Attachment 2.2 Sequencing results
... phases of vertebrate development which are inconsistent with AHR being expressed only as a metabolic response to environmental chemicals. So AHR’s function goes beyond just being a regulator of xenobiotic metabolism. [1] AHR is expressed in a variety of human tissues as described above. Highest expr ...
... phases of vertebrate development which are inconsistent with AHR being expressed only as a metabolic response to environmental chemicals. So AHR’s function goes beyond just being a regulator of xenobiotic metabolism. [1] AHR is expressed in a variety of human tissues as described above. Highest expr ...
The P53 pathway and metabolism: implications in tissue
... WHY DO WE AGE AND GET CANCER? ARE THESE 2 PROCESSES INTRINSICALLY LINKED? p53 ...
... WHY DO WE AGE AND GET CANCER? ARE THESE 2 PROCESSES INTRINSICALLY LINKED? p53 ...
A caudal mRNA gradient controls posterior development in the wasp
... in patterning ancestral insects than in Drosophila. In the intermediate germ Gryllus embryo, cad plays a major role in thoracic and gnathal patterning by activating transcription of the gap genes hunchback (hb) and Krüppel (Kr). This role in gap gene activation is played by bcd and maternal hb in Dr ...
... in patterning ancestral insects than in Drosophila. In the intermediate germ Gryllus embryo, cad plays a major role in thoracic and gnathal patterning by activating transcription of the gap genes hunchback (hb) and Krüppel (Kr). This role in gap gene activation is played by bcd and maternal hb in Dr ...
Rfam Documentation
... What is your definition of an RNA family? We will group sequences into a “family” where we can identify sequence or secondary structure conservation using our covariation models. This is decided when we build our seed alignment and search the CM against rfamseq. From the resulting searches we decide ...
... What is your definition of an RNA family? We will group sequences into a “family” where we can identify sequence or secondary structure conservation using our covariation models. This is decided when we build our seed alignment and search the CM against rfamseq. From the resulting searches we decide ...
Metatranscriptome_Pipeline_Tutorial.doc
... the use of the pipeline by going through the various steps to illustrate the complexity of the process and the underlying tools and files used and generated by the pipeline. To illustrate the process we are going to use sequence reads generated from the rumen of a cow. These are 100bp paired end rea ...
... the use of the pipeline by going through the various steps to illustrate the complexity of the process and the underlying tools and files used and generated by the pipeline. To illustrate the process we are going to use sequence reads generated from the rumen of a cow. These are 100bp paired end rea ...
Gene Expression Profiles of Cold
... shown), were sequenced and BLAST analyzed. The cDNAs were divided into five categories according to the nature of deduced amino acid sequences: 17% are involved in cell wall metabolism, 12% are various transporters, 6% participate in signaling pathways, 34% are components of other cellular processes ...
... shown), were sequenced and BLAST analyzed. The cDNAs were divided into five categories according to the nature of deduced amino acid sequences: 17% are involved in cell wall metabolism, 12% are various transporters, 6% participate in signaling pathways, 34% are components of other cellular processes ...
Human mitochondrial transfer RNAs: Role of pathogenic
... brought the advantage of ATP production via the process of oxidative phosphorylation. However, the presence in mitochondria of an electron transport chain that continuously reduces oxygen to build up an electrochemical gradient required for ATP synthesis has a potentially deleterious side effect for ...
... brought the advantage of ATP production via the process of oxidative phosphorylation. However, the presence in mitochondria of an electron transport chain that continuously reduces oxygen to build up an electrochemical gradient required for ATP synthesis has a potentially deleterious side effect for ...
Real-time PCR Handbook
... choose an RT that not only provides high yields of full-length cDNA, but also has good activity at high temperatures. High-temperature performance is also very important for denaturation of RNA with secondary structure. In one-step qRT-PCR, an RT that retains its activity at higher temperatures allo ...
... choose an RT that not only provides high yields of full-length cDNA, but also has good activity at high temperatures. High-temperature performance is also very important for denaturation of RNA with secondary structure. In one-step qRT-PCR, an RT that retains its activity at higher temperatures allo ...
34386 - PubAg
... release the same blend of volatile terpenoids and indole as released when they are damaged by caterpillar feeding (9). Maize seedlings contain indole as an intermediate in at least two biosynthetic pathways. The BX1 enzyme catalyzes the conversion of indole-3-glycerol phosphate (IGP) to indole, whic ...
... release the same blend of volatile terpenoids and indole as released when they are damaged by caterpillar feeding (9). Maize seedlings contain indole as an intermediate in at least two biosynthetic pathways. The BX1 enzyme catalyzes the conversion of indole-3-glycerol phosphate (IGP) to indole, whic ...
MyGene.info Documentation
... Optional, can be a comma-separated fields to limit the fields returned from the matching gene hits. The supported field names can be found from any gene object (e.g. gene 1017). Note that it supports dot notation as well, e.g., you can pass “refseq.rna”. If “fields=all”, all available fields will be ...
... Optional, can be a comma-separated fields to limit the fields returned from the matching gene hits. The supported field names can be found from any gene object (e.g. gene 1017). Note that it supports dot notation as well, e.g., you can pass “refseq.rna”. If “fields=all”, all available fields will be ...
Analysis of the Molecular Basis of Flowering Time Variation in
... those carrying Col FLC alleles (Koornneef et al., 1994; Lee et al., 1994). In the presence of active alleles of FRI or loss-of-function mutations of the autonomous pathway FLC RNA levels from the Ler alleles are much lower than FLC levels from a Col allele (Schlappi, 2001). C24 and Niederzenz access ...
... those carrying Col FLC alleles (Koornneef et al., 1994; Lee et al., 1994). In the presence of active alleles of FRI or loss-of-function mutations of the autonomous pathway FLC RNA levels from the Ler alleles are much lower than FLC levels from a Col allele (Schlappi, 2001). C24 and Niederzenz access ...
Evolutionary dynamics of nematode operons
... million years (Myr). Of the 1133 C. elegans operons annotated, 12 did not have protein sequences in at least one of the constituent genes, and therefore, were not used in our analysis. For an additional 98 operons, their gains and losses could not be unambiguously determined for various reasons (see ...
... million years (Myr). Of the 1133 C. elegans operons annotated, 12 did not have protein sequences in at least one of the constituent genes, and therefore, were not used in our analysis. For an additional 98 operons, their gains and losses could not be unambiguously determined for various reasons (see ...
Perspectives in Diabetes Glucokinase Gene Structure
... two different promoters in the glucokinase gene therefore provides a genetic basis for the differential regulation of this enzyme in the liver and p-cell. Hepatic glucokinase, which is stimulated by an increase in the plasma insulin concentration, affects the rate of glucose usage by the liver, ther ...
... two different promoters in the glucokinase gene therefore provides a genetic basis for the differential regulation of this enzyme in the liver and p-cell. Hepatic glucokinase, which is stimulated by an increase in the plasma insulin concentration, affects the rate of glucose usage by the liver, ther ...
Horizontal gene transfer from flowering plants to Gnetum
... start of group II introns (Tables 3 and 4 and Figs. 1 and 6). Angiosperm-type nad1 exons found in Gnetum share five nucleotide substitutions with angiosperms that are not found in other seed plants. They are also characterized by a fournucleotide (GATA) insertion that makes exon b nonfunctional (Fig ...
... start of group II introns (Tables 3 and 4 and Figs. 1 and 6). Angiosperm-type nad1 exons found in Gnetum share five nucleotide substitutions with angiosperms that are not found in other seed plants. They are also characterized by a fournucleotide (GATA) insertion that makes exon b nonfunctional (Fig ...
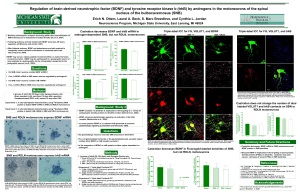
If so, is trkB mRNA in SNB motor neurons
... If so, do glutamatergic terminals or the glutamatergic post- synaptic region contain either BDNF or trkB receptor proteins? BDNF mRNA (black silver grains) in SNB (A) and RDLN (B) motoneurons from a sham GDX rat detected with 35S-labeled cRNA probes. SNB and RDLN motoneurons were counterstained wit ...
... If so, do glutamatergic terminals or the glutamatergic post- synaptic region contain either BDNF or trkB receptor proteins? BDNF mRNA (black silver grains) in SNB (A) and RDLN (B) motoneurons from a sham GDX rat detected with 35S-labeled cRNA probes. SNB and RDLN motoneurons were counterstained wit ...
The Influence of Anticodon–Codon Interactions and Modified Bases
... concentration resulting from a single tRNA gene. As discussed in the introduction, this is a reasonable approximation in species where concentrations have been measured. It has been shown that some tRNAs may be charged less efficiently than others with their appropriate amino acid (Elf et al. 2003). ...
... concentration resulting from a single tRNA gene. As discussed in the introduction, this is a reasonable approximation in species where concentrations have been measured. It has been shown that some tRNAs may be charged less efficiently than others with their appropriate amino acid (Elf et al. 2003). ...
ACTIN-RELATED PROTEIN6 Regulates Female Meiosis By
... the SWR1 ATPase, enhance complex function (Mizuguchi et al., 2004). Homologs of all yeast and 11 human SWR1 subunits have been identified in Arabidopsis thaliana (March-Díaz and Reyes, 2009; Meagher et al., 2009), indicating that Arabidopsis has the SWR1 complex. Additionally, H2A.Z deposition at man ...
... the SWR1 ATPase, enhance complex function (Mizuguchi et al., 2004). Homologs of all yeast and 11 human SWR1 subunits have been identified in Arabidopsis thaliana (March-Díaz and Reyes, 2009; Meagher et al., 2009), indicating that Arabidopsis has the SWR1 complex. Additionally, H2A.Z deposition at man ...
"RNA Interference in Caenorhabditis elegans".
... one might detect suppression or enhancement of phenotypes in mutant strains. In C. elegans, RNAi can be induced by delivering dsRNA by microinjection (see Basic Protocol 1 and Alternate Protocol 1), feeding (see Basic Protocols 2, 3, and 4 and Alternate Protocol 2), or soaking (see Basic Protocol 5) ...
... one might detect suppression or enhancement of phenotypes in mutant strains. In C. elegans, RNAi can be induced by delivering dsRNA by microinjection (see Basic Protocol 1 and Alternate Protocol 1), feeding (see Basic Protocols 2, 3, and 4 and Alternate Protocol 2), or soaking (see Basic Protocol 5) ...
What is p53
... oncogenic forms of human papillomavirus (HPV E6). In cells, p53 can associate with a 90-kD protein, identified as the product of the mdm-2 oncogene, which is amplified in some types of tumors. When bound to mdm-2, p53 can no longer function as an activator of transcription. p53 plays multiple roles ...
... oncogenic forms of human papillomavirus (HPV E6). In cells, p53 can associate with a 90-kD protein, identified as the product of the mdm-2 oncogene, which is amplified in some types of tumors. When bound to mdm-2, p53 can no longer function as an activator of transcription. p53 plays multiple roles ...
Alternatively Spliced Genes
... from left to right. The RNA helices are indicated by Roman numerals, and nucleotide positions by Arabic numerals. The shaded nucleotides are binding sites for Sm proteins, and regions marked with black lines form base-pairing interactions with the pre-mRNA. (Modified with permission from Burge, C.B., ...
... from left to right. The RNA helices are indicated by Roman numerals, and nucleotide positions by Arabic numerals. The shaded nucleotides are binding sites for Sm proteins, and regions marked with black lines form base-pairing interactions with the pre-mRNA. (Modified with permission from Burge, C.B., ...
Mouse Lefty2 and Zebrafish Antivin Are Feedback
... The murine “lefty” locus is composed of two highly conserved genes, lefty1 and lefty2, which are tightly linked on chromosome 1 (Meno et al., 1997). To examine the role of lefty2 in gastrulation and L–R asymmetry, we generated mutant mice deficient in lefty2. To inactivate lefty2, we constructed a t ...
... The murine “lefty” locus is composed of two highly conserved genes, lefty1 and lefty2, which are tightly linked on chromosome 1 (Meno et al., 1997). To examine the role of lefty2 in gastrulation and L–R asymmetry, we generated mutant mice deficient in lefty2. To inactivate lefty2, we constructed a t ...
Partial report - GEP Community Server
... The annotation of the TSS for the D. biarmipes onecut gene is based on minimizing the change in the size of the initial transcribed exon compared to the D. melanogaster ortholog (i.e. parsimony). The blastn alignment of the initial transcribed exon of the A isoform of onecut (onecut:3) from D. melan ...
... The annotation of the TSS for the D. biarmipes onecut gene is based on minimizing the change in the size of the initial transcribed exon compared to the D. melanogaster ortholog (i.e. parsimony). The blastn alignment of the initial transcribed exon of the A isoform of onecut (onecut:3) from D. melan ...
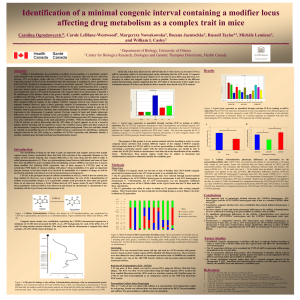
C3H/HeJ
... between APN, an in-house strain with low CYP1A2 expression, and C3H/HeJ, a laboratory strain expressing normal CYP1A2 levels, determined that this phenotype is mediated by three quantitative trait loci (QTL) localized to chromosomes 1, 4 and 9, as previously reported. The QTL on chromosome 9 co-loca ...
... between APN, an in-house strain with low CYP1A2 expression, and C3H/HeJ, a laboratory strain expressing normal CYP1A2 levels, determined that this phenotype is mediated by three quantitative trait loci (QTL) localized to chromosomes 1, 4 and 9, as previously reported. The QTL on chromosome 9 co-loca ...
a complex voyage to the X chromosome
... between the X chromosome and autosomes. Following the recognition of the X chromosome, the MSL complex may spread to active genes on the X chromosome by recognizing marks of transcription and possibly additional DNA elements. The roles of sequence elements and transcriptional marks in targeting the ...
... between the X chromosome and autosomes. Following the recognition of the X chromosome, the MSL complex may spread to active genes on the X chromosome by recognizing marks of transcription and possibly additional DNA elements. The roles of sequence elements and transcriptional marks in targeting the ...
Codon usage bias from tRNA`s point of view
... The selection-mutation-drift theory of codon usage plays a major role in the theory of molecular evolution by explaining the co-evolution of codon usage bias and tRNA content in the framework of translation optimization. Because most studies have focused only on codon usage, we analyzed the tRNA gen ...
... The selection-mutation-drift theory of codon usage plays a major role in the theory of molecular evolution by explaining the co-evolution of codon usage bias and tRNA content in the framework of translation optimization. Because most studies have focused only on codon usage, we analyzed the tRNA gen ...
Non-coding RNA

A non-coding RNA (ncRNA) is an RNA molecule that is not translated into a protein. Less-frequently used synonyms are non-protein-coding RNA (npcRNA), non-messenger RNA (nmRNA) and functional RNA (fRNA). The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene.Non-coding RNA genes include highly abundant and functionally important RNAs such as transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as RNAs such as snoRNAs, microRNAs, siRNAs, snRNAs, exRNAs, and piRNAs and the long ncRNAs that include examples such as Xist and HOTAIR (see here for a more complete list of ncRNAs). The number of ncRNAs encoded within the human genome is unknown; however, recent transcriptomic and bioinformatic studies suggest the existence of thousands of ncRNAs., but see Since many of the newly identified ncRNAs have not been validated for their function, it is possible that many are non-functional. It is also likely that many ncRNAs are non functional (sometimes referred to as Junk RNA), and are the product of spurious transcription.