* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Current Approaches to Protein Purification Richard
Monoclonal antibody wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Signal transduction wikipedia , lookup
Size-exclusion chromatography wikipedia , lookup
Paracrine signalling wikipedia , lookup
Biosynthesis wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression wikipedia , lookup
Genetic code wikipedia , lookup
Gel electrophoresis wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Biochemistry wikipedia , lookup
Magnesium transporter wikipedia , lookup
Point mutation wikipedia , lookup
Expression vector wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Homology modeling wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Metalloprotein wikipedia , lookup
Interactome wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Proteolysis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
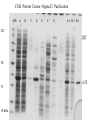
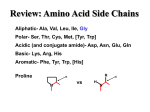
Protein Purification (from a lecture by Dr. Richard Burgess, University of Wisconsin, Madison, at the CSH protein purification course). Object: to separate a particular protein from all other proteins and cell components There are many proteins (over 4300 genes in E. coli) A given protein can be 0.001-20% of total protein Other components: nucleic acids, carbohydrates, lipids, small molecules Enzymes are found in different states and locations: soluble, insoluble, membrane bound, DNA bound, in organelles, cytoplasmic, periplasmic, nuclear Study Question You are given a shoe box full of an assortment of small objects including: Ping Pong balls Sugar cubes Paper clips 1/2” brass screws Iron filings 1. List the properties of each of these components that might help you fractionate them. 2. Devise the most efficient method you can for getting pure paper clips. 20 Naturallyoccurring Amino Acids Acidic: D, E, (C, Y) Basic: K, R, H Hydrophobic: I, L, V, W, F Polar: S, T, N, Q Other: G, A, M, P Overview of Protein Purification Types of Separations Protein Properties Protein Inactivation/Stabilization Protein Analysis and Purification Analytical Separations Gel-electrophoresis IEF 2D-gels Preparative Separations Various chromatographic methods Total E. coli Proteins - 2-Dimensional Gel Main Types of Molecular Interactions Hydrogen Bonds N H----N N-H + low temperature high temperature N H----O C donor acceptor N strength is very dependent on geometry and distance (2.6-3.1 A) Hydrophobic Interactions (waxy residues: Ileu, Leu, Val, Phe, Trp) HH HH H H H H H2O high salt high temperature low salt Ionic Interactions (charged residues:Asp- Glu- S- Lys+ Arg+ His+) +... low ionic strength Cl- + Na+ - high ionic strength Variables that Affect Molecular Forces Temperature Ionic strength Ion type Polarity of solvent (dielectric constant) pH Protein Properties - Handles for Fractionation Size (110 Da/amino acid residue) smallest most proteins largest Amino acids: 30 100 1,000 15,000 MW (kDa): 3.3 11 110 1,600 Multi-subunit complexes can contain 5-30 subunits Shape globular (sphere) asymmetric (cigar) Effects frictional properties, effective radius, movement through pores Centrifuge Gel filtration Sediments slower Elutes earlier Appears smaller Appears larger Protein Properties - Handles for Fractionation Net charge Ionizable group pKa pH2 pH7 pH12 C-terminal (COOH) Aspartate (COOH) Glutamate (COOH) Histidine (imidazole) N-terminal (amino) Cysteine (SH) Tyrosine (phenol) Lysine (amino) Arginine (guanido) 4.0 4.5 4.6 6.2 7.3 9.3 10.1 10.4 12.0 oooooooo---------------------------------------oooooooooo------------------------------------ooooooooooo-----------------------------------+++++++++++++oooooooooooooooooooo +++++++++++++++oooooooooooooooooo ooooooooooooooooooooooo----------------oooooooooooooooooooooooooo------------++++++++++++++++++++++++oooooooo ++++++++++++++++++++++++++++++o Isoelectric point pI = pH where protein has zero net charge Typical range of pI = 4-9 Charge distribution + - + - uniform - + - + + versus + + + clustered - - - - Protein Properties-Handles for Fractionation Hydrophobicity Hydrophobic residues usually are buried internally The number and distribution on the surface vary Can use Hydrophobic Interaction Chromatography H HH hydrophobic patch Solubility Varies from barely soluble (<mg/ml) to very soluble (>300 mg/ml) Varies with pH, ionic strength/type, polarity of solvent, temperature Least soluble at isoelectric point where there is least charge repulsion Ligand and metal binding Affinity for cofactors, substrates, effector molecules, metals, DNA When ligand is immobilized on a bead, you have an affinity bead Separation Processes that can be Used to Fractionate Proteins Separation Process Basis of Separation Precipitation ammonium sulfate polyethyleneimine (PEI) isoelectric solubility charge, size solubility, pI Chromatography gel filtration (SEC) size, shape ion exchange (IEX) charge, charge distribution hydrophobic interaction(HIC) hydrophobicity DNA affinity DNA binding site immunoaffinity (IAC) specific epitope chromatofocusing pI Electrophoresis gel electrophoresis (PAGE) charge, size, shape isoelectric focusing (IEF) pI Centrifugation Ultrafiltration sucrose gradient size shape, density ultrafiltration (UF) size, shape Typical Protein Purification Scheme Protein Inactivation/Stabilization Buffers Solution Components Protein Sources for Purification Traditional natural sources Bacteria, animal and plant tissue Cloning recombinant proteins into overexpression vector/host systems for intracellular production (E. coli the most used) In vitro protein synthesis Transcription/translation systems Total E. coli Proteins - 2-Dimensional Gel Determining the protein sequence from gel (proteomics) What You Can Learn from Amino Acid Sequence 1. Molecular weight of the polypeptide chain 2. Charge versus pH; Isoelectric point 3. Extinction coefficient 4. Hydrophobicity & membrane spanning regions 5. Potential modification sites 6. Conserved motifs that suggest cofactor affinity What You Can’t Learn from Amino Acid Sequence 1. Function 2. 3-Dimensional structure; Shape 3. Multi-subunit features 4. Ammonium sulfate precipitation properties 5. Surface features (hydrophobic patches, charge distribution, antigenic sites) Conclusion: Protein Purification is still very empirical! Engineering Proteins for Ease of Purification and Detection Once you have a gene cloned and can over-express the protein, you can alter protein to improve the ease of purification or detection You can fuse a tag to the N-or C- terminus of your protein You can decide to remove the tag or not Basic strategies Add signal sequence that causes secretion into culture medium Add protein that helps the protein refold and stay soluble Add sequence that aids in precipitation Add an affinity handle (by far the most used is the His-tag) Add sequence that aids in detection CSH Protein Course -Sigma32 Purification MW A B C D E F G A/3 B/3 D/3 225 bb’ 50 35 10 kDa s32