* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Explore the 3D Structure of Insulin
Point mutation wikipedia , lookup
Genetic code wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Catalytic triad wikipedia , lookup
Peptide synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
Homology modeling wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Metalloprotein wikipedia , lookup
Biosynthesis of doxorubicin wikipedia , lookup
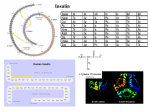
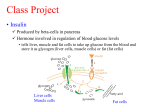
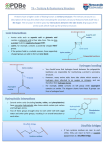
Explore the 3D Structure of Insulin pdb101.rcsb.org The Insulin hormone controls blood glucose levels. To learn more about the function of insulin, visit pdb101.rcsb.org and read the Molecule of the Month on Insulin. Primary Structure To learn more about the primary, secondary, tertiary and quaternary levels of protein structure, watch the What is a Protein? video at bit.ly/1JkBKgZ. Cut out the five strips on the dotted line. Make sure to separate the conjoined strips. The green, orange, and yellow tabs have to remain attached to their respective strips. 2. Protein Chains: 2.1. Chain A: Tape together the two pieces of the A chain (green) so that residue 11 is back-to-back in the two pieces, as shown in the picture. Proteins are polymer CHAINS of amino acid residues. Each number on the model represents one amino acid residue. 2.2. Chain B: Tape together the three pieces of the B chain (blue) by overlapping the gray tabs at the ends, so that all the numbers are visible on one side and all the numbers are in order. Insulin has 2 chains. Chain A is green; chain B is blue. Secondary Structure 1. Preparation: 3. Alpha Helices: N-H and C-O groups forming hydrogen bond Hydrogen bonds Alpha Helix Some sections of amino acid chains curl and form ALPHA HELICES due to HYDROGEN BONDS between N-H and C-O groups. The helix regions on the model are marked on the chains with darker shades of green and blue. 3.1. Chain A: Starting with residues 1 and 5 form a hydrogen bond between C-O and N-H, and tape together. 3.4. Chain B: Starting with residues 7 and 11, form a hydrogen bond. Keep creating the alpha helix until you reach the end of the darker colored area. 3.2. Keep creating the hydrogen bonds until you reach the end of the darker colored area. When you reach residue 6, make sure the green tab is on the outside of the helix. 3.3. Create another alpha helix starting with residues 17 and 21. Keep creating hydrogen bonds until you reach the end of the darker colored area. Turn 4. Turns: Some amino acids give a TURN to the chain. The turns are marked on chain B of the paper model with the white dashed lines. 4.1. Fold outwards on the white dashed lines to create the turns. You can secure the folds with tape. Turn Tertiary and Quaternary Structure Note: Chain B also includes a beta strand marked with a blue arrow. 5. Disulfide Bridges: The structure of Insulin is stabilized by 3 DISULFIDE BRIDGES. A disulfide bridge is formed when a sulfur atom from the residue cysteine forms a single covalent bond with a sulfur atom from a second cysteine. 5.1. The first disulfide bridge connects 2 5.2. Connect the yellow tabs, matching cysteines in chain A. To form the bridge, their shape while making sure the colored connect the green tabs as seen on the photo. sides are facing the same way. Complete model: PDB ID 1trz sulfur Æ atom The second and third disulfide bridges (represented by orange and yellow tabs) are formed between chains A and B. Disulfide Bridge Æy u 3D 5.3. Connect the orange tabs matching their shape while making sure the colored sides are facing the same way. Go to pdb101.rcsb.org (Learn > Paper models) to DOWNLOAD additional copies of this model, to watch a VIDEO DEMONSTRATION of how to build it, and to access a DIGITAL ACTIVITY PAGE allowing for further exploration of the 3D model. PDB-101 is the educational portal of RCSB Protein Data Bank (rcsb.org) N H 28 N H 27 N H 26 N H 25 O C 15 24 O C N H N H 14 23 O C N H N H 13 22 O C N H N H 12 21 O C N H N H 11 N H 10 N H N H 8 N H N H 7 O C 1 N H CHAIN B C O O C 21 H N N H O 2 O C O C N H 9 O C N H 3 O C O 20 H N 2 N H C 1 O C N H 4 O C O 19 H N 3 N H C N H O C 5 O C O 18 H N 4 N H C O C O C N H O 6 O C 16 17 H N 5 O C N H C O C O C 16 N H O C O C O C O H N 6 O C N H C O C O C 17 N H O C O O 15 H N 7 O C N H C CHAIN A O 14 8 H N O C O C N H C PDB ID 1trz: E. Ciszak, G. D. Smith (1994) Crystallographic evidence for dual coordination around zinc in the T3R3 human insulin hexamer. Biochemistry 33: 1512-1517 O C 18 N H O C 11 12 13 C N H 9 H N O C N H 10 H N O C N H 11 O C C The template for this paper model is based on the human insulin structure (PDB entry 1trz, chains A and B). # rcsb.org 30 29 O C 19 N H O C # N H # N H N H 20 O C O C # O C pdb101.rcsb.org Explore the 3D Structure of Insulin PDB-101 is the educational portal of RCSB Protein Data Bank (rcsb.org)