* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download ppt file
Magnesium transporter wikipedia , lookup
Citric acid cycle wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Point mutation wikipedia , lookup
Catalytic triad wikipedia , lookup
Metalloprotein wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Genetic code wikipedia , lookup
Protein structure prediction wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
Biochemistry wikipedia , lookup
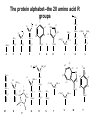
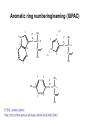
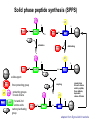
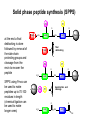
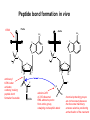
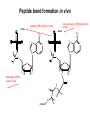
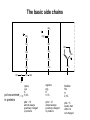
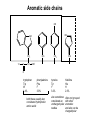
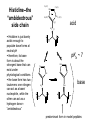
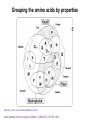
Basic protein structure and stability I: Formation of peptide bonds/ properties of amino acids Biochem 565, Fall 2008 08/25/08 Proteins are the primary functional manifestation of the information in genomes DNA sequence transcription RNA sequence translation protein sequence atgcaaactctttctgaacgcctcaagaagaggcgaattgcgttaaaaatgacgcaaaccgaa ctggcaaccaaagccggtgttaaacagcaatcaattcaactgattgaagctggagtaaccaa gcgaccgcgcttcttgtttgagattgctatggcgcttaactgtgatccggtttggttacagtacgg aactaaacgcggtaaagccgcttaa augcaaacucuuucugaacgccucaagaagaggcgaauugcguuaaaaaugacgcaaacc gaacuggcaaccaaagccgguguuaaacagcaaucaauucaacugauugaagcuggagua accaagcgaccgcgcuucuuguuugagauugcuauggcgcuuaacugugauccgguuug guuacaguacggaacuaaacgcgguaaagccgcuuaa MQTLSERLKKRRIALKMTQTELATKAGVKQQSIQLIEAGVT KRPRFLFEIAMALNCDPVWLQYGTKRGKAA protein structure protein function alpha carbon O H2N CH C O OH H3N R O The zwitterionic form is the predominant form at neutral pH side chain O C a-amino acids-the building blocks of proteins C R carboxylic acid group amino group CH H3N O C H R The alpha carbon is a chiral center--natural proteins are made of L amino acids (shown above) as opposed to D The protein alphabet--the 20 amino acid R groups NH3 O O 2 O 2 CH2 1 HN C O 2 1 N 1 C CH2 CH3 CH2 CH2 CH2 CH2 H CH2 CH A C E F G H I D H2N NH2 C CH3 S CH2 CH2 1 O NH2 2 O C C CH2 HC a P 2 H3C CH2 1 CH CH2 CH2 2 CH3 CH2 L 2 1 H3C K 1 OH NH 3 NH 1 HO CH2 CH2 CH2 OH CH2 CH2 CH2 Q R S 2 3 CH3 2 CH H2 C CH2 H2C N NH2 2 CH2 CH3 SH M 2 1 1 CH2 1 H3C CH3 1 1 2 CH2 CH N T V W Y Aromatic ring numbering/naming (IUPAC) 2 1 HN 3 7a H2 C CH 3a 7 NH2 C 4 1 a N 2 O OH NH2 H2 C CH 3 C 4 6 OH 5 5 6 4 HO 1 3 2 a 5 HN NH2 H2 C CH a C OH IUPAC nomenclature: http://www.chem.qmw.ac.uk/iupac/AminoAcid/index.html O O Proteins are made by controlled polymerization of amino acids water is eliminated O two amino acids condense to form... H2N CH O C OH H2N CH R1 OH R2 N or amino terminus H2N ...a dipeptide. If there are more it becomes a polypeptide. Short polypeptide chains are usually called peptides while longer ones are called proteins. C O CH C R1 O NH CH C R2 peptide bond is formed residue 1 residue 2 C or carboxy terminus OH + HOH Solid phase peptide synthesis (SPPS) P2 P1 O Fmoc HN O AA2 Fmoc OH AA1 HN Resin A activation deblocking Fmoc P2 P1 O Fmoc HN O AA2 AA1 H2N Resin Resin A solid support Fmoc P1 fmoc protecting group P2 AA1 AA2 A A protecting groups for side chains 1st and 2nd amino acids carbonyl activating group P2 P1 O Fmoc HN AA2 repeat steps for each amino acid in peptide, then deblock, deprotect, cleave off resin coupling O NH AA1 Resin adapted from Sigma-Aldrich website Solid phase peptide synthesis (SPPS) P2 P1 O Fmoc HN at the end a final deblocking is done followed by removal of the side-chain protecting groups and cleavage from the resin to recover the peptide NH AA1 Fmoc final deblocking P2 P1 O AA2 H2N SPPS using Fmoc can be used to make peptides up to 70-100 residues in length (chemical ligation can be used to make longer ones) AA2 O Resin P2 O NH AA1 AA2 Resin deprotection and cleavage P1 O H2N Resin O NH AA1 OH Peptide bond formation in vivo P-site t-RNA A-site NH2 NH2 N N N N O O P N O P O H H O H H O OH H H H H OH O H O R1 NH peptide N O O O O N N O aminoacyl t-RNA ester activates carbonyl, making peptide bond formation favorable O H2N adenine 2451 R2 of 23S ribosomal RNA abstracts proton from amino group, catalyzing nucleophilic attack H chemical protecting groups are not necessary because the ribosomal machinery ensures selective positioning and activation of the reactants Peptide bond formation in vivo new aminoacyl t-RNA comes into A-site peptidyl t-RNA shifts to P-site P-site A-site NH2 NH2 N N O O P N O O P O O O O H N N O O H OH H H O OH H H H H N N OH deacylated t-RNA leaves P-site O H N O R2 HN peptide H R1 H N Properties of the amino acid side chains • • • • • • size acid-base equilibria hydrophobicity/polarity tautomerism oxidation/reduction of cysteine chemical reactivity (next lecture) Sizes of amino acids a.a A R D N C E Q G H I L K vol (Å3) 88.6 173.4 111.1 114.1 108.5 138.4 143.8 60.1 153.2 166.7 166.7 168.6 surface area(Å2) a.a vol (Å3) surface area(Å2) M 162.9 185 115 F 189.9 210 225 P 112.7 145 150 S 89.0 115 160 T 116.1 140 135 W 227.8 255 Y 193.6 230 190 V 140.0 155 180 75 195 volume: Zamyatin A Prog Biophys Mol Biol 24, 107 (1972) surface area: Chothia C J Mol Biol 105, 1 (1975) 175 170 200 Acid-base titration curves of ionizable side chains base 1 eq. OHadded Asp and Glu Cys His+ Lys+ Tyr pKa Arg+ 0 3 4 5 6 7 8 9 10 11 12 13 14 physiological pH pH acid The basic side chains NH2 H2N NH3 C NH CH2 HN HO C CH2 CH2 CH2 CH2 CH2 CH2 CH NH CH2 NH2 O pct occurrence in proteins lysine Lys K 5.9% arginine Arg R 5.1% histidine His H 2.3% pKa ~ 10 almost always positively charged in proteins pKa ~ 12 almost always positively charged in proteins pKa ~ 6 means that often it is not charged The acidic side chains O O O HO C O C C CH2 CH2 CH2 CH and these carboxylic acid side chains are closely related to their amide versions... HO NH2 NH2 O C O C C CH2 CH2 CH2 CH NH2 O O aspartate Asp D 5.3% NH2 glutamate Glu E 6.3% generally negative charged in proteins because conjugate carboxylic acids have pKa of about 4 asparagine Asn N 4.3% glutamine Gln Q 4.3% Shifting of side chain titration curves base 1 NH H2 C N eq. OHadded His+ pKa NH H2 C N H 0 3 4 5 6 7 8 9 10 11 12 13 14 physiological pH pH acid Poorly populated but highly reactive forms of amino acids base form of lysine not highly populated in general at physiological pH, but is a reactive nucleophile, and if present even in minuscule amounts may do chemistry H NH3 NH2 CH2 CH2 R N O – H+ CH2 CH2 CH2 pH 7 CH2 H N C O CH CH2 NH H N C O H R CH2 CH2 CH2 CH2 CH2 CH NH HO C O CH H2O NH2 Kyte-Doolittle hydropathy of amino-acid residues side chain Ile Val Leu Phe Cys Met Ala Gly Thr Trp hydropathy index 4.5 4.2 3.8 2.8 2.5 1.9 1.8 -0.4 -0.7 -0.9 side chain Ser Tyr Pro His Glu Gln Asp Asn Lys Arg hydropathy index -0.8 -1.3 -1.6 -3.2 -3.5 -3.5 -3.5 -3.5 -3.9 -4.5 Many attempts have been made to quantify polarity, nonpolarity (hydrophobicity) of amino-acid residues in terms of scales. Kyte-Doolittle is a classic one. It is based on transfer free energies from nonpolar solvents to water combined with measurements of the tendency of residues to be buried in proteins. nonpolar--blue; polar--red; ambiguous--purple Kyte J & Doolittle RF J Mol Biol 157, 105-32 (1982) The aliphatic amino acids (plus methionine) CH3 CH3 S CH3 CH3 CH3 H HO C CH H 3C CH H 3C H 3C H CH CH2 CH2 CH2 CH2 HO NH2 O C NH O glycine Gly G 7.2% sometimes considered a "polar" amino acid alanine Ala A 7.8% the aminoacid equivalent of vanilla ice cream valine Val V 6.6% isoleucine Ile I 5.3% these are branched at the beta-carbon leucine Leu L 9.1% the most common type in proteins proline Pro P 5.2% methionine Met M 2.2% only amino acid with side-chain fused to backbone in two places to make a ring not aliphatic because of sulfur but is similar in character in many ways (nonaromatic, nonpolar residue) Aromatic side chains OH HN NH N CH2 CH2 -O C CH CH2 CH2 NH3+ O tryptophan Trp W 1.4% phenylalanine Phe F 3.9% both these usually also considered hydrophobic amino acids tyrosine Tyr Y 3.2% histidine His H 2.3% also sometimes considered an uncharged polar residue often not grouped with other aromatics and also can be charged/polar The polar uncharged side chains OH NH2 HN NH2 OH CH2 HO C CH O serine Ser S 6.8% HO H SH O O N C CH2 CH2 CH2 CH3 CH2 C CH2 CH2 histidine His H 2.3% tyrosine Tyr Y 3.2 % pKa ~ 6 for conjugate acid means that sometimes it's charged in proteins not really very polar NH2 threonine Thr T 5.9% also has some hydrophobic character due to methyl cysteine Cys C 6.8% not as polar as its sister serine but much easier to ionize to anion asparagine Asn N 4.3% glutamine Gln Q 4.3% these two have a bifunctional character in the sense of having both hydrogen bond donor and acceptor groups pKa ~ 10 means that it can be deprotonated 3 (2) Histidine--the “ambidextrous” side chain Histidine is just barely acidic enough to populate base forms at neutral pH therefore, its base form is about the strongest base that can exist under physiological conditions the base form has two tautomers: one nitrogen can act as a base/ nucleophile, while the other can act as a hydrogen donor-”ambidextrous” 2 (1) HN NH 4 (2) 1 (1) acid 5 () H2C N H O pKa ~ 7 HN N N NH base CH2 H2C N H N H O O predominant form in model peptides Cysteine and cystine Pairs of cysteines frequently undergo oxidation to a disulfide bonded form called “cystine” disulfide formation 2 CH2SH 1/2O2 CH2S 2 cysteines SCH2 H 2O cystine more hydrophobic than cysteine disulfide exchange R1 S R2S SR2 R2 S R1S disulfide exchange occurs through the thiolate anion at neutral to basic pH SR2 Key points about the character of amino acid side chains • amino acids don’t fall neatly into classes--they are different combinations of small/large, charged/uncharged, polar/nonpolar properties • how we casually speak of them can affect the way we think about their behavior. For example, if you think of Cys as a polar residue, you might be surprised to find it in the hydrophobic core of a protein unpaired to any other polar group. But this does happen. • the properties of a residue type can also vary with conditions/environment Grouping the amino acids by properties from http://www.russell.embl-heidelberg.de/aas/ which adapted it from Livingstone & Barton, CABIOS, 9, 745-756, 1993.