WHITE PANICLE1, a Val-tRNA Synthetase
... nski et al., 2000; Guo et al., 2010). All cells must contain a complete set of 20 AARSs. Eukaryotic cells possess more than the basal set of 20 AARS genes, because organelles such as plastids and mitochondria possess their own translational systems (Duchêne et al., 2009). In Arabidopsis, 45 expresse ...
... nski et al., 2000; Guo et al., 2010). All cells must contain a complete set of 20 AARSs. Eukaryotic cells possess more than the basal set of 20 AARS genes, because organelles such as plastids and mitochondria possess their own translational systems (Duchêne et al., 2009). In Arabidopsis, 45 expresse ...
Vegetative incompatibility in filamentous fungi: Podospora and
... display a cellular function. Different fungal species have at least partially different sets of het genes. For instance, the mating-type locus that functions as a het locus in N. crassa is not a het locus in numerous other ascomycetes (see references in [12•]), and the P. anserina homologue of the N ...
... display a cellular function. Different fungal species have at least partially different sets of het genes. For instance, the mating-type locus that functions as a het locus in N. crassa is not a het locus in numerous other ascomycetes (see references in [12•]), and the P. anserina homologue of the N ...
MGF 360-17R Missing
... Trunc – 014 [MGF 110-7L/MGF 360-6L Fusion Protein]: This gene is a fusion between the MGF 110-7L ortholog and MGF 360-6L. The carboxy terminus of this fusion is shown as a small box between the 3L and 4L orthologs as the MGF 110 orthologs are outside of the scope of this diagram. The annotated ortho ...
... Trunc – 014 [MGF 110-7L/MGF 360-6L Fusion Protein]: This gene is a fusion between the MGF 110-7L ortholog and MGF 360-6L. The carboxy terminus of this fusion is shown as a small box between the 3L and 4L orthologs as the MGF 110 orthologs are outside of the scope of this diagram. The annotated ortho ...
Gene conversion and purifying selection shape nucleotide variation
... When a sample from a male showed two nucleotide peaks at one or more sites (“heterozygous” sites) in the L or M opsin gene, the individual should have two or more loci of the gene with different sequences. In this case, we conservatively inferred two loci for this gene. When a female showed heterozy ...
... When a sample from a male showed two nucleotide peaks at one or more sites (“heterozygous” sites) in the L or M opsin gene, the individual should have two or more loci of the gene with different sequences. In this case, we conservatively inferred two loci for this gene. When a female showed heterozy ...
Archives of Microbiology
... of the VnfDG products are altered by this gene fusion, nor whether the unique addition of 21 nucleotides in the vnfDG fusion area of Anabaena sp. CH1 is of functional signiWcance. In a neighbor-joining analysis of all available deduced VnfDG sequences, those from cyanobacteria clustered next to thos ...
... of the VnfDG products are altered by this gene fusion, nor whether the unique addition of 21 nucleotides in the vnfDG fusion area of Anabaena sp. CH1 is of functional signiWcance. In a neighbor-joining analysis of all available deduced VnfDG sequences, those from cyanobacteria clustered next to thos ...
Control of ribosome traffic by position-dependent
... The translation rate Rx at a position x depends on the codon at this position. We use the codondependent rates determined in ref. [11], as summarized in table 1. To simplify the model, we assign one of the following three rates to each codon; A (∼35 codons/sec), B (∼8 codons/sec), and C-rate (∼4.5 c ...
... The translation rate Rx at a position x depends on the codon at this position. We use the codondependent rates determined in ref. [11], as summarized in table 1. To simplify the model, we assign one of the following three rates to each codon; A (∼35 codons/sec), B (∼8 codons/sec), and C-rate (∼4.5 c ...
msb201035-sup
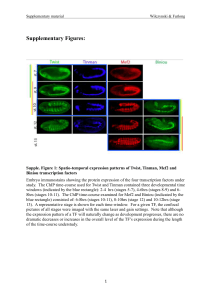
... X-axis represents developmental time in hours. Box-plot colors indicate temporal TFbinding classes (green: early, yellow: continuous, red: late). Temporal classes show very different distributions of signals between early and late time-points. For example, there is very little TF occupancy (ChIP sig ...
... X-axis represents developmental time in hours. Box-plot colors indicate temporal TFbinding classes (green: early, yellow: continuous, red: late). Temporal classes show very different distributions of signals between early and late time-points. For example, there is very little TF occupancy (ChIP sig ...
Käfer, E. and D. Luk
... Käfer 1981 Mutat. Res. 80:43-64) and UV survival was determined in several experiments (Figure 1). The obtained results permit a rough comparison between new and "old" alleles. and also between mutants in different genes. For example, the new allele mus(FK129 ) of mus-9 more closely resembles mus(FK ...
... Käfer 1981 Mutat. Res. 80:43-64) and UV survival was determined in several experiments (Figure 1). The obtained results permit a rough comparison between new and "old" alleles. and also between mutants in different genes. For example, the new allele mus(FK129 ) of mus-9 more closely resembles mus(FK ...
What Size Are Your Genes?
... different forms of our gene of interest, as represented by the yellow, purple, and blue dyes. Agarose gel electrophoresis separates the gene fragments into discrete bands, each comprising molecules of the same size. How can these results be used to determine the lengths of different fragments? Remem ...
... different forms of our gene of interest, as represented by the yellow, purple, and blue dyes. Agarose gel electrophoresis separates the gene fragments into discrete bands, each comprising molecules of the same size. How can these results be used to determine the lengths of different fragments? Remem ...
(type I) and mannose-resistant F8 (P) fimbriae of Escherichia coli
... shown in this paper the genetic determinants of F8 fimbriae were m a p p e d at a position between 17--20 on the chromosome of strain 2980 of E. coli O18:K5. Therefore, it seems that the gene clusters coding for type I fimbriae are located at fixed positions on the chromosomes of different strains. ...
... shown in this paper the genetic determinants of F8 fimbriae were m a p p e d at a position between 17--20 on the chromosome of strain 2980 of E. coli O18:K5. Therefore, it seems that the gene clusters coding for type I fimbriae are located at fixed positions on the chromosomes of different strains. ...
Conserved syntenic clusters of protein coding genes are missing in
... Results: Using comparative genomics based on extensive searches of 60 avian genomes, we have found that birds lack approximately 274 protein coding genes that are present in the genomes of most vertebrate lineages and are for the most part organized in conserved syntenic clusters in non-avian saurop ...
... Results: Using comparative genomics based on extensive searches of 60 avian genomes, we have found that birds lack approximately 274 protein coding genes that are present in the genomes of most vertebrate lineages and are for the most part organized in conserved syntenic clusters in non-avian saurop ...
Drosophila Muller F Elements Maintain a Distinct Set of Genomic
... The placements of the fosmid end reads were specified in the reads. placed file in each CAF1 assembly. The F and D element scaffolds were partitioned into a list of overlapping fosmids based on the reads.placed file for each species. This set of fosmids was obtained from the Drosophila Genomics Resourc ...
... The placements of the fosmid end reads were specified in the reads. placed file in each CAF1 assembly. The F and D element scaffolds were partitioned into a list of overlapping fosmids based on the reads.placed file for each species. This set of fosmids was obtained from the Drosophila Genomics Resourc ...
Conserved syntenic clusters of protein coding genes are missing in birds
... many conserved genes that could not be found in the previous assembly (for example, [20]), and yielded significant BLAT-alignments for approximately 96% of genes from a positive control search set consisting of randomly selected lizard gene models with known orthologs in birds). Lastly, this subset ...
... many conserved genes that could not be found in the previous assembly (for example, [20]), and yielded significant BLAT-alignments for approximately 96% of genes from a positive control search set consisting of randomly selected lizard gene models with known orthologs in birds). Lastly, this subset ...
Classifying Gene Expression Data using an Evolutionary Algorithm
... difficult than the binary classification. The main difficulties in solving microarray classification are the availability of very small amount of samples compared to the number of genes in the sample and the extremely large search space of solutions. Moreover, the variation in microarray experiment ...
... difficult than the binary classification. The main difficulties in solving microarray classification are the availability of very small amount of samples compared to the number of genes in the sample and the extremely large search space of solutions. Moreover, the variation in microarray experiment ...
Alfred Henry Sturtevant - National Academy of Sciences
... occurs. The factors responsible were isolated by Sturtevant and by Muller around 1915 and were shown to act as dominant cross-over suppressors. The first clue to the nature of these factors came in 1921, when Sturtevant compared the chromosome maps of Drosophila melanogaster with those of D. simulan ...
... occurs. The factors responsible were isolated by Sturtevant and by Muller around 1915 and were shown to act as dominant cross-over suppressors. The first clue to the nature of these factors came in 1921, when Sturtevant compared the chromosome maps of Drosophila melanogaster with those of D. simulan ...
Threshold phenomena versus cell heredity in the
... for sex-linked genes for internal evidence concerning their origin. This paper deals with the genes of the house mouse for tabby {Ta; Falconer, 1953), brindled (Mobr; Fraser, Sobey & Spicer, 1953) and striated (Str; Phillips, 1963). The test for the validity of the L.H. is essentially a comparison b ...
... for sex-linked genes for internal evidence concerning their origin. This paper deals with the genes of the house mouse for tabby {Ta; Falconer, 1953), brindled (Mobr; Fraser, Sobey & Spicer, 1953) and striated (Str; Phillips, 1963). The test for the validity of the L.H. is essentially a comparison b ...
Development of novel computational tools based on
... gene transfer, defined as a mechanism that promotes the transfer of foreign genomic segments between lineages was found to be relatively common in prokaryotes and less common in higher-order organisms. This mechanism effectively contributes to the evolution and diversity of bacterial species by the ...
... gene transfer, defined as a mechanism that promotes the transfer of foreign genomic segments between lineages was found to be relatively common in prokaryotes and less common in higher-order organisms. This mechanism effectively contributes to the evolution and diversity of bacterial species by the ...
PDF
... for sex-linked genes for internal evidence concerning their origin. This paper deals with the genes of the house mouse for tabby {Ta; Falconer, 1953), brindled (Mobr; Fraser, Sobey & Spicer, 1953) and striated (Str; Phillips, 1963). The test for the validity of the L.H. is essentially a comparison b ...
... for sex-linked genes for internal evidence concerning their origin. This paper deals with the genes of the house mouse for tabby {Ta; Falconer, 1953), brindled (Mobr; Fraser, Sobey & Spicer, 1953) and striated (Str; Phillips, 1963). The test for the validity of the L.H. is essentially a comparison b ...
Polyploidy Enhances F Pollen Sterility Loci
... However, it was unclear how polyploidy affected allelic interactions between pollen sterility loci during pollen mother cell (PMC) meiosis in intersubspecific autotetraploid rice hybrids. Microarray and RNA sequencing (RNA-Seq)-based transcriptome profiling are helpful tools for characterizing molecul ...
... However, it was unclear how polyploidy affected allelic interactions between pollen sterility loci during pollen mother cell (PMC) meiosis in intersubspecific autotetraploid rice hybrids. Microarray and RNA sequencing (RNA-Seq)-based transcriptome profiling are helpful tools for characterizing molecul ...
Homeotic genes controlling flower development in Antirrhinum
... identity of whorls of organs they will be referred to as whorl identity genes (note that they do not affect the numbering of the whorls but only the identity of organs in the whorls). Similar classes of genes are observed in Arabidopsis (Bowman et al. 1989; Haughn and Sommerville, 1988; see also Mey ...
... identity of whorls of organs they will be referred to as whorl identity genes (note that they do not affect the numbering of the whorls but only the identity of organs in the whorls). Similar classes of genes are observed in Arabidopsis (Bowman et al. 1989; Haughn and Sommerville, 1988; see also Mey ...
Genes - Gerstein Lab Publications
... The pseudogene population (denoted G) arising from the decay of protein-coding genes in the worm is estimated to comprise 2,168 sequences which is about 12% of the total gene complement (G). This is only an initial estimate of the pseudogene population, that may be ...
... The pseudogene population (denoted G) arising from the decay of protein-coding genes in the worm is estimated to comprise 2,168 sequences which is about 12% of the total gene complement (G). This is only an initial estimate of the pseudogene population, that may be ...
Essential gene

Essential genes are those genes of an organism that are thought to be critical for its survival. However, being essential is highly dependent on the circumstances in which an organism lives. For instance, a gene required to digest starch is only essential if starch is the only source of energy. Recently, systematic attempts have been made to identify those genes that are absolutely required to maintain life, provided that all nutrients are available. Such experiments have led to the conclusion that the absolutely required number of genes for bacteria is on the order of about 250-300. These essential genes encode proteins to maintain a central metabolism, replicate DNA, translate genes into proteins, maintain a basic cellular structure, and mediate transport processes into and out of the cell. Most genes are not essential but convey selective advantages and increased fitness.