* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 7.014 Section Problem:
Fatty acid metabolism wikipedia , lookup
Citric acid cycle wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Western blot wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Point mutation wikipedia , lookup
Peptide synthesis wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Genetic code wikipedia , lookup
Metalloprotein wikipedia , lookup
Protein structure prediction wikipedia , lookup
Catalytic triad wikipedia , lookup
Biochemistry wikipedia , lookup
Biosynthesis wikipedia , lookup
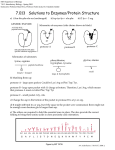
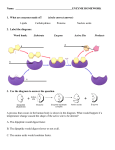
7.014 Section Problem: There is a class of related enzymes called serine proteases that all use the same mechanism to cleave peptide bonds. Each member of this family cleaves protein substrates at a different location – that is, each enzyme cleaves protein substrates after a different amino acid(s). Formally, they catalyze the following reaction: A-B-C-D-X-E-F-G-H + H2O --------------->A-B-C-D-X protein substrate + protein fragment E-F-G-H protein fragment Where A through G are any amino acid and X is one of the specific amino acids uniquely recognized by the enzyme. If this process repeats over and over, the substrate will be completely degraded into small peptide fragments. Each member of the family cleaves after a specific amino acid of the substrate because that amino acid is recognized by a binding pocket of the enzyme that is specifically designed to bind that particular amino acid. a) One of these protease enzymes, trypsin, cleaves after lys or arg. What would be the product(s) if each of the following molecules were treated with trypsin? i) leu-thr-phe-ala-ser ii) trp-tyr-lys-ala-phe iii) lys-arg-lys-arg The structures of three hypothetical proteases are shown below (not to scale – the actual enzyme is much larger than the binding pocket). If the amino acid side chain of the substrate binds well to the “recognition pocket” the substrate will bind and be cleaved. GENERIC ENZYME: ACTIVE SITE peptide bond cleavage gly gly gly RECOGNITION POCKET specific substrate regognition protease A gly gly asp protease B ile val phe protease C b) The specificities of each protease (the amino acids that it likes to cut after) are listed below. Match the enzyme with the specificity & explain. i) lysine, arginine ii) phenylalanine, tryptophan, tyrosine iii) glycine, alanine c) How could you design a similar enzyme to cleave after aspartic acid? d) Speculate on the effect of changing the aspartic acid in protease B to a glutamic acid. e) There are three amino acids required for the active site to function and three amino acids involved in substrate recognition – why then do these enzymes typically contain more than 200 amino acids? f) Suppose you make a solution of protease A. A small sample taken when the solution was made (time = 0) is capable of cleaving 100 mmol of protein substrate per minute. This rate of substrate cleavage is called the “activity” of the enzyme. You then take identical small samples from this solution at regular intervals over the next few hours as the solution stands at room temperature and measure their activity (the rate at which they cleave the substrate protein). You find that the activity of the enzyme drops rapidly as time passes. However, if you add a large excess of casein (a protein found in milk which has no enzymatic activity), the protease loses its activity much more slowly. These data are sketched below: activ ity 100 protease A + casein protease A alone 0 0 tim e Explain these observations. g) If you monitor the reaction of any of these proteases as they degrade a protein, you observe that the protease does not cut all the recognition sites in the substrate at once – the recognition sites on the surface of the substrate protein are the first to be cut. Explain. h) How might your answer to parts (f) and (g) be combined to design a long-lasting protease A molecule? STRUCTURES OF AMINO ACIDS GENERIC AMINO ACID: O O Individual amino acids are linked through these groups to form the backbone of the protein. O O H H C CH3 H O CH2 SH H NH3 + H N C O + C CH2 NH3 + N C H H H O C CH2CH2 S CH3 H NH3 + C O CH2CH3 H C H CH2 H CH3 NH3 OH + THREONINE (thr) C CH2 H H H NH3 + H H O CH2CH2CH2CH2 LYSINE (lys) H O O C H C C H O O C CH2 OH NH3 + H TYROSINE (tyr) H OH SERINE (ser) C H CH2 NH3 + PROLINE (pro) O C NH3 + C H C CH2 CH2 H N CH 2 H + H H H CH3 O O H C TRYPTOPHAN (trp) C CH3 H O C LEUCINE (leu) N C GLYCINE (gly) O NH3 + H C H NH3 + H PHENYLALANINE (phe) O H NH2 C CH2 NH3 + METHIONINE (met) O O C H C C O O O O CH2CH2 C C C C C GLUTAMINE (glN) O C H O O NH3 + O ASPARTIC ACID (asp) O C CH2 C NH3 + O H O ISOLEUCINE (ile) C H C O C NH2 C NH3 CH3 + H HISTIDINE (his) H CH2CH2 H ASPARAGINE (asN) GLUTAMIC ACID (glu) CYSTEINE (cys) C NH3 + O C CH2 C O NH3 + O NH2 C O C NH2 + C C O C O O C H CH2CH2CH2 N H O O C H C Peptide bonds O O ARGININE (arg) O O Side chain, unique to each differnt amino acid NH3 + ALANINE (ala) H C R NH3 + C NH3 + O C O C O H R1 H O H R 3 C C N C C C N N C R H 2 H H O Protein Synthesis H C CH3 C NH3 H + CH3 VALINE (val) NH3+ Solutions to: Enzymes/Protein Structure a) i) leu-thr-phe-ala-ser (unchanged) GENERIC ENZYME: ii) trp-tyr-lys + ala-phe iii) 2 lys + 2 arg Schematics of enzymes: (side chains shown in black) ACTIVE SITE peptide bond cleavage gly gly gly gly ile - asp gly protease A val phe protease C protease B RECOGNITION POCKET specific substrate regognition Schematics of substrates: lysine, arginine peptide backbone phenylalanine, tryptophan, tyrosine glycine, alanine + long & + charged small large & hydrophobic b) Matching them up: protease A - large open pocket. Could be lys/arg or phe/trp/tyr. protease B - large open pocket with (-) charge at bottom. Therefore, lys/arg, which means that protease A must cut after phe/trp/tyr protease C - small pocket. Cuts after Gly, ala. c) Change the asp in the bottom of the pocket in protease B to a lys or arg. d) It might still bind lys or arg, but if the space in the pocket were constrained, there might not be enough room because glu is longer than asp. e) The others are required to hold the essential ones in place. f) Protease A is a protein, therefore other protease A molecules can cleave it and thereby inactivate it. Having casein around decreases the chance that a protease A molecule will cleave a protease A molecule, because it will be more likely to cleave the more numerous casein molecules. g) The protease enzyme must be able to bind to the target amino acids. If they are buried inside the target protein, the protease can't "see" them and therefore can't cut at them. Eventually, the structure of the target protein gets so broken down that the inside amino acids are exposed to the protease. h) To protect protease A from degradation by other protease A molecules, bury all the phe/trp/tyr inside the protein so that they cannot be recognized and cleaved.