* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 6. Protein Folding
Genetic code wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Gene expression wikipedia , lookup
Biosynthesis wikipedia , lookup
Paracrine signalling wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Magnesium transporter wikipedia , lookup
Point mutation wikipedia , lookup
Expression vector wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Biochemistry wikipedia , lookup
Metalloprotein wikipedia , lookup
Interactome wikipedia , lookup
Homology modeling wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
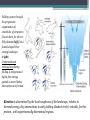
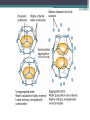
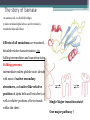
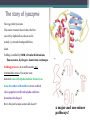
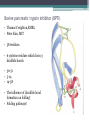
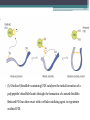
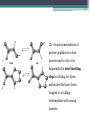
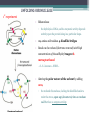
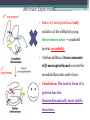
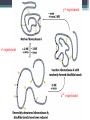
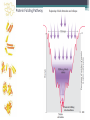
6. Folding and flexibility By Assist. Prof. Betul Akcesme • “Prediction of three dimensional structure of a protein from its amino acid sequence is the major unsolved problem in structural molecular biology” Introduction a polypeptide chain acquires its correct 3D structure to achieve the biologically active native state Some proteins folds spontaneously, others require assistance such as enzymes PROTEIN FOLDING ,chaperones) CHAPERONS 1.Binds to partially folded polypeptide prevents making unwanted bonding - promotes the folding 2. After a sequence acquires most of its correct secondary structure, it has a looser tertiary structure 3. FINAL native state MOLTEN GLOBULAR STATE Molten globular state to native state spontaneously How to predict the 3D structure from primary structure? Major unsolved problem native state protein is not static: secondary elements, domains undergo small movements in space: individual atoms or collectively; Functional activities of protein : ligand binding trigger conformation changes. Globular proteins are only marginally stable Denatured state (unfolded) Native state (folded) Energy difference: 5 – 15 kcal/mol (a single H-bond: 2 – 5 kcal/mol) Two major contributors to the energy difference: Enthalpy and Entropy Decrease for spontaneous process ENTHALPY energy of noncovalent interactions including the hydrophobic interactions, hydrogen bonds, and ionic bonds. These interactions are maximized to produce globular molecule in native state Covalent interactions ? These non‐covalent interactions are therefore stronger and more frequent in the native state and hence their energy contribution, enthalpy, is much larger. The enthalpy difference between native and denatured states can reach several hundred kcal/mol. ENTROPY native state is order, denatured states is disorder –each denatured structure is unique. entropy between the native ordered state and the denatured state can also reach several hundred kcal/mol but in the opposite direction to the enthalpy difference. Denatured state Native state FREE ENERGY difference between states are very small! • Fig.6.22 from Mathews, van Holde & Ahern, Biochemistry, 3rd ed. (2000): Thermodynamics of protein folding • summary of overall interplay between DH and the -TDS components (both hydrophobic effect and conformational entropy) • A simple reaction profile for two state reaction. The reaction can be described in terms entirely analogous to reaction kinetics where G is the conformational stability of the folded protein (GU − GF). For folding to occur the change in free energy must be negative The marginal stability of the native state is biologically and functionally important! Globular proteins in correct quantities at appropriate times. THEREFORE It is equally important to be able to degrade proteins as it is to be able to synthesize them. Globular proteins in living cells usually have a rapid turnover ,which means their native states is marginally stable. Some enzymes need structural flexibility, which would be inconsistent with a rigidly stabilized structure. Kinetic factors are important for folding “Calculation for searching all possible conformation in a random fashion to get the lowest energy is impossible!” Assuming each peptide has only three (α,β,L) conformation and time scale is one pico-second (10-12) (for a polypeptide chain of 150 amino acid residues) 3150 (=1068) possible conformations Time required is1068 x 10-12 = 1056 seconds (= 1048 years) cf) actual folding time: 0.1–1000 seconds both in vivo & in vitro Levinthal Paradox To occur on this short time scale, the folding process must be directed in some way through a “kinetic pathway of unstable intermediates ” to escape sampling a large number of irrelevant conformations Kinetic trapping in a wrong minimum Kinetic factors are important for folding process it is possible that the observed folded conformation is not the one with lowest free energy but rather the most stable of those conformations that are kinetically accessible! • Protein might be trapped in a local low energy state with a high energy barrier that prevents it from reaching the GLOBAL ENERGY MINIMUM. ( give different fold) • Structure prediction with energy calculation is (might) WRONG! HOW A LIVING CELL CAN PREVENT THE FOLDING PATHWAY FROM BECOMING BLOCK AT THE INTERMEDIATE STAGE? The most common obstacles to correct folding (1) Aggregation of the intermediates through exposed hydrophobic groups (2) Formation of incorrect disulfide bonds (3) Isomerization of proline residues Can be overcome by CHAPERONES! Needs “Special proteins that assist the folding protein – chaperone” • Energy minima in protein folding. • Anfinsen’s thermodynamic hypothesis suggests that the threedimensional structure of a native protein under physiological conditions is one in which the free energy of the system is the lowest (Anfinsen, C. B. Science 1973,181, 223–230). Thus, while proteins may sample other local minima during folding, there is only one true minimum-energy fold. Alternative way to remove kinetic barrier • α lytic protease (bacterial enzyme) • It is synthesized and folded in vivo but not active protein with prosegment of 77 residues This segment is excised after folding to PRODUCE ACTIVE ENZYME. Unfolded α lytic protease without this segment can not refold easily ADDING PROSEGMENT can induce refolding The capacity of folding exits in unfolded enzyme BUT there is a BARRIER present in folding pathway. PROSEGMENT REMOVES THIS KINETIC BARRIER by lowering the free energy of transition state for folding ( like enzyme catalyzes chemical reactions) Molten globules are intermediates in folding driving force not very known BUT • packing of core not formed •Interior side chains are mobile •Loops and surface elements remain unfolded MOLTEN GLOBULE UNFOLDED single native form is reached, forming native interactions, including hydrophobic packing in the interior & fixing surface loops FOLDED Up to 1 second Within a few milliseconds also some native like alpha helix and beta strand positions Ensemble of conformations are reduced Folding occurs through the progressive organization of ensembles of structures [shown here for the srcSH3 domain (left)] on a funnel-shaped Free energy landscape (right). Conformational entropy loss during folding is compensated by the free energy gained as more Native interactions are formed. Kinetics is determined by the local roughness of the landscape, relative to thermal energy. Key interactions in early folding (dashed circle) coincide, for this protein, with experimentally determined regions. Burying hydrophobic side chains is a key event Main mystery of protein folding is… …The collapse of the unfolded state to generate the molten globule embodies • WHAT IS THE DRIVING FORCE BEHIND THE CHOICE OF NATIVE TERTIARY FOLD FROM A RANDOMLY ORIENTED POLPEPTIDE CAHIN? 1. Secondary structure formation cannot be the thermodynamic driving force of protein folding… BECAUSE stable hydrogen bonds can also be formed to water molecules in the unfolded state. (There is very little change in Free Energy) 2. Large Free energy change by bringing hydrophobic side chains out of contact with water and into contact with each other in the interior of a globular entity. Most likely scenario that the polypeptide chain begins to form compact shape with Hydrophobic side chains partially buried very early in the folding….. • This scenario has several important consequences ▫ Thus it vastly reduces the number of possible conformations that need to be searched because only those that are sterically accessible within this shape can be sampled. ▫ Furthermore, when side chains are buried, their polar backbone –NH and –CO groups are also buried in a hydrophobic environment, hence unable to form hydrogen bonds to water – BUT they bond to each other – so you get alpha and beta structures. CONCLUSION Secondary structure elements NOT DRIVING FORCE for the formation of Molten Globule But they are consequences of burying hydrophobic side chains… Core Questions • Looking at the protein sequences of globular proteins, one finds that hydrophobic side chains are usually scattered along the entire sequence, seemingly randomly. • In the native state of folded protein, ½ of these side chains are buried, and the rest are scattered on the surface of the protein, surrounded by hydrophilic side chains. • The buried hydrophobic side chains are not clustered in the sequence. • What causes these residues to be selectively buried during the early and rapid formation of the molten globule? • If we wish to solve the folding problem, this must be answered first. Both single and multiple folding pathways have been observed The unfolded state consists of many rapidly interconverting conformationally different molecules, U1...Un. High energy unfolded state Low energy folded state The molten globule is an ensemble of structurally related molecules, M1...Mm, which are rapidly interconverting and which slowly change to a single unique conformation, the folded state F. The conversion from the molten globule state to the folded state is slow and passes through a high energy transition state, T. Both single and multiple folding pathways have been observed • Ui‘s --- unfolded states, many of them. • Mi’s --- molten globule states. Has most secondary structures, but less compact. • Converging to F. During this relatively slower process it passes a high energy transition state T. • These facts have been verified by NMR, hydrogen exchange, spectroscopy, and thermo-chemistry. • • Alan Fersht, Cambridge • The techniques is based on investigation of the effects on the energetics of folding of single mutation in a protein known structure EXAMPLE Mutation ALA to GLY in solvent exposed face at an alpha helix , the helix is already formed in the intermediate state both destabilize both intermediate state and the native state, as well as transition state On the other hand, mutations destabilize the native state but do not affect the energy of the intermediate or transition state at all the helix is formed after the transition state. The story of barnase 110 amino acids, no disulfide bridges. 3 amino terminal alpha helices, and C terminal 5stranded antiparallel sheet. Effects of all mutations are examined, detailed residue characterization : its folding intermediates and transition states Folding process: intermediate molten globule state: already with most of native secondary structures, and native like relative position of alpha helix and beta sheet, as well as relative positions of beta strands within the sheet. Single Major transition state! One major pathway ! The story of lysozyme Hen egg-white lysozyme The native structure has 2 lobes, the first one with 5 alpha helices, the second is mainly a 3-stranded antiparallel beta sheet. Folding is studied by NMR, Circular dichronism, fluorescence, hydrogen- deuterium exchanges Folding process: At 20 milliseconds, two intermediate states of lysozyme were detected: one with alpha domain formed, no beta, the other with neither; in fact a third (less popular) with both alpha and beta domains developed. But is this just because some folds slower? 2 major and one minor pathways! The most common obstacles to correct folding (1) Aggregation of the intermediates through exposed hydrophobic groups (2) Formation of incorrect disulfide bonds (3) Isomerization of proline residues Can be overcome by CHAPERONES! Needs “Special proteins that assist the folding protein – chaperone” • ENZYMES THAT HAVE ROLE IN FOLDING! Enzymes assists formation of proper disulfide bonds during folding • Unfold proteins have no disulfide bonds. • Formation of disulfide bonds require enzyme to help (oxidation of cysteine residues). ▫ In bacteria, periplasmic space Catalyzed by disulfide bridge forming enzyme (Dsb) ▫ In eukaryotes, disulfide bond formation occurs in E.R. before transport to cell surface ▫ (not found in cytosol because of reducing environment but IN membrane or secreted) Protein disulfide isomerase(PDI) Catalyze internal disulfide exchange to remove folding intermediates with incorrectly formed disulfide bridge. Bovine pancreatic trypsin inhibitor (BPTI) • Thomas Creighton,EMBL • Peter Kim, MIT • 58 residues. • 6 cysteine residues which form 3 disulfide bonds. • 30-51 • 5-14 • 14-38 • The influence of disulfide bond formation on folding! • Folding pathways! Bovine pancreatic trypsin inhibitor (BPTI) • Trapping disulfide bonded intermediates as a method for studying the folding pathways. First bond randomly form … with 30-51 being more stable, 60%, Second bond (if the first is 30-51): 5-14, 5-38, 14-38 all possible. 14-38 native, others unfold. Formation of 5-55 occurs very slowly- buried inside the folded intermediate and not accessible to oxidizing agents • Adding the enzyme protein disulfide isomerase significantly increases the rate of folding of protein BPTI. • Prokaryote: Dsb (similar to thioredoxin) • Eukaryote: PDI (Protein disulfide isomerase) • Reduced (SH-containing) PDI catalyzes the rearrangement of a polypeptide’s non-native disulfide bonds via disulfide interchange reactions to yield native disulfide bonds. • (b) Oxidized (disulfide-containing) PDI catalyzes the initial formation of a polypeptide’s disulfide bonds through the formation of a mixed disulfide. Reduced PDI can then react with a cellular oxidizing agent to regenerate oxidized PDI. Isomerization of proline residues • Trans‐peptide with C=O and –NH groups pointing in opposite directions was 1000 times preferred to the cis‐peptide.(C=O and –NH groups pointing in same directions) • However, when the 2ndresidue is a proline the cis conformer is only 4 times less stable. cis=-proline peotide in many proteins • Mainly the Pro trans conformer,(few steric collusions) but the cis version is found in tight turns and are sometimes essential for conformational flexibility. • In the native protein the cis‐proline arrangements are stabilized by tertiary structure interactions, but in the unfolded state there is an equilibrium between cis/trans isomers. • When refolding occurs, you can get the proline‐peptide bond in the wrong form. From a kinetic standpoint, cis‐trans proline isomerization is a very slow process that can prevent the progress of protein folding by trapping one or more proline residues crucial for folding in the non‐native isomer, especially when the native protein requires the cis isomer. • More prolines—more chance. As noted…cis‐trans isomerization is a slow process and in vitro is often the rate limiting folding step. •NEED to get things in right conformations! • Cis –trans isomerization of proline peptides is a slow process and in vitro it is frequently the rate limiting step in folding for those molecules that have been trapped in a folding intermediate with wrong isomers. • In vivo, rates of this process are enhanced by enzymes initially called peptidyl prolyl isomerases. • First one found cyclophilin • *impacts the rate of cis‐trans isomerization of proline peptides by a factor of a million over non enzmatic reaction Proline containing tetrapeptide Invloved in immunosupression by inhibiting T cell proliferation after binding immunosupressive drugs ( unrelated to their isomerase activity) Graft rejection! (involved in immunosupression) The active site is on the outside of beta barrel 1) CAN A PROTEIN FOLD SPONTANEOUSLY TO ITS NATIVE STATE? YES ! Anfinsen's dogma (also known as the thermodynamic hypothesis) is a postulate in molecular biology championed by the Nobel Prize winner Christian B. Anfinsen. 2) DOES A PROTEIN SEARCH ALL POSSIBLE OUTCOMES TO FIND THE FINAL CONFORMATION , THE ONE OF LOWEST FREE ENERGY? NO! How the protein reaches this structure is really the subject of the field of protein folding, which has a related dogma called Levinthal's paradox. The Levinthal paradox states that the number of possible conformations available to a given protein is astronomically large, such that even a small protein of 100 residues would require more time than the universe has existed to explore all possible conformations and choose the appropriate one, it would also arguably make computational prediction of protein structures under the same basis unfeasible if not impossible. Anfinsen Experiment Protein Folding • Protein folding considers the question of how the process of protein folding occurs, i. e. • unfolded native state. • Importance: • – Predict 3D structure from primary sequence • – Avoid misfolding related to human diseases • – Design proteins with novel functions ANFINSEN: AMINO ACID SEQUENCE DETERMINES PROTEIN SHAPE The dogma states that, at least for small globular proteins, the native structure is determined only by the protein's amino acid sequence. At the environmental conditions (temperature, solvent concentration and composition, etc.) at which folding occurs, the native structure is a unique, stable and kinetically accessible minimum of the free energy. Hypothesis “protein amino acid sequence determines the final shape a protein assumes in a water solution” UNFOLDING RIBONUCLEASE 1st experiment • Ribonuclease ▫ the hydrolysis of RNA, and its enzymatic activity depends entirely upon the protein being in a particular shape. • 124‐amino acid residues. 4 disulfide bridges • Bonds can be reduced (electrons removed) with high concentrations of the sulfhydryl reagent bmercaptoethanol –S—S—becomes –SHHS— • Altering the polar nature of the solvent by adding urea, ▫ the reduced ribonuclease, lacking the disulfide bonds to resist the stress, open up (denatures) into a random coil that has no enzyme activity. 1st experiment When Anfinsen did this, he observed that the ribonuclease protein slowly regained its enzymatic activity. • Free of the reducing agent, the sulfhydryl groups (--SH) of the cysteines were being oxidized by dissolved oxygen from the air, and the protein was refolding into the catalytically active shape. • This could only mean that the folding was indeed directed by the amino acid sequence Anfinsen Experiment 2nd experiment • Remove β-mercaptoethanol only, oxidation of the sulfhydryl group, then remove urea → scrambled protein, no activity • Further addition of trace amounts 3rd experiment of β-mercaptoethanol converts the scrambled form into native form. • Conclusion: The native form of a protein has the thermodynamically most stable structure. 3rd experiment 1st experiment 2nd experiment Conclusion • The original work led Anfinsen to propose his "Thermodynamic Hypothesis", which states that the native conformation of a protein is adopted spontaneously. • In other words, there is sufficient information contained in the protein sequence to guarantee correct folding from any of a large number of unfolded states. • This demonstrated that the shape that a protein realizes in solution is dictated by amino acid sequence information, which is expressed in terms of thermodynamic stability Protein Folding Pathway