* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Inflammation in the Lung
Survey
Document related concepts
Rheumatic fever wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
DNA vaccination wikipedia , lookup
Sociality and disease transmission wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
Adaptive immune system wikipedia , lookup
Inflammation wikipedia , lookup
Neonatal infection wikipedia , lookup
Hospital-acquired infection wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Immune system wikipedia , lookup
Infection control wikipedia , lookup
Immunosuppressive drug wikipedia , lookup
Hygiene hypothesis wikipedia , lookup
Transcript
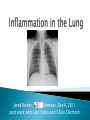
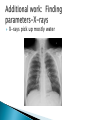
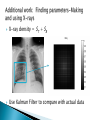
Jared BarberSeminar, Oct 4, 2011 Joint work with Ivan Yotov and Gilles Clermont Background on pneumonia and inflammation ODE Model ◦ Model ◦ Desired behaviors ◦ Results PDE Model ◦ Model additions ◦ Results Conclusions Future Work A condition where inflammation in the lung compromises lung function A leading cause of death in elderly, very young, chronically ill, and third world Caused by bacteria, virus, fungi, parasites ◦ Bacteria associated with most severe cases ◦ Flu can cause pneumonia Associated with coughing, fever, chills, lack of breath, confusion in elderly. Treated with fluids, antibiotics, oxygen therapy, breathing treatments Players ◦ Pathogen-introduced via air Bacteria-b ◦ Immune cells Neutrophils-n Macrophages-m m ◦ Cytokines Pro-inflammatory-cp Anti-inflammatory-ca Process (to right) cp b n ca Parameters chosen so that ◦ Healthy steady state is stable ◦ Neutrophils outkill/outnumber macrophages ◦ At maximal anti-inflammation levels, immune response is reduced by 75-80% ◦ Pro and anti-inflammatory cytokine levels are of the same order ◦ Anti-inflammatory cytokines delayed wrt proinflammatory cytokines There are some rarer desired behaviors that are not currently reproducible by the model 0.005 0 0 50 -4 2 x 10 100 150 cp 200 1 Bacterial infection is cleared by local immune response without needing to activate macrophages and neutrophils 0 0 50 100 150 ca 200 50 100 150 200 150 200 150 200 -5 5 x 10 0 0 -5 b 0.01 4 0.005 2 0 0 50 -4 2 x 10 100 150 cp 200 2 50 100 150 ca 200 50 100 200 -5 x 10 0 0 -5 2 0 50 100 n x 10 1 0 0 4 x 10 -4 1 5 0 m x 10 150 t in mhrs 0 0 50 100 t in hrs 0.1 0 0 50 b 2 0.1 1 0.05 0 50 100 150 cp 200 0 0.02 0.1 0.01 0 50 b 100 150 ca 200 0 0.1 0.2 1 0.05 0.1 50 100 150 cp 200 0 0.02 0.1 0.01 0 50 0.05 0.01 100 150 200 0 0 50 100 150 200 0 50 0 0 50 t in hrs n 0.2 Note: We have all desired behaviors 0.1 50 100 150 m 0.02 200 t in hrs 0.1 0 100 150 ca 50 0 m 0.2 0 0 50 200 0 0 50 100 150 100 150 200 100 150 200 150 200 n 2 0 0 m 0.2 0 0 200 Bacterial infection grows initially and then is destroyed by activated immune cells which subsequently decay to zero 0 100 150 ca 200 100 t in hrs 0.5 0 0 50 b 20 0.4 10 0.2 Bacterial infection is initially reduced but recovers once anti-inflammatory cytokines kick in 0 0 50 100 150 cp 200 0 0.1 0.5 0.05 0 50 b 100 150 ca 200 0 0.4 0.4 10 0.2 0.2 50 100 150 cp 200 0 0.1 0.5 0.05 0 50 100 150 ca 200 t in hrs 0.4 0.05 100 150 200 0 50 100 150 200 t in hrs n 0.2 0 50 100 150 m 0.1 0 0.4 0.2 0 50 m 1 0 0 200 0 0 50 100 150 50 100 150 200 0 50 100 150 200 150 200 n 20 0 0 200 m 1 0 0 100 150 ca 200 0 0 50 100 t in hrs 0.2 0 0 50 b 20 0.4 10 0.2 100 150 ca Introduction of additional bacteria later on can turn a healthy situation (Simulation 2) into an unhealthy one 0 0 50 100 150 cp 200 0 0 50 100 200 150 200 150 200 150 200 m 0.4 0.05 0.2 0 0 50 b 100 150 ca 200 0 0.4 0.2 10 0.2 0.1 0 50 100 150 cp 200 0 0 50 100 150 200 150 200 150 200 m 0.4 0.05 0.2 0 0 50 100 150 ca 200 0 0.2 0.2 0.1 0 50 100 150 m 0.05 50 100 n 0.4 0 0 200 0 0 50 100 50 100 n 20 0 0 0 0 50 100 Patients seen are usually Type II or Type III We want O(Type II) ≈ O(Type III), not the case 2 Type III Region 1.8 bacteria initial condition 1.6 Type II Region 1.4 1.2 1 0.8 Type I Region 0 1 2 3 4 5 6 7 8 relative speed at which immune cells act 9 10 Diffusion ◦ All species ◦ Smaller species (cytokines) diffuse more than larger species (inflammatory cells) Chemotaxis ◦ Macrophages migrate towards regions of high bacterial and cytokine concentration ◦ Neutrophils migrate towards regions of high cytokine concentration Lung made up of three components: ◦ Air/Alveolar region (A-90% of the lung) ◦ Blood (B-5% of the lung) ◦ Tissue (T-5% of the lung) Inflammation indicator function n( x, y, z ) kc p n c p ( x, y, z ) ( x, y , z ) 1 kca ca ( x, y, z ) Local saturation function 1 ST ,ref ST ( x, y, z ) 1 1 mT ( x, y, z ) Saturation for other components: 1 ST ,ref ST ( x, y , z ) 1 1 mT ( x, y, z ) 1 ST ( x, y , z ) S A ( x, y, z ) S A,ref 1 S T , ref 1 ST ( x, y , z ) S B ( x, y, z ) S B ,ref 1 S T , ref Effective diffusion/chemotaxis coefficients depend on air, blood, and tissue saturation: D( x, y, z ) DA S A ( x, y, z ) DB S B ( x, y, z ) DT ST ( x, y, z ) ( x, y, z ) A S A ( x, y, z ) B S B ( x, y, z ) T ST ( x, y, z ) For macrophages: Dm ( x, y, z ) 0.00015 S A ( x, y, z ) 0 S B ( x, y, z ) 0.00045 ST ( x, y, z ) Bacterial infection is cleared and immune system returns to original steady state Note: Actual Comp Domain 20x20 cm Time: Each profile 2hrs apart B 5 0.08 0.3 4 0.06 0.2 3 0.04 2 0 0.1 0.02 1 0 50 100 0 0 50 100 0 0 50 100 ST 0.05 0.6 0.04 1 0.4 0.03 0.02 0.5 0.2 0.01 0 0 50 100 0 0 50 100 0 0 50 t = 104 100 Note: Actual Comp Domain 20x20 cm Bacterial infection is not cleared and system proceeds to death Note: Actual Comp Domain 20x20 cm Bacterial infection is not cleared and system proceeds to death 10 9 Type III Region PDEs 8 bacteria initial condition Note: Actual Comp Domain 20x20 cm 7 6 PDE Divider? 5 4 Type II Region PDEs 3 ODE Divider 2 1 0 1 2 3 4 5 6 7 relative speed at which immune cells act 8 9 10 Type I Region Both models can produce desired behavior PDE model allows more Type II simulations ◦ PDE system starts with less bacterial load PDE vs ODE ◦ Diffusion lessens virility of bacterial growth ◦ Chemotaxis allows inflammatory cells to gang up on the bacteria PDE model gives much more flexibility X-rays pick up mostly water X-ray density = ST + SB Use Kalman Filter to compare with actual data Further refine ODE model to obtain more desired behaviors Consider including other members ◦ Damage ◦ Adaptive immune response For chemotaxis and diffusion coefficients, ◦ Maximize number of physiologically realistic simulations ◦ Find conditions to limit pattern formation from occurring ◦ Use smaller initial size of infection Obtain average values of neutrophils and cytokines in addition to x-rays to use for parameter estimation