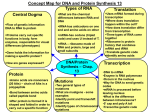
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Section 8: Genetic Mutations, Ribosome Structure
Cre-Lox recombination wikipedia , lookup
Community fingerprinting wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Protein structure prediction wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Non-coding RNA wikipedia , lookup
Messenger RNA wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Gene expression wikipedia , lookup
Expanded genetic code wikipedia , lookup
List of types of proteins wikipedia , lookup
Bottromycin wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Molecular evolution wikipedia , lookup
Epitranscriptome wikipedia , lookup
Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline TOPICS • • • Ribosome structure Classes of mutation Mechanism of action of the tetracyclines SUMMARY Up to this point, our focus has been on antibiotics that exert prokaryotic specificity by targeting molecules that are not present in eukaryotes (or in the case of gramicidin, no robust prokaryotic specificity is exerted). The RNA polymerase and protein synthesis inhibitors highlight examples of antibiotics that target prokaryotic-specific regions of conserved molecules present in both prokaryotes and eukaryotes. We primed students for the upcoming discussion upon introduction of trimethoprim. Now we elaborate on the effect of mutations on structure. To understand the mechanism of action of the translational inhibitors, such as the tetracycline class of antibiotics, students must understand the consequences of mutations on RNA, how these mutations can affect amino acid composition, the effects of those changes on structure and how structure relates to function. We show that even subtle differences in structure between prokaryote and eukaryote versions of the ribosome can result in altered binding affinity of antibiotic for target. While we do not specifically discuss the transcriptional inhibitor rifampicin, it too targets a prokaryotic region of a conserved molecule (RNA polymerase). LEARNING GOALS • • • • • Explain the molecular mechanism by which translational inhibitors kill bacteria. Indicate the key molecules that interact with the ribosome and the function of each interaction. Provide a molecular explanation for how the transcriptional and translational inhibitors achieve prokaryotic specificity. Compare a silent, nonsense and missense mutation in terms of DNA alteration, result on mRNA structure and polypeptide structure and function. Explain how point mutations in a gene might result in a functional, yet structurally altered gene product. PRE-CLASS PREPARATION Students should read about or review the mechanistic process of translation, classes of mutation, DNA replication, and nucleic acid synthesis. Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline PRE-CLASS ASSESSMENT 1. During nucleic acid synthesis, the new, incoming nucleotide is always added to the: A. 5’ phosphate B. 5’ –OH C. 3’ phosphate D. 3’ –OH Nucleic acids are always synthesized in the 5’ to 3’ direction. 2. Which of the following mutations would be MOST likely to have a harmful effect on an organism? A. A base-pair substitution in the middle of the coding sequence. B. A deletion of three nucleotides in the middle of the coding sequence. C. A single nucleotide deletion in the middle of an intron. D. A single nucleotide deletion near the end of the coding sequence. E. A single nucleotide insertion just after the start of the coding sequence. Of course we can only determine the answer empirically, but we can speculate based on knowledge of information flow in the cell. In all cases, we are assuming that a polypeptide is the gene product. One might ask the students if and how their answers would change if the gene product were an RNA. A. A single base-pair substitution would keep the reading frame in tact, so the mutation would likely be less harmful than a mutation that completely alters the reading frame or truncates or eliminates the polypeptide. B. While more nucleotides are removed than in (A), this change would also keep the reading frame intact. C. The introns don’t contain coding sequence, so this would not be likely to have an adverse effect unless the sequence was regulatory in nature. D. This change would cause a frameshift, but since it occurs at the end of the coding region, the effect is probably less severe than the deletion in the middle of the coding sequence (fewer amino acids affected). E. This is likely to be the most severe because it would alter the reading frame of the entire protein, resulting in the production of all the wrong amino acids. 3. Choose the correct statement: A. Mutations in the DNA are sometimes beneficial to the individual and the population. B. Mutations in the DNA are sometimes beneficial to the individual, but always harmful to the population. C. Mutations in the DNA are always harmful to the individual but sometimes beneficial to the population. D. Mutations in the DNA are always harmful to both the individual and the population. Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline This question assesses students understanding of the fact that mutations are not always harmful and that they drive evolution. Students may not be able to answer this question correctly prior to class, but if an unacceptable number of students answers incorrectly, instructors can remind students of the question prior to class and indicate that it will be asked again at the end of class. GUIDE TO THE POWERPOINT SLIDES Outline • • • • • • • Introduction/active Learning Ribosome structure Tetracycline binding and prokaryotic specificity Acquisition of genetic mutations Classes of genetic mutations Effect of mutation on structure and function Review of transcription vs. translation Introduction/active Learning GUIDING QUESTIONS • What interactions might be blocked by tetracycline interacting with its target? • Is the target molecule present in prokaryotes and eukaryotes? Active Learning Activity type: Shout-out Tetracycline kills bacteria by inhibiting translation. Students are shown a schematic figure of the ribosome and other molecules used in translation with the following question: • What interactions might be blocked by tetracycline interacting with its target? This question is designed to encourage students to consider the mechanism of translation in greater detail than they have up to this point by asking what inter-molecular interactions are critical the process and, therefore, would be targets for disruption by tetracycline. Consider?what?interac=ons?might?be?blocked?by? tetracycline?interac=ng?with?its?target? to h"p://upload.wikimedia.org/wikipedia/commons/b/b1/Ribosome_mRNA_transla=on_en.svg? Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline There are a variety of different answers that are possible. Some potential answers include: • mRNA interacts with (binds to) ribosome (30S subunit-5’ mRNA recognition) • 30S interacts with AUG start site of mRNA • 30S and 50S interact • tRNA interacts with 70S ribosome (A site = aminoacyl tRNA entry site) • Anticodon of tRNA binds to codon of mRNA • Peptide bond formation catalyzes in the active site of the ribosome (=P site peptidyltransferase) • Spent (uncharged) tRNAs dissociate/exit (E site) • Tetracycline;binds;to;the;30S;(small);subunit;nucleoBdes; through;nonCcovalent;bonding;interacBons; • Growing polypeptide chain needs to be released • The;binding;alters;the;structure;of;the;small;subunit;such;that;; the;tRNA;anBcodon;and;mRNA;codon;interacBons;are; (exit tunnel) obscured; Actual answer: Tetracycline binds to the 30S (small) subunit nucleotides through non-covalent bonding interactions The binding alters the structure of the small subunit such that the tRNA anticodon and mRNA codon interactions are obscured h"p://faculty.ccbcmd.edu/courses/bio141/ lecguide/unit2/control/tetres.html; Ribosome structure We provide structural slides from the literature to illustrate the tertiary structure of ribosomes as it pertains to function, and the very specific noncovalent binding interactions required to achieve tetracycline binding. The goal is for students to connect how molecular structure determines conformation and function. This%figure%shows%the%general%shape%of%the%whole%ribosome% Just as polypeptides use non-covalent bonding interactions to adopt functional folded structure (conformation), RNA molecules can also exert intra-molecular non-covalent bonding, primarily in the form of hydrogen bonds between complementary base pairs. The rRNA associates with the small polypeptides through non-covalent bonding interactions. This “folded” structure of the rRNA plus polypeptides confers the shape and function of the molecule. The large and small subunit associate only in the presence of mRNA which passes through a “tunnel” created by the mature ribosome. If:we:zoom:in:on:the:RNA:porBon,:we:see:that: individual:nucleoBdes:undergo:baseFpairing:to:form: “stems”:and:“loops”: • The$large$and$small$subunit$associate$only$in$the$presence$of$mRNA$ • The$mRNA$passes$through$a$“tunnel”$created$by$the$mature$ribosome$ • This$tunnel$contains$the$ac;ve$A,$P,$and$E$sites$where$charged$tRNA$ molecules$enter$(A$site),$amino$acids$are$transferred$to$the$growing$ chain$(P$site)$and$uncharged$tRNAs$exit$(E$site)$ Protein$synthesis$inhibitor$ an;bio;cs,$such$as$tetracycline,$ bind$to$the$30S$or$50S$ ribosomal$subunit.$$ Where$on$these$subunits$do$ you$think$they$bind?$ h"p://rna.ucsc.edu/rnacenter/images/figs/ecoli_23s.jpg: Small World Initiative ICell nstructor Guide 1146 Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline This tunnel contains the active A, P, and E sites where charged tRNA molecules enter (A site), amino acids are transferred to the growing chain (P site) and uncharged tRNAs exit (E site). ell 146 & The&primary&binding&site&of&tetracycline&(yellow)&is&to& the&A&site&of&the&ribosome& Green,&blue&and&cyan&represent& relevant&helices&of&the&16S&rRNA& Model&depic*ng&possible&tetracycline&binding& interac*ons&with&16S&rRNA&& Cell 1146 The&point&here&is¬&to&memorize&this&specific&example,&but&to&use&it&as& context&to&understand:& • The&role&of&nonDcovalent&bonding&interac*ons&in&biology& • The&rela*onship&of&structure&to&func*on& • How&the&ribosome&func*ons&at&the&molecular&level& tetracycline& From&Broderson&(2000)&Cell&103:1143N&1154& h<p://commons.wikimedia.org/wiki/ File:Tetracycline_structure.svg& tetracycline& Broderson&(2000)&Cell&103:1143D&1154& At this point, students should be able to meet the learning goal: ü Indicate the key molecules that interact with the ribosome and the function of each interaction. Tetracycline binding and prokaryotic specificity GUIDING QUESTION: • How does tetracycline achieve prokaryotic specificity if it targets a molecule conserved in both prokaryotes and eukaryotes? Transcription, translation and DNA synthesis are highly conserved processes, present across phylogeny. We ask, how then, do antibiotics that target these functions achieve prokaryotic specificity? The answer lies in the fact that there are slight differences in the structure of the targets; however, while there are structural differences between prokaryotic and eukaryotic targets, function of the target molecule is retained. We begin by presenting the DNA differences between the prokaryotic and eukaryotic ribosome sequences that alter ribosome structure to the extent that tetracycline binding specificity is affected without affecting function. These special DNA mutations are what allow utility of antibiotics that target functions that are conserved in both eukaryotes and prokaryotes. The transcriptional inhibitor rifampicin utilizes this strategy as well, although we do not include slides depicting its mechanism of action. Note that the original paper for discussion listed in Section 7 (Fujii et al. (1995) provides an opportunity to discuss the mechanism of action of rifampicin. Active Learning Activity type: Think-Pair-Share Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline We use an example from the literature to illustrate that tetracycline achieves its prokaryotic specificity due to a higher binding affinity for the prokaryotic ribosome structure. Interpret'this'graph'' Question: Interpret this graph (see image on slide) [3H]Tet'refers'to' tetracycline'in'which' hydrogen'is'replaced' with'the'radioac9ve' [3H]'form.'' How#does#the#informa,on#gained#in#this#experiment#differ# from#that#in#the#previous#experiment?# In#vitro# transla,on#with# purified# ribosomes#and# poly#U#RNA# template# 100%# corresponds#to# number#of#phe# amino#acids# incorporated# with#no# tetracycline# present.## Cell 1146 V1* V2* V3* V4* V5* V6* V7* 1243* 1294* Cell 1146 1117* Tetracycline*binding* 1054* 1197D1198* Numbers*refer*to*single* nucleo3de*posi3ons*in*the* 16S*rRNA* 986* The$process$of$transla/on$is$conserved$between$prokaryotes$ and$eukaryotes;$key$nucleo/de$differences$in$ribosomal$RNA$ retain$func/on$of$the$ribosome,$but$create$differen/al$ binding$opportuni/es$for$an/bio/cs$ Budkevich T V et al. Nucl. Acids Res. 2008;36:4736-4744 1173* The second slide measures ribosome function in the presence of tetracycline. An in vitro translation assay was performed using a polyU RNA template together with purified ribosomes to produce a polyphenylalanine peptide (codon UUU). Prokaryotic ribosome function decreases markedly in the presence of tetracycline. Budkevich T V et al. Nucl. Acids Res. 2008;36:4736-4744 1043* An in vitro binding assay was performed between a radioactive ([3H]) form of tetracycline and either purified prokaryotic (70S) or eukaryotic (80S) ribosomes. The data show higher levels of tetracycline bound to the prokaryotic (70S) ribosome compared to the amount of tetracycline bound to the eukaryotic (80S) ribosome. V8* V9* V10* tetracycline* Broderson*(2000)*Cell*103:1143D*1154* Lafontaine$and$Tollervey$(2001)Nature)Reviews)Molecular)Cell)Biology)2,$514H520$ In the slide set, we elaborate on nucleotide differences between the eukaryotic and prokaryotic rRNA sequences. These nucleotide differences are unique in that they lead to slightly different structures without altering overall functional capability. Even subtle differences in structure can result in altered binding affinity of antibiotic for target. Transcrip)onal,inhibitors,exploit,subtle, differences,in,structure,between,prokaryo)c, and,eukaryo)c,RNA,polymerase,structure,, Image,from,Centre,d’études,d’agents,Pathogènes,et,Biotechnologies,pour,la,Santé, Montpellier*+*France,, h<p://www.cpbs.cnrs.fr/spip.php?ar)cle81&lang=fr, Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline This information also becomes relevant for our discussion of molecular phylogeny in Section 9 and will be relevant to students when they characterize the identity of their isolates using 16S PCR amplification, sequencing and BLAST comparisons. At this point, students should be able to meet the learning goal: ü Provide a molecular explanation for how the transcriptional and translational inhibitors achieve prokaryotic specificity. Acquisition of genetic mutations GUIDING QUESTION: • How do these nucleotide changes arise and why are they maintained? Presentation of ribosomal structural detail provides an excellent segue into the topic of nucleotide changes and their potential effect on conformation and function. DNA$polymerase$makes$an$occasional$ mistake$ 3’ C 5’ G T G CT AAC 5’ GTGAAT GG C 3’ CACTTACC G T AA G C C G 3’ 5’ Briefly, we present slides depicting how mutations in DNA replication are transmitted to subsequent generations of cells and how these mutations can affect polypeptide structure after translation. For example, a single nucleotide change can result in serine substitution in place an alanine. We do not discuss the mechanism of DNA replication; for those wishing to do so, this would be an idea placement of that topic. A"serine"residue"is"subs+tuted"for"alanine" DNA" 5’ ATG GCT TGC 3’ 3’ TAC CGA ACG 5’ transcrip+on" 5’ AUGGCUUGC 3’ transla+on" N Met–Ala–Cys C 5’ ATG TCT TGC 3’ 3’ TAC AGA ACG 5’ transcrip+on" 5’ AUGUCUUGC 3’ transla+on" N Met–Ser–Cys C • Are)the)proper7es)of)alanine)and)serine)similar?)) • Do)you)think)that)this)single)amino)acid)subs7tu7on)will) affect)the)conforma7on)of)the)polypep7de?) • If)so,)how?)If)not,)why?) • If)conforma7on)is)altered,)will)func7on)be)altered?) of Alanine) Serine) WikiCommons)user:)Borb) Classes of genetic mutations We provide schematic slides to illustrate the different classes of mutation: point mutations, deletions, and insertions. We also discuss the different classes of point mutations (normal, silent, nonsense, and missense) and their potential outcomes. It is also important to stress the key concept that point mutations drive evolution. Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline We can use a three-letter word analogy to consider the effects of some DNA mutations on amino acid sequence HIT THE BAT Normal “polypeptide” sequence HIT CTH EBA T Single nucleotide insertion RESULT: “frameshift” of codon usage Classes&of&point&muta/ons& NORMAL& SILENT& NONSENSE& MISSENSE& TGC ACG TGT ACA TGA ACT TGG ACC transcrip/on& UGC transla/on& HIT CAT THE BAT Three nucleotide insertion RESULT: single amino acid inserted; codon usage remains “in frame” HIT CHE BAT Single nucletide substitution (point mutation) RESULT: silent, missense or nonsense (see next slide) Cys Correct& amino&acid& transcrip/on& UGU transla/on& Cys No&change& in&amino& acid& sequence& transcrip/on& transcrip/on& UGA UGG transla/on& transla/on& STOP Premature& STOP&codon& introduced& Trp Single&amino& acid&change& Effect of mutation on structure and function Examina'on)of)Red)Blood)Cells) )Sickle)Cell)Anemia)) )Normal)) ) 2012)Northeast)Regional)Summer)Ins'tute)) ) ) We have inserted a modified version of a “teachable tidbit” from the National Academies Northeast Regional Summer Institute of 2012 to demonstrate the effects of a single amino acid change on hemoglobin in the disease sickle cell anemia. While this deviates from our theme of antibiotics, it is a well-studied example that students usually find interesting. For an additional example involving resistance to the antibiotic ciprofloxacin, refer to the discussion paper packet Number 7 (also found at the end of Section 10). More information on the sickle cell anemia “tidbit” can be found at the Yale Center for Scientific Teaching website: http://cst.yale.edu/biology-and-chemistry-interface-tidbits Active Learning Activity type: Think-Pair-Share and electronic response Think-Pair-Share: In the first activity, students use their knowledge regarding polarity to determine which amino acid side chains are hydrophilic and which are hydrophobic. ThinkEPairEShare( 1.(Which(amino(acid(side(chains((R(groups)(are(hydrophobic?( 2.(Which(amino(acid(side(chains((R(groups)(are(hydrophilic?( Valine((Val,(V)(( Glutamate((Glu,(E)(( Serine((Ser,(S)( Leucine((Leu,(L)( Lysine((Lys,(K)(( Threonine((Thr,(T)( 2012(Northeast(Regional(Summer(InsJtute(( ( Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline We then ask students to consider: What percentage of the 574 amino acids in hemoglobin would have to be changed to cause sickle cell anemia? Even if students are familiar with this example, it is dramatic to consider (or reconsider) the large implications for such a seemingly small change. Note that there are two β subunits that compose the mature hemoglobin molecule, so there are actually two amino acid changes per hemoglobin. Each%blue% dot=one%amino% acid% α=141%amino%acids% % β=146%amino%acids% β% α% α% β% Approximately%what%percentage%of%the%574%amino%acids%in%hemoglobin% do%you%think%you%would%need%to%change%to%cause%sickle%cell%anemia?% 2012%Northeast%Regional%Summer%InsItute%% % Electronic Response: Next, analogous to the question posed previously about where tetracycline might be expected to bind to the ribosome, we ask: Where in this folded protein might this change have occurred? A. Near the oxygen binding site B. At the interface between subunits C. On the surface Remember:(An(external(hydrophilic(amino(acid(is( replaced(with(a(hydrophobic(one(( If the comparison were totally analogous to the tetracycline/ribosome example, we might expect the amino acid change to occur where the different subunits interact, but in this case, the change is on the surface on the molecule. Because a hydrophilic amino acid is exchanged for a hydrophobic one, this region of the surface no longer interacts with water and preferentially interacts with other altered hemoglobin molecules. We indicate to the students that the answer must be determined through experimental analyses and that all answers would be logical predictions. Glutamate( (hydrophilic)( Valine( (hydrophobic)( 2012(Northeast(Regional(Summer(InsCtute(( ( At this point, students should be able to meet the learning goals: ü Explain the molecular mechanism by which transcriptional and translational inhibitors kill bacteria. ü Compare a silent, nonsense, and missense mutation in terms of DNA alteration, result on mRNA structure, and result on polypeptide structure and function. ü Explain how point mutations in a gene might result in a functional, yet structurally different gene product. Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline Review transcription vs. translation Active Learning Activity type: Think-Pair-Share We close with an activity designed to highlight the effects of mutation in RNA synthesis compared to DNA synthesis and the implications of DNA replication errors for prokaryotes compared to eukaryotes. Question: Consider the effects of a nucleotide synthesis error that results in complete loss of protein function and that the protein is required for life of the cell. • First, consider the effect on an individual prokaryotic cell if the mutation was introduced by RNA polymerase in transcription. • Next, consider the effect on an individual prokaryotic cell if the mutation was introduced by DNA polymerase in DNA replication. Students tend to think of RNA synthesis as a single reaction, yet transcription occurs continuously from a single gene, such that the effects of a single mutation in RNA synthesis are diluted. We use a visual representation to highlight this fact. On the other hand, an error in DNA replication will affect all gene products in a bacterial cell (and half of those in a eukaryotic cell since there are two copies of each gene). With%an%error%in%DNA%replica1on% Compare(the(effects(of(an(error(in(transcrip1on.(.(.( Modified from Figure 7-2 Essential Cell Biology (© Garland Science 2010) Modified from Figure 7-2 Essential Cell Biology (© Garland Science 2010) In addition, mutations in RNA are not heritable. Errors in DNA synthesis, on the other hand have dire consequences for the cell. In multicellular organisms, defects must occur in germline (sex) cells to be heritable to the next generation (but the error can be propagated to subsequent daughter cells, leading to defects in development or cancer). Mammals have two copies of each chromosome (diploid), and only one is acquired by each gamete. Therefore, a particular mutation in a eukaryotic cell may not be inherited. All DNA mutations in bacteria are heritable; they only have one chromosome and each cell division creates offspring. Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline Summary Statement: Antibiotics must target essential (conserved) functions, yet they must be prokaryotespecific to be most useful. Specificity is achieved in two main ways: • Antibiotic targets a molecule that is essential for prokaryotes but not for eukaryotes. • Antibiotic targets a structural change in a conserved molecule. The structural difference is the result of nucleotide differences between prokaryote and eukaryote DNA that alters structure of the gene product but retains function. POST-CLASS ASSESSMENT 1. What is the most likely effect of a mutation in the DNA that inserts a three nucleotide stop codon into the beginning of the coding sequence of a gene? A. The polypeptide and the mRNA will be shorter than usual. B. The polypeptide will be shorter than usual but the mRNA will not be affected. C. The polypeptide will be unaffected, but the mRNA will be shorter than usual. D. In eukaryotes, only the polypeptide will be affected, but in prokaryotes the mRNA may also be affected if several genes are under the control of the same promoter. The transcript, in general, is unaffected by codon changes. 2. Consider a bacterial operon that contains the coding sequence for three genes, (A, B, C), all under the control of a single promoter where A is the most upstream gene followed by B then C. How will a three nucleotide insertion in the most conserved region of the promoter most likely affect expression of gene C? A. Protein C will probably not be produced. B. The gene C mRNA will be produced but Protein C will be non-functional. C. The gene C mRNA will be produced but Protein C will not be produced. D. The expression of gene C will not be affected. Because the genes are arranged in an operon, they are all under control of the same promoter. Insertion of three nucleotides at the conserved region of the promoter will likely negatively affect RNA polymerase holoenzyme binding, leading to loss of expression of the entire operon (no mRNA, no protein). Small World Initiative Instructor Guide Section 8: Genetic Mutations, Ribosome Structure, and Tetracycline ORIGINAL RESEARCH PAPERS FOR DISCUSSION The set of papers below give students a glimpse into the incremental nature in which scientific models are tested and functions are elucidated. We illustrate the process of discovery of the mechanism of action of tetracycline. We suggest that instructors focus on one or a few key figures from each paper. The first paper in the series is from 1953 (the same year the structure of DNA was published) demonstrating that aureomycin (the first tetracycline identified) affects protein synthesis. The second paper demonstrates binding to the 30S subunit of the ribosome while the third paper shows binding to ribosomes in a cell-free system to inhibit protein synthesis. The fourth and fifth papers work to elucidate the specific region to which tetracycline binds. The final paper determines specific tetracycline binding sites using atomic structure data and predicts the mechanism of action based on these data. Gale, E. and Folkes, J. (1953) The Assimilation of Amino-Acids by Bacteria. Biochem. 53:493-498. Connamacher, R. and Mandel, G. (1965) Binding of Tetracycline to the 30S Ribosomes and to Polyuridylic Acid. Biochemical and Biophysical Research Communications 20:98103. Day, L. (1966) Tetracycline Inhibition of Cell-Free Protein Synthesis. J. Bact. 91:19171923. Gottesman, M. (1967) Reaction of Ribosome-bound Peptidyl Transfer Ribonucleic Acid with Aminoacyl Transfer Ribonucleic Acid or Puromycin. J. Biol. Chem. 242:5564-5571. Goldman, R., Hasan, T., Hall, C., Strycharz, W., and Cooperman, B. (1983) Photoincorporation of Tetracycline Into Escherichia coli Ribosomes. Identification of the Major Proteins Photolabeled by Native Tetracycline and Tetracycline Photoproducts and Implications for the Inhbitory Action of Tetracycline on Protein Synthesis. Biochem. 22:359-368. Brodersen, D., Clemons, W., Carter, A., Morgan-Warren, R., Wimberly, B., and Ramakrishnan, V. (2000) The Structural Basis for the Action of the Antibiotics Tetracycline, Pactamycin, and Hygromycin B on the 30S Ribosomal Subunit. Cell 103:1143-1154. RESOURCES Budkevich et al. (2008) Features of 80S mammalian ribosome and its subunits. Nucleic Acids Res. 36:4736-4744. Tetracycline binding to ribosome experiment. Pray, L. (2008) DNA replication and causes of mutation. Nature Education 1:214. DNA replication and causes of mutation.