* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Proteins
Cooperative binding wikipedia , lookup
Rosetta@home wikipedia , lookup
Protein design wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Immunoprecipitation wikipedia , lookup
Protein purification wikipedia , lookup
Structural alignment wikipedia , lookup
Protein folding wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein domain wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Homology modeling wikipedia , lookup
Circular dichroism wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Alpha helix wikipedia , lookup
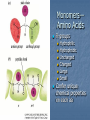
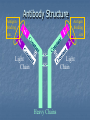
Proteins Protein Function Catalysis Structure Movement Defense Regulation Transport Antibodies Monomers— Amino Acids R-groups Hydrophilic Hydrophobic Uncharged Charged Large Small Confer unique chemical properties on each aa PROTEIN LEVELS OF STRUCTURE PRIMARY STRUCTURE Is a unique characteristic of every protein Is encoded by the nucleotide sequence of DNA Is thus a form of genetic information Is read from the amino terminus to the carboxyl terminus Nature of Protein Sequences Sequences and composition reflect the function of the protein: Membrane proteins have more hydrophobic residues. Homologous proteins from different organisms have similar sequences. e.g., cytochrome c is highly conserved Cytochrome c SECONDARY STRUCTURE I: THE -HELIX Helix If N-terminus is at bottom, then all peptide N-H bonds point “down” and all peptide C=O bonds point “up”. N-H of residue n is Hbonded to C=O of residue n+4. Secondary Structure II: The -Strand approx. 3.4 A Several -strands assemble into a -sheet (a tertiary structural element) TERTIARY STRUCTURE 3-D structure. Form follows function!! Native vs denatured Determinants of tertiary structure Amino acid sequence Environment in which the protein resides Stabilizing Interactions Hydrogen Bonds Electrostatic interactions (“salt-bridges” or ion pairs) van der Waals interactions (dipole-dipole and dispersion) Hydrophobic interactions Disulfide bridges Protein Denaturation •Denaturants--Anything stabilizing interactions • • • • Heat Salts pH Organic solvents that can disrupt Quaternary Structure ANTIBODIES Extremely specific Definitions: Antigen Epitope (antigenic determinant) Hapten FLUORESCEIN – a hapten Antibody Structure Antigen binding site V V V Light Chain Antigen binding site V SS SS Heavy Chains Light Chain Antibody Structure Antibody Structure Recognition and Binding The N-terminal region of antibody light chains and heavy chains form the antigen binding site The variability in amino acid sequence provides the structural basis for the diversity of antigen-binding sites Antigen Binding Antigen 1 Antigen 3 Polyclonal vs Monoclonal Abs 107-109 genetically distinct lymphocytes, each producing a single type of Ab. Polyclonal—normal immune response. Several Abs, recognition of various epitopes with varying affinities. Monoclonal Monoclonal Ab Production Given: Normal cells—Mortal Transformed cells—Immortal Two Pathways of DNA Synthesis Major Salvage—Requires HGPRT 8-azaguanine—HGPRT poison. Aminopterin---Interferes w/ major pathway PEG---promotes cell fusion HAT Selection 1) 2) 3) Select HGPRT- mutant myeloma by treatment with 8azaguanine Fuse HGPRT- mutant myeloma with normal cells using PEG Select with aminopterin 1) 2) 3) 4) Normal? Myeloma? Hybridoma? Screen for desired monoclonal. MAbs in the Lab Macs extremely useful in molecular biology and medicine Applications Affinity columns Western blots ELISA (Enzyme Linked ImmunoSorbent Assay) Back The Future? Single Chain Antibodies Catalytic antibodies Bifunctional antibodies Etc. Features of a single-chain antibody (sFv). N VL VH Linker C The linker typically consists of a flexible/soluble peptide (for example, [GGGGS]6) CL CH1 CH2 CH3 CH1 CH2 CH3 CL Consists of the variable light (VL) chain of an antibody joined via a linker to the variable heavy (VH) domain. The sFv maintains the antigen binding specificity (but not always the affinity) of the parent antibody. Back