63. (PMID 23335809) Byrnes GB et al. Are the so‐called low penetrance breast cancer genes, ATM, BRIP1, PALB2, and CHEK2, high risk for women with strong family histories? Breast Cancer Res. 2008;10(3):208. (PMID 18557994) Canto MI et al. International Cancer of the Pancreas Screening (CAPS) Consortium summit on the management of patients with increased risk for familial pancreatic cancer. Gut. 2013 Mar;62(3):339‐47. (PMID 23135763) Casadei S et al. Contribution of inherited mutations in the BRCA2‐interacting protein PALB2 to familial breast cancer. Cancer Res. 2011 Mar 15;71(6):2222‐9. (PMID 21285249) Chen S and Parmigiani G. Meta‐anlaysis of BRCA1 and BRCA2 penetrance. J Clin Oncol. 2007 Apr;25(11):1329‐33. (PMID 17416853) Chompret A et al. P53 germline mutations in childhood cancers and cancer risk for carrier individuals. Br J Cancer. 2000 Jun;82(12):1932‐7. (PMID 10864200) Claus EB et al. The genetic attributable risk of breast and ovarian cancer. Cancer. 1996 Jun 1;77(11):2318‐24. (PMID: 8635102) Cybulski C et al. CHEK2 is a multiorgan cancer susceptibility gene. Am J Hum Genet. 2004 Dec;75(6):1131‐5. (PMID 15492928) Dong X et al. Mutations in CHEK2 associated with prostate cancer risk. Am J Hum Genet. 2003 Feb;72(2):270‐80. (PMID 12533788) Easton DF. How many more breast cancer predisposition genes are there? Breast Can Res. 1999 Aug;1(1):14‐17. (PMID 11250676) Einarsdóttir K et al. Effect of ATM, CHEK2 and ERBB2 TAGSNPs and haplotypes on endometrial cancer risk. Hum Mol Genet. 2007 Jan 15;16(2):154‐64. (PMID 17164260) Errko H et al. A recurrent mutation in PALB2 in Finnish cancer families. Nature. 2007 Mar 15;446(7133):316‐9. (PMID 17287723) Fitzgerald RC et al. Hereditary diffuse gastric cancer: updated consensus guidelines for clinical management and directions for future research. J Med Genet. 2010 Jul;47(7):436‐44. (PMID 20591882) Ford D et al. Genetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. The Breast Cancer Linkage Consortium. Am J Hum Genet. 1998 Mar;62(3):676‐89. (PMID 9497246) Gonzalez PD et al. Beyond Li Fraumeni Syndrome: clinical characteristics of families with p53 germline mutations. J Clin Oncol. 2009 Mar 10;27(8):1250‐6. (PMID 19204208) Guilford P. E‐cadherin germline mutations in familial gastric cancer. Nature. 1998 Mar 26;392(6674):402‐5. (PMID 9537325) Graeser MK et al. Contralateral Breast Cancer Risk in BRCA1 and BRCA2 Mutation Carriers. J Clin Oncol. 2009 Dec 10;27(35): 5887‐92. (PMID 19858402) Han FF et al. The effect of CHEK2 variant I157T on cancer susceptibility: evidence from a meta‐analysis. DNA Cell Biol. 2013 Jun;32(6):329‐35. (PMID 23713947) Hisada M et al. Multiple primary cancers in families with Li‐Fraumeni syndrome. J Natl Cancer Inst. 1998 Apr 15;90(8):606‐11. (PMID 9554443) Hobert JA and Eng C. PTEN hamartoma tumor syndrome: An overview. Genet Med 2009:11(10):687– 694. (PMID 19668082) Jones S et al. Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science. 2009 Apr 10;324(5924):217. (PMID 19264984) Kaurah P. Founder and recurrent CDH1 mutations in families with hereditary diffuse gastric cancer. JAMA. 2007 Jun 6;297(21):2360‐72. (PMID 17545690) Kilpivaara O et al. CHEK2 I157T associates with familial and sporadic colorectal cancer. J Med Genet. 2006 Jul;43(7):e34. (PMID 16816021) King MC et al. Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science. 2003 Oct;302(5645):643‐6. (PMID 14576434) Leide A et al. Cancer Risks for Male Carriers of Germline Mutations in BRCA1 or BRCA2: A Review of the Literature. J Clin Oncol. 2004 Feb 15;22(4):735‐42. (PMID 14966099) Levine DA et al. Fallopian Tube and Primary Peritoneal Carcinomas Associated With BRCA Mutations. J Clin Oncol. 2003 Nov 15;21(22):4222‐7. (PMID 14615451) Liu C et al. The CHEK2 I157T variant and colorectal cancer susceptibility: a systematic review and meta‐analysis. Asian Pac J Cancer Prev. 2012;13(5):2051‐5. (PMID 22901170) NCCN Guidelines. Gastric Cancer. (URL: http://www.nccn.org/professionals/physician_gls/f_guidelines.asp) [February 2016 accessed]. NCCN Guidelines. Genetic/Familial High‐Risk Assessment: Breast and Ovarian. (URL: http://www.nccn.org/professionals/physician_gls/f_guidelines.asp) [February 2016 accessed]. Information Sheet on Breast High/Moderate Risk panel Page 5 of 7 © GeneDx Revision Date: 07/2016 NCCN Guidelines. Genetic/Familial High‐Risk Assessment: Colorectal. (URL: http://www.nccn.org/professionals/physician_gls/f_guidelines.asp) [February 2016 accessed]. Olivier M et al. Li‐Fraumeni and related syndromes: correlation between tumor type, family structure, and TP53 genotype. Cancer Res. 2003 Oct 15;63(20):6643‐50. (PMID 14583457) Pennington KP et al. BRCA1, TP53, and CHEK2 germline mutations in uterine serous carcinoma. Cancer. 2013 Jan;119(2):332‐8. (PMID 22811390) Petridis C et al. Germline CDH1 mutations in bilateral lobular carcinoma in situ. Br J Cancer. 2014 Feb 18;110(4):1053‐7. (PMID 24366306) Pharoah PD et al. Incidence of gastric cancer and breast cancer in CDH1 (E‐cadherin) mutation carriers from hereditary diffuse gastric cancer families. Gastroenterology. 2001 Dec;121(6):1348‐53. (PMID 11729114) Pharoah PD et al. Polygenic susceptibility to breast cancer and implications for prevention. Nat Genet. 2002 May;31(1):33‐6. (PMID 11984562) Rahman N et al. PALB2, which encodes a BRCA2‐interacting protein, is a breast cancer susceptibility gene. Nat Genet. 2007 Feb;39(2):165‐7. (PMID 17200668) Renwick A et al. ATM mutations that cause ataxia‐telangiectasia are breast cancer susceptibility alleles. Nat Genet. 2006 Aug; 38(8):873‐5. (PMID 16832357) Risch HA et al. Population BRCA1 and BRCA2 mutation frequencies and cancer penetrances: a kin‐cohort study in Ontario. J Natl Cncer Inst. 2006 Dec;98(23):1694‐706. (PMID 17148771) Roberts NJ et al. ATM Mutations in Patients with Hereditary Pancreatic Cancer. Cancer Discov. 2012 Jan;2(1):41‐6. (PMID 22585167) Ruijs MWG et al. TP53 germline mutation testing in 180 families suspected of LieFraumeni syndrome: mutation detection rate and relative frequency of cancers in different familial phenotypes. J Med Genet. 2010 Jun;47(6):421‐8. (PMID 20522432) Saslow D et al. American Cancer Society Guidelines for Breast Screening with MRI as an Adjunct to Mammography. CA Cancer J Clin. 2007 May‐Jun; 57(3):185. (PMID 17392385) Schrader KA et al. Germline mutations in CDH1 are infrequent in women with early‐onset or familial lobular breast cancers. J Med Genet. 2011 Jan;48(1):64‐8. (PMID 20921021) Seppala EH et al. CHEK2 variants associate with hereditary prostate cancer. Br J Cancer. 2003 Nov 17;89(10):1966‐70. (PMID 14612911) Slater EP et al. PALB2 mutations in European familial pancreatic cancer families. Clin Genet. 2010 Nov;78(5):490‐4. (PMID 20412113) Suchy J et al. CHEK2 mutations and HNPCC‐related colorectal cancer. Int J Cancer. 2010 Jun 15;126(12):3005‐9. (PMID 19876921) Surveillance, Epidemiology, and End Results (SEER) Program of the National Cancer Institute. SEER Cancer Statistics Review, 1975‐2012: Lifetime Risk Tables (URL: http://surveillance.cancer.gov/devcan) [February 2016 accessed]. Tavtigian SV et al. Rare, evolutionarily unlikely missense substitutions in ATM confer increased risk of breast cancer. Am J Hum Genet. 2009 Oct;85(4):427‐46. (PMID 19781682) The CHEK2 Breast Cancer Consortium. CHEK2*1100delC and susceptibility to breast cancer: a collaborative analysis involving 10,860 breast cancer cases and 9,065 controls from 10 studies. Am J Hum Genet. 2004 Jun;74(6):1175‐82. (PMID 15122511) Thompson D et al. Cancer risks and mortality in heterozygous ATM mutation carriers. J Natl Cancer Inst. 2005 Jun 1; 97(11):813‐22. (PMID 15928302) Van der Groep P, van der Wall E, and van Diest PJ. Pathology of hereditary breast cancer. Cell Oncol (Dordrecht). 2011 Apr;34(2):71‐88. (PMID 21336636) Walsh T et al. Mutations in 12 genes for inherited ovarian, fallopian tube, and peritoneal carcinoma identified by massively parallel sequencing. Proc Natl Acad Sci U S A. 2011 Nov 1;108(44):18032‐7. (PMID 22006311) Gene Most commonly associated cancers BRCA1 Breast, Ovarian, Pancreatic, Prostate, Endometrial serous carcinoma^ Breast, Ovarian, Pancreatic, Prostate, Endometrial serous carcinoma^ Associated recessive syndrome Management Guidelines High Risk Genes NM_007294.3 BRCA2 NM_000059.3 CDH1 NM_004360.3 PALB2 NM_024675.3 PTEN NM_000314.4 TP53 NM_000546.5 ATM NM_000051.3 CHEK2 NM_007194.3 NCCN‐BR/OV Fanconi anemia NCCN‐BR/OV, CAPS# Gastric, Breast, Colon (signet ring)^ NCCN‐BR/OV, NCCN‐Gastric Breast, Pancreatic (possibly high risk)^ Fanconi anemia NCCN‐BR/OV, CAPS# Breast, Thyroid, Endometrial Breast, Sarcoma, Brain, Hematologic malignancies, Adrenocortical, among others* Moderate Risk Genes NCCN‐BR/OV NCCN‐BR/OV Breast, Colon^, Pancreatic^ Ataxia telangiectasia NCCN‐BR/OV Breast, Prostate (possibly high risk)^, Colon^ NCCN‐BR/OV, NCCN‐CRC Key Red font denotes significantly increased cancer risk. We consider significantly increased risk to be a relative risk of 4 or higher in relation to the general population risk. This translates to the following lifetime cancer risks: ≥50% breast cancer, ≥20% colon cancer, ≥11% endometrial cancer, ≥8% melanoma, ≥7% renal cancer, ≥6% pancreatic cancer, ≥6% ovarian cancer, ≥5% thyroid cancer, ≥4% gastric or small bowel cancer. Information Sheet on Breast High/Moderate Risk panel Page 6 of 7 © GeneDx Revision Date: 07/2016 Blue font denotes moderately increased cancer risk. We consider moderately increased risk to be a relative risk between 2 and 4 in relation to the general population risk. This translates to the following lifetime cancer risks: 24‐49% breast cancer, 32‐60% prostate cancer, 5‐11% endometrial cancer, 3‐6% pancreatic cancer. ^ Gene‐specific risk for this cancer type is not well‐defined. * High overall risk of cancer: 75% lifetime risk for males to develop cancer, nearly 100% risk for females. # CAPS ‐ International Cancer of the Pancreas Screening (CAPS) Consortium summit on the management of patients with increased risk for familial pancreatic cancer (Canto 2013). Information Sheet on Breast High/Moderate Risk panel Page 7 of 7 © GeneDx Revision Date: 07/2016