A mixed group ll/group III twintron in the Euglena
... structural analysis, we proposed that this 409 nt intron was a 'mixed' twintron in which a group II intron was internal to a group HI intron (15). To test this hypothesis, the polymerase chain reaction (PCR) was used to amplify rps3 pre-mRNA processing intermediates. The locations of the cDNA and PC ...
... structural analysis, we proposed that this 409 nt intron was a 'mixed' twintron in which a group II intron was internal to a group HI intron (15). To test this hypothesis, the polymerase chain reaction (PCR) was used to amplify rps3 pre-mRNA processing intermediates. The locations of the cDNA and PC ...
Cleavage, Deprotection and Isolation of Peptides after Fmoc Synthesis
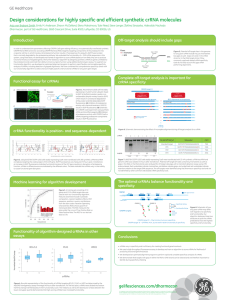
... Asn(Mbh/Tmob), Gln(Mbh/Tmob), as well as for peptides constructed on a PAL or Rink Amide resin (See Figure 2). While Reagent R may give higher cleavage yields, it is highly noxious and may not be preferable to handle. Reagent R or B are also recommended when extended cleavage times (those in excess ...
... Asn(Mbh/Tmob), Gln(Mbh/Tmob), as well as for peptides constructed on a PAL or Rink Amide resin (See Figure 2). While Reagent R may give higher cleavage yields, it is highly noxious and may not be preferable to handle. Reagent R or B are also recommended when extended cleavage times (those in excess ...
Mendel`s Accountant: A New Population Genetics Simulation Tool
... given by m = σ[(l − 1)µ + µx], where l is the index of the linkage subunit on which the mutation occurs and x is the value of the random number, with 0.0 ≤ x ≤ 1.0, that specifies the mutation’s fitness effect. We apply the modulo function with modulus µ to the absolute value of m to recover x. We div ...
... given by m = σ[(l − 1)µ + µx], where l is the index of the linkage subunit on which the mutation occurs and x is the value of the random number, with 0.0 ≤ x ≤ 1.0, that specifies the mutation’s fitness effect. We apply the modulo function with modulus µ to the absolute value of m to recover x. We div ...
Enzyme Mechanisms
... Subtilisin: externals very different from mammalian serine proteases; triad same ...
... Subtilisin: externals very different from mammalian serine proteases; triad same ...
somatic hypermutation of the 5' noncoding region of the Frequent MARTINOrrI*t,
... abnormal chromosomal band 3q27, the site of the BCL6 gene (4). BCL6 sequences representing exons 2-10 and spanning the entire coding region of the gene were examined by PCR/SSCP analysis using the set of primers illustrated in Fig. 1. No SSCP variants were observed in these sequences in 22 DLCL case ...
... abnormal chromosomal band 3q27, the site of the BCL6 gene (4). BCL6 sequences representing exons 2-10 and spanning the entire coding region of the gene were examined by PCR/SSCP analysis using the set of primers illustrated in Fig. 1. No SSCP variants were observed in these sequences in 22 DLCL case ...
Systematic and Applied Microbiology - digital
... These sequences of strains BIFI-56 and BIFI-58 showed a 100% identity to the same fragment of the16S rRNA sequence from the E. faecium type strain (ATCC 19434, DSM 20477).Therefore the 16S rRNA sequence revealed that both strains were E. faecium isolates. In order to know if both strains were atypic ...
... These sequences of strains BIFI-56 and BIFI-58 showed a 100% identity to the same fragment of the16S rRNA sequence from the E. faecium type strain (ATCC 19434, DSM 20477).Therefore the 16S rRNA sequence revealed that both strains were E. faecium isolates. In order to know if both strains were atypic ...
1. dia
... 1 g ClTrt-resin + 2 mmol Fmoc-Aaa(X)-OH + 8 mmol DIEA in 3-5 mL DCM, for 1.5 h then 0.8 mL MeOH to block the unreacted groups washing with DCM, iPrOH, MeOH, ether The final cleavage results in peptides with COOH group at the C-terminus Cleavage with 90-95% TFA + scavangers results in free peptides C ...
... 1 g ClTrt-resin + 2 mmol Fmoc-Aaa(X)-OH + 8 mmol DIEA in 3-5 mL DCM, for 1.5 h then 0.8 mL MeOH to block the unreacted groups washing with DCM, iPrOH, MeOH, ether The final cleavage results in peptides with COOH group at the C-terminus Cleavage with 90-95% TFA + scavangers results in free peptides C ...
Constructing Sequences for Oxytocin and Vasopressin
... The structure of oxytocin is very similar to that of the vasopressin (also known commonly as arginine vasopressin): Both are nonapeptides (peptides with nine amino acids) with a disulfide bridge and their amino acid sequence differs at only two positions. The two genes are located on the same chromo ...
... The structure of oxytocin is very similar to that of the vasopressin (also known commonly as arginine vasopressin): Both are nonapeptides (peptides with nine amino acids) with a disulfide bridge and their amino acid sequence differs at only two positions. The two genes are located on the same chromo ...
Similarity Searches on Sequence Databases: BLAST
... compositions, e.g. many proteins contain patches known as low-complexity regions: such as segments that contain many prolines or glutamic acid residues. • If BLAST aligns two proline-rich domains, this alignment gets a very good E-value because of the high number of identical amino acids it contains ...
... compositions, e.g. many proteins contain patches known as low-complexity regions: such as segments that contain many prolines or glutamic acid residues. • If BLAST aligns two proline-rich domains, this alignment gets a very good E-value because of the high number of identical amino acids it contains ...
Silica Particles
... Cells , Tissue, Viral NA isolation RNA Isolation Lysis Blood, Cells , Tissue, for (RT)-PCR Bacteria,Yeast Blood, Serum , Plasma Lysis Lysis ...
... Cells , Tissue, Viral NA isolation RNA Isolation Lysis Blood, Cells , Tissue, for (RT)-PCR Bacteria,Yeast Blood, Serum , Plasma Lysis Lysis ...
The Language of Life
... organisms that never shared the universal genetic code All changes in stop codons must include three changes: – Replacement of stop codons that do not code for stop anymore with those that still do – Production of new tRNAs with anticodons that recognize the codon as not stop anymore – Modification ...
... organisms that never shared the universal genetic code All changes in stop codons must include three changes: – Replacement of stop codons that do not code for stop anymore with those that still do – Production of new tRNAs with anticodons that recognize the codon as not stop anymore – Modification ...
Cells: building blocks of living organisms
... DNA molecules generally form right-hand side helices in B form, while RNA are A form, also right-hand side. A left-hand side helix exists that is called Z DNA. DNA molecules cannot exist as a single strand, they are degraded, i.e. cut into pieces. A DNA molecule is made of two complementary strands ...
... DNA molecules generally form right-hand side helices in B form, while RNA are A form, also right-hand side. A left-hand side helix exists that is called Z DNA. DNA molecules cannot exist as a single strand, they are degraded, i.e. cut into pieces. A DNA molecule is made of two complementary strands ...
Cloning and Expression of Cellulosimicrobium cellulans β
... by agarose gel (0.8% wt/v) electrophoresis. For restriction endonuclease analysis, recombinant plasmid pTE353-βG was isolated from L. plantarum transformant. To detect the β-1,3-glucanase gene, recombinant plasmid was cleaved with SacI+EcoRI restriction endonuclease mixture and analysed by agarose g ...
... by agarose gel (0.8% wt/v) electrophoresis. For restriction endonuclease analysis, recombinant plasmid pTE353-βG was isolated from L. plantarum transformant. To detect the β-1,3-glucanase gene, recombinant plasmid was cleaved with SacI+EcoRI restriction endonuclease mixture and analysed by agarose g ...
The evolution of meiotic sex and its alternatives
... are resolved without recombination (exchange of flanking regions, see red versus blue arrows). (Online version in colour.) natural selection [7], for directly repairing DNA double-strand breaks (DSBs) [8], or for removal of oxidative DNA damage in germline cells [9,10]. Prophase I would be needed fo ...
... are resolved without recombination (exchange of flanking regions, see red versus blue arrows). (Online version in colour.) natural selection [7], for directly repairing DNA double-strand breaks (DSBs) [8], or for removal of oxidative DNA damage in germline cells [9,10]. Prophase I would be needed fo ...
Now we turn to the study of chemical kinetics. Kinetics is the study of
... second factor is the concentrations of reactants and products. This should make qualitative sense. In order for two compounds to react, they have to meet. If the concentration of the reactants is higher, then they will be closer together, and it won't take as long for them to come together and react ...
... second factor is the concentrations of reactants and products. This should make qualitative sense. In order for two compounds to react, they have to meet. If the concentration of the reactants is higher, then they will be closer together, and it won't take as long for them to come together and react ...
Clin Infect Dis. - Repositorio Académico UPC
... software 115, which plotted the negative derivative of fluorescence with respect to temperature (–d(F)/dT vs T). To make sure that we did not have inhibition of amplification because of the presence of contaminants in stool samples, we randomly selected some stool samples from children, mixed them w ...
... software 115, which plotted the negative derivative of fluorescence with respect to temperature (–d(F)/dT vs T). To make sure that we did not have inhibition of amplification because of the presence of contaminants in stool samples, we randomly selected some stool samples from children, mixed them w ...
1 Single molecule sequencing of THCA synthase reveals
... intermittent haplogroups. We cannot rule out the possibility of other diverged copies of inactive THCAS stochastically amplifying given the PCR reactions were set up independently. Gradient PCR at lower annealin ...
... intermittent haplogroups. We cannot rule out the possibility of other diverged copies of inactive THCAS stochastically amplifying given the PCR reactions were set up independently. Gradient PCR at lower annealin ...
Engineered bacteriophage-defence systems in bioprocessing
... question exhibit distinct phagesensitivity profiles, meaning that they are attacked by different groups of phages. ...
... question exhibit distinct phagesensitivity profiles, meaning that they are attacked by different groups of phages. ...
Origin of metabolism
... We identify as modules, either subsets of chemicals and reactions, or subsets of functions, that are re-used in many contexts with a conserved internal structure. At the small molecule substrate level, module boundaries are often associated with the most complex reaction mechanisms, catalyzed by hig ...
... We identify as modules, either subsets of chemicals and reactions, or subsets of functions, that are re-used in many contexts with a conserved internal structure. At the small molecule substrate level, module boundaries are often associated with the most complex reaction mechanisms, catalyzed by hig ...
The nonenzymatic subunit of pseutarin C, a
... the deduced amino acid sequence. However, we observed a few differences that are summarized in Table 1. Such differences may indicate the presence of more than one isoform of this protein. Edman degradation of some of the peptides showed heterogeneity. For example, peptide Hc101-104 (LIQY; Hc indica ...
... the deduced amino acid sequence. However, we observed a few differences that are summarized in Table 1. Such differences may indicate the presence of more than one isoform of this protein. Edman degradation of some of the peptides showed heterogeneity. For example, peptide Hc101-104 (LIQY; Hc indica ...
The genomic landscape of meiotic crossovers and gene
... eLife digest Most living organisms package their DNA into bundles called chromosomes. These chromosomes generally form pairs, with each chromosome in the pair containing the same number of genes. The genes also come in the same order, but the exact sequence of DNA bases within the genes can be diffe ...
... eLife digest Most living organisms package their DNA into bundles called chromosomes. These chromosomes generally form pairs, with each chromosome in the pair containing the same number of genes. The genes also come in the same order, but the exact sequence of DNA bases within the genes can be diffe ...
Deoxyribozyme
_DNAzyme.png?width=300)
Deoxyribozymes, also called DNA enzymes, DNAzymes, or catalytic DNA, are DNA oligonucleotides that are capable of catalyzing specific chemical reactions, similar to the action of other biological enzymes, such as proteins or ribozymes (enzymes composed of RNA).However, in contrast to the abundance of protein enzymes in biological systems and the discovery of biological ribozymes in the 1980s,there are no known naturally occurring deoxyribozymes.Deoxyribozymes should not be confused with DNA aptamers which are oligonucleotides that selectively bind a target ligand, but do not catalyze a subsequent chemical reaction.With the exception of ribozymes, nucleic acid molecules within cells primarily serve as storage of genetic information due to its ability to form complementary base pairs, which allows for high-fidelity copying and transfer of genetic information. In contrast, nucleic acid molecules are more limited in their catalytic ability, in comparison to protein enzymes, to just three types of interactions: hydrogen bonding, pi stacking, and metal-ion coordination. This is due to the limited number of functional groups of the nucleic acid monomers: while proteins are built from up to twenty different amino acids with various functional groups, nucleic acids are built from just four chemically similar nucleobases. In addition, DNA lacks the 2'-hydroxyl group found in RNA which limits the catalytic competency of deoxyribozymes even in comparison to ribozymes.In addition to the inherent inferiority of DNA catalytic activity, the apparent lack of naturally occurring deoxyribozymes may also be due to the primarily double-stranded conformation of DNA in biological systems which would limit its physical flexibility and ability to form tertiary structures, and so would drastically limit the ability of double-stranded DNA to act as a catalyst; though there are a few known instances of biological single-stranded DNA such as multicopy single-stranded DNA (msDNA), certain viral genomes, and the replication fork formed during DNA replication. Further structural differences between DNA and RNA may also play a role in the lack of biological deoxyribozymes, such as the additional methyl group of the DNA base thymidine compared to the RNA base uracil or the tendency of DNA to adopt the B-form helix while RNA tends to adopt the A-form helix. However, it has also been shown that DNA can form structures that RNA cannot, which suggests that, though there are differences in structures that each can form, neither is inherently more or less catalytic due to their possible structural motifs.