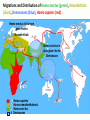
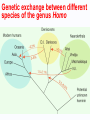
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture PPT - Carol Lee Lab
Human–animal hybrid wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Genetic engineering wikipedia , lookup
Human genome wikipedia , lookup
Adaptive evolution in the human genome wikipedia , lookup
Designer baby wikipedia , lookup
Public health genomics wikipedia , lookup
Population genetics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome (book) wikipedia , lookup
Genome evolution wikipedia , lookup
Koinophilia wikipedia , lookup
Human genetic variation wikipedia , lookup
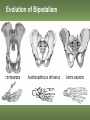
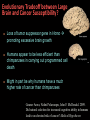
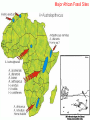
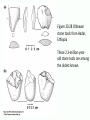
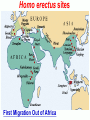
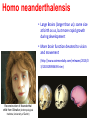
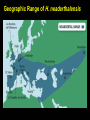
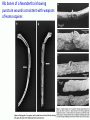
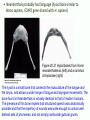
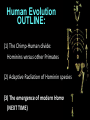
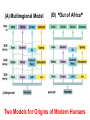
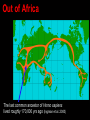
Human Evolution "Light will be thrown on the origin of man and his history” Charles Darwin, 1859 Human Evolution OUTLINE: (1) The Chimp-Human divide: Hominins versus other Primates (2) Adaptive Radiation of Hominin species (3) The emergence of modern Homo (NEXT TIME) Human Evolution OUTLINE: (1) The Chimp-Human divide: Hominins versus other Primates (2) Adaptive Radiation of Hominin species (3) The emergence of modern Homo (NEXT TIME) When did Humans Evolve? First Homo: ~2.5 mya First Australopithecine: 4-4.5 mya Chimps-Humans diverged: ~5 mya Adaptations to a Savanna Climate Change? Drier, more savanna, less forest Lead to Adaptive Radiations of early humans? How do we differ from other Animals? Animal Phylogeny First of all, we are •Bilaterally Symmetric •Triploblasts •Coelomates •Deuterostomes •And Chordates Phylogeny of mitochondrial cytochrome oxidase II alleles in humans and the African Great Apes (Ruvolo et al. 1994) Overall, genetic data reveal that among primates, we are most closely related to Chimpanzees. Most molecular phylogenies place chimpanzees as our closest relative. However, a minority of molecular phylogenies do yield alternate phylogenies (i.e. human-gorilla or gorilla-chimp clades) due to incomplete lineage sorting. Review in textbook by Herron (Chapter 20). A rough cladogram based on dental and skull characters Cladogram of Hominan Species (Based on morphology) Not certain exactly which Australopithecine led to Homo Evolution of key anatomical features Brain vs body size Dramatic increase in brain size (relative to body size) during human evolution Pilbeam and Gould, 1974 Consequences of Bipedalism • Pelvis tilt • Stress on knees, ankles, and back • Non-grasping feet • Freeing of hands Evolution of Bipedalism Sexual dimorphism has been declining during the course of Hominid evolution Changes in sexual mating system??? in the process of polygamy monogamy? Reduction in Jaw size Reduction in tooth size But insufficient reduction in tooth size or number to fit into jaw!!! (how many of you had braces?) Evolution doesn’t achieve perfection… just enough to leave enough offspring Evolution of Orthodontic problems Genetic differences between humans and other primates Humans: 23 pairs of chromosomes Gorillas & Chimps: 24 pairs Human chromosome 2 is derived from the fusion of two chromosomes that remain separate in the other great apes. Hacia (2001) H = Human C = Chimp As you can see from the previous table, the genetic differences between humans and our closest living relatives (chimpanzee, gorilla) are miniscule (tiny) • Much of the genetic differences are due to differences in gene regulation, specifically Evolution of Development • As we learned in the last lecture, depending on where the evolutionary changes are in the developmental program, even a small number of developmental genetic alterations could have profound changes on phenotype Rapid evolution in the Homo lineage Some examples of rapid evolutionary genetic changes in Homo Most of the functional evolutionary differences between Humans and other great apes affect gene regulation (which affects development) Evolution of trans-acting elements Transcription factors (trans-regulatory changes) Epigenetic trans-regulators (such as microRNAs) Genomic deletions (those that affect function mostly affect gene regulation) Rapid Evolution in Homo Evolution of gene regulation Evolution of Development Humans display a 3–5 times faster evolutionary rate in divergence of developmental patterns, compared to chimpanzees. Particularly in brain tissue… affected in many cases by trans-acting elements Trans-regulatory evolution and the costs As we know from previous lectures, it is thought that regulatory evolutionary changes that are due to cisregulatory elements would occur more readily (than trans), because of the increased pleiotropy of transacting elements However, trans-regulatory changes can have profound (large) impacts because of the many genes they regulate Also, due to the pleiotropy, trans-regulatory changes could have high costs (leading to tradeoffs) Changes in developmental genes and patterns of gene expression are greater in brain tissue than other tissues in humans relative to other primates Most different in the Brain tissue between Humans and other primates (Enard et al. 2002) Alleles unique to Homo Enard et al. Nature 2002 FOXP2: gene implicated in language (a transcription factor) 2 amino acid substitutions in Homo relative to chimps Neanderthals share the same derived allele Microcephalin (MCPH1): a gene that regulates brain size during development and has experienced positive selection in Homo Thought a derived allele (Haplogroup D) introgressed into H. sapiens by mating with extinct Homo species (Next Lecture, http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1635020/) Most differences between Homo and Chimps are due to evolutionary changes in gene expression, quantitative changes, …rather than the presence of unique alleles PNAS 2009, 106: 22358-22363 http://www.pnas.org/content/106/52/22 358.abstract Trans-regulatory evolution in the Humans (focusing mainly on the brain) Nowick et al (2009) identified 90 TF genes with significantly different expression levels in human and chimpanzee brains, among which the rapidly evolving KRAB-zinc finger genes are markedly over-represented. KRAB-zinc fingers have on average accumulated more amino acid differences between humans and chimpanzees than other genes, indicating that they may have contributed disproportionately to the phenotypic differences between these species KRAB-ZNFs affect transcription; they are transcription factors with an N-terminal KRAB domain and C-terminal zinc fingers The functions of many of these TFs are unknown Differential gene expression between humans and chimps • Most expression differences were seen in testis • 25% of transcription factors examined (18 out of 79) in this study showed changes in expression in the human brain Nowick et al. 2009 Evolution of transcription factors in the Human Brain (Nowick et al. 2009) The TFs form 2 tight gene networks (Fig 3), coordinatelyregulated in expression in the brain (co-regulating 2 sets of functional categories of genes, Fig. 5), indicating dramatic shifts within larger biological pathways The TFs are involved in regulation of energy metabolism, vesicle transport, and related functions required to build and maintain the larger and more complex human brain Human TFs are more interconnected with each other than those of chimpanzees Network of transcription factor (TF) genes that show evolutionary shifts in the human brain Module 1: Dominated by TF genes up-regulated in human brain Module 2: Dominated by downregulated TFs Both modules are enriched for primate-specific KRAB-ZNF genes Network of transcription factor (TF) genes that show evolutionary shifts in the human brain Module 1: Dominated by TF genes up-regulated in human brain Module 2: Dominated by downregulated TFs The coordinated regulation suggests that expression of these transcription factors is regulated by a common regulatory element or interacting elements (perhaps transcription factors)… So, maybe: Master transcription factors regulate transcription factors downstream Increases in links of human transcription factors So, what functions are these transcription factors regulating? What are the differences between humans and chimps? Functions of genes associated with transcription factors that underwent evolution in the human brain (2 functional modules) Module 1- Upregulated in Humans relative to chimpanzee: TF-associated genes highly enriched for functional categories involved in transcription, ubiquitination, and vesicular transport Module 2- Downregulated in Humans relative to chimpanzee: TF-associated genes, over-represented with functional categories corresponding to translation, mitochondrial function and energy metabolism Functions of genes associated with transcription factors that underwent evolution in the human brain (now shown in comparison to the chimpanzee) Evolution at the level of epigenetic trans-regulators Evolution of transcription factors and epigenetic trans-regulators (micro RNAs) in the brain Human vs Chimp divergence Figure 3. Trans-effects on developmental pattern divergence in the prefrontal cortex of the brain (Somel et al. 2011 Plos Biology) Genomic deletions in the Human relative to chimpanzee McClean et al. 2011 Nature Specific Deletions in the Human genome (McClean et al. 2011 Nature) • 510 deletions in humans relative to chimpanzees • The deletions are almost exclusively in non-coding regions and affect gene regulation • The deletions are enriched near genes involved in steroid hormone signaling and neural function • Deletions of tissue-specific enhancers may thus accompany both losses and gains of traits in the human lineage, and provide specific examples of the kinds of regulatory alterations and inactivation events long proposed to have an important role in human evolutionary divergence. Specific Deletions in the Human genome (McClean et al. 2011 Nature) Primate penile spines. Philip Reno, Stanford University • One deletion in Homo removes a sensory vibrissae and penile spine enhancer from the human androgen receptor (AR) gene, a molecular change correlated with the anatomical losses of androgen-dependent sensory vibrissae (whiskers) and penile spines (penis spines) in Homo (loss at ~700,000 yrs ago) • Another deletion removes a forebrain subventricular zone enhancer near the tumor suppressor gene: growth arrest and DNA-damage inducible, gamma (GADD45G), a loss correlated with expansion of specific brain regions in humans Evolution of the Homo brain The accelerated evolution of human brain expression appears to mainly involve remodeling of developmental patterns (evolution of development) Much of the changes are due to trans-regulatory evolution (transcription factors and micro RNAs) Some changes due to gene deletions in the Homo lineage (mostly regulatory regions, like the enhancers mentioned) Evolutionary Tradeoffs associated with rapid brain evolution Particularly since much of the evolution of the brain appears to be due to trans-regulatory evolution, …greater pleiotropic constraint of trans-acting factors (leading to consequences for other functions) Gene deletions could also lead to costs Evolutionary Tradeoff between Large Brain and Cancer Susceptibility? Loss of tumor suppressor gene in Homo promoting excessive brain growth Humans appear to be less efficient than chimpanzees in carrying out programmed cell death Might in part be why humans have a much higher rate of cancer than chimpanzees Gaurav Arora, Nalini Polavarapu, John F. McDonald. 2009. Did natural selection for increased cognitive ability in humans lead to an elevated risk of cancer? Medical Hypotheses Rapid evolution of the brain in Homo Rapid evolution new opportunities, but also new problems: Evolution of many psychiatric disorders Humans are unique among animals in being susceptible to certain neuropathologies Neurodegeneration with Aging Alzheimer's disease in the later stages of life Even healthy aging in humans is marked by variable degrees of neural deterioration and cognitive impairment, such as shrinking of the brain not found in aging chimpanzees Schizophrenia Many genes that are expressed in Schizophrenia are genes that experienced rapid positive selection in Homo Khaitovich et al. 2008. Metabolic changes in schizophrenia and human brain evolution. Genome Biology. http://genomebiology.com/2008/9/8/R124 Human Evolution OUTLINE: (1) The Chimp-Human divide: Hominins versus other Primates (2) Adaptive Radiation of Hominin species (3) The emergence of modern Homo (NEXT TIME) Human Evolution Resembles a Messy Bush rather than a continuous line The line leading to us was not always from the most “sophisticated” species at a given time For instance, Homo probably arose from the gracile australopithecines rather than from the larger brained strong robust ones Among Homo, we descended from a lineage that had smaller brains than the Neanderthals Phylogeny of Hominan Species (Based on morphology) Not certain exactly which Australopithecine led to Homo A rough cladogram based on dental and skull characters Fossil evidence of Hominin Lineages Major African Fossil Sites ? • We descended from the more gracile line (not sure which, exactly) • And not necessarily the larger brained “Lucy” Gracile Australopithecines ~2.4-2.8 mya A. africanus ~3.0-3.9 mya A. afarensis A. afarensis Robust Australopithecines A. aethiopicus ~1.9-2.7 mya A. boisei ~1.4-2.3 mya A. robustus Larger brain than gracile Australopithecines ~1.0-2.0 mya Muscular Massive jaws, sagittal crest (pointy skull to support massive jaw muscles), seed eaters Did not lead to Homo line Also called Paranthropus Hominins as prey Leopard canines fit punctures in Australopithecine skull from Swartkrans, near Johannesburg, South Africa Homo habilis “handy man” ~1.6-1.9 mya Tool User Figure 20.28 Oldowan stone tools from Hadar, Ethiopia These 2.3-million-yearold stone tools are among the oldest known ? Homo ergaster = erectus ~1.5-1.8 mya First Migration Out of Africa First use of Fire (South African cave ~1 million yrs ago) Homo erectus sites First Migration Out of Africa Emergence of diverse Homo species across a broad geographic range • Homo erectus spread out of Africa throughout Eurasia and gave rise to multiple species of Homo in different geographic regions • Multiple sister taxa of Homo then evolved in different geographic regions (H. sapiens, H. neaderthalensis, Denisovans, archaic Homo in Africa) • The multiple sister species of Homo then came into contact as the species migrated ? • We descended from the more gracile line (not sure which, exactly) • And not necessarily the larger brained Homo neanderthalensis ~28,000-300,000 yrs ago Large Brains (larger than ours) Occurred outside of Africa Complex Culture Reconstruction of Neanderthal child from Gibraltar (Anthropological Institute, University of Zurich) Originally called Homo sapiens neanderthalensis. Because of its larger brain, we assumed that it had to be the same species as us Homo neanderthalensis Did Homo sapiens (we) intermate with Neanderthals? Why did they go extinct? • Evolved outside of Africa (mostly Europe) • Complex culture: more primitive tools initially, but then after H. sapiens invaded Europe out of Africa, they then adopted H. sapiens tool technology • Overlapped in geography with H. sapiens in Europe for about 10,000 years Reconstruction of Neanderthal child from Gibraltar (Anthropological Institute, University of Zurich) • Extinct ~25,000 yrs ago Homo neanderthalensis • Melanocortin 1 receptor (Mcr1) allele mutations (loss of function) indicate that at least some had red hair and fair skin (different mutation found in H. sapiens)… likely to be independent evolution in low UV environment Reconstruction of Neanderthal child from Gibraltar (Anthropological Institute, University of Zurich) Homo neanderthalensis • Large Brains (larger than us): same size at birth as us, but more rapid growth during development • More brain function devoted to vision and movement (http://www.sciencedaily.com/releases/2013/0 3/130319093639.htm) Reconstruction of Neanderthal child from Gibraltar (Anthropological Institute, University of Zurich) Geographic Range of H. neaderthalensis Rib lesion is consistent with injury by a long-range projectile weapon traveling along a ballistic trajectory Generally, projectile weapons are more commonly associated with H. sapiens. Rib bones of a Neanderthal showing puncture wounds consistent with weapons of Homo sapiens Homo neanderthalensis • Buried dead, had rituals • Art, radiocarbon dated to ~43,500 and 42,300 years ago in Spain, before H. sapiens thought to have colonized this region (older than H. sapiens art, 30,000 yr old Chauvet cave paintings) Spain's Nerja caves • Neanderthals probably had language (hyoid bone similar to Homo sapiens, FOXP2 gene shared with H. sapiens) Figure 20.31 Hyoid bones from Homo neanderthalensis (left) and a common chimpanzee (right) The hyoid is a small bone that connects the musculature of the tongue and the larynx, and allows a wider range of tongue and laryngeal movements. The bone found in Neanderthals is virtually identical to that of modern humans. The presence of this bone implies that structured speech was anatomically possible and that the repertory of sounds was wide enough to contain welldefined sets of phonemes, and not simply inarticulate guttural grunts. Denisovans New species of Homo In March 2010, a finger bone fragment of a juvenile female that lived about 41,000 years ago was found in Denisova Cave in Altai Krai, Russia; a tooth and toe bone belonging to different members of the same species have since been found. This region was also inhabited at about the same time by Neanderthals and perhaps modern humans. Denisovans ranged from Siberia to Southeast Asia Why did our congeners (all other species of Homo) go extinct? Hypotheses: climate change? We killed them directly We domesticated dogs 40,000 and as a consequence became superior at hunting Questions Adaptive radiation of Hominin lineages (Australopithecines and Homo) led to multiple hominin species that overlapped temporally and geographically So, given this overlap in space and time, where did modern Homo sapiens originate, and from which species? And did other species of Homo contribute to the genomic composition of Homo sapiens? Human Evolution OUTLINE: (1) The Chimp-Human divide: Hominins versus other Primates (2) Adaptive Radiation of Hominin species (3) The emergence of modern Homo (NEXT TIME) Where did Modern Humans Come From? (A) Multiregional Model (B) “Out of Africa” Two Models for Origins of Modern Humans Predictions (A) Multiregional Model (B) “Out of Africa” Large Genetic Differences among human populations (1.5 Million Years) Small Genetic Differences among human populations (small effective population size) Genetic Diversity Equal Everywhere Most Diversity in Africa (founder effect) Modern Humans Based on mitochondrial data alone it appeared that the “Out of Africa” hypothesis was correct, with no introgression with local species of Homo Vigilante et al. 1991 Science Phylogeny of mitochondrial DNA Ancestral alleles are in Africa A phylogeny of modern H. sapiens using mitochondrial DNA • Most of the Genetic Diversity is in Africa • Ancestral alleles are in Africa • Non-Africans are nested within the African clade (nonAfrican alleles are a subset of African alleles) • Supports the scenario that H. sapiens originated in Africa, and a small subset migrated out of Africa Out of Africa The last common ancestor of Homo sapiens lived roughly 170,000 yrs ago (Ingman et al. 2000) How are we related to our co-occurring sister taxa? (which tended to split off earlier from our ancestors, so they are often called “archaic”) Neanderthals vs Modern Humans Summary from several early studies based on DNA sequence data : Igman et al. (2000), Krings et al. (2000), Ovchinnikov et al. (2000), Hoffreiter et al. (2001), Green et al. (2006), Green et al. (2008) According to this data set, Homo sapiens and Homo neanderthalensis are quite distinct, separated by (~500,000 yrs); that is, we split from a common ancestor ~500,000 years ago Neanderthals form a separate Sister clade to Modern Humans; so, they are a related sister species, NOT our direct ancestors People call the other Homo species (Neanderthals, Denisovans, African Homo erectus descendant) “archaic”, but they are out sister taxa, NOT ancestral • In the initial genetic studies (mtDNA and partial genome sequencing), Neanderthal genes were absent in current human populations • These preliminary results suggested there was no genetic contribution from Neanderthals • And suggested that they went extinct without leaving any genetic signature in the current populations of Homo However, more comprehensive genome sequencing reveals genetic interbreeding between H. sapiens and H. neanderthalensis (work from Svante Pääbo’s lab, Science, 2010) http://www.sciencemag.org/content/328/5979/710.full Overall, genome sequence is 99.5% to ~99.9% identical with H. sapiens The Neanderthal line began to diverge from Homo sapiens by about 800,000 years ago and that we were "genetically distinct" by 300,000 years ago However, about 1-4% of DNA in Modern Europeans and Asians was inherited from Neanderthals No evidence of interbreeding between the two species in Africa (Neanderthals did not occur in Africa, but originated in Europe) Ozzy Osbourne's Genome Reveals Some Neandertal Lineage By Katherine Harmon What genetic oddities does rock's Prince of Darkness and beheader of bats have entangled deep in his genetic code? Knome, the company that analyzed Ozzy's full genome, divulges some of the details in a Q&A http://www.scientificamerican.com/a rticle.cfm?id=ozzy-osbourne-genome Homo neanderthalensis • Asymmetric Gene Flow: No evidence for gene flow in the direction from modern humans to Neanderthals. (only from Neanderthals to Homo sapiens) • This result would not be unexpected if contact occurred between a small colonizing population of H. sapiens and a much larger resident population of Neanderthals. • While modern humans share some nuclear DNA with the extinct Neanderthals, the two species do not share any mitochondrial DNA, which in primates is always maternally inherited. • This observation has prompted the hypothesis that H. sapiens females x male Neanderthals were able to generate fertile offspring, whereas the progeny of female Neanderthals and male H. sapiens were either rare, absent or sterile. Introgression of Neanderthal genes leading to local adaptation? • The Neanderthal DNA in modern human populations includes some of the genes for our HLA immune system (MHC loci). • It has been suggested that this gave early modern human immigrants to Europe and Asia critical protection to diseases that had not existed in their African homeland. • Mating with Neanderthals might have aided the migrating Homo sapiens adapt to local pathogens. • The initial genetic studies were not wrong, but simply told only part of the story • 1-4% introgression from Neandertals to H. sapiens represents only a small part of the genome, and easy to miss in partial genome sequence data. Neanderthals vs Modern Humans • So, the phylogeny above is correct across most of the genome • Except, that the whole genome sequence data indicate some small amount of introgression (1-4%) of DNA from Neanderthals to nonAfrican humans Denisovans Genome sequencing indicates that Denisovans are more closely related to Neanderthals than to Homo sapiens The estimated average time of divergence between Denisovan and Neanderthal sequences is 640,000 years ago, and that between both of these and the sequences of modern Africans is 804,000 years ago. About 1% of Chinese and up to 6% of the DNA of Melanesians and Australian Aborigines are derived from Denisovans Callaway, 2011. Ancient DNA reveals secrets of human history, Nature http://www.nature.com/news/2011/110809/full/476136a.html Hybridization with other Homo to acquire immunity alleles Half of the HLA (human leukocyte antigen, encode for MHC) alleles of modern Eurasians represent archaic HLA haplotypes likewise inferred to have introgressed from Denisovans or Neanderthals For example, HLA type allele HLA-B*73 introgressed into humans in west Asia from Denisovans These alleles, of which several encode unique or strong ligands for natural killer cell receptors, now represent more than half the HLA alleles of modern Eurasians and also appear to have been later introduced into Africans. The apparent over-representation of these alleles suggests a positive selective pressure for their retention in the human population. Abi-Rached et al. 2011. The Shaping of Modern Human Immune Systems by Multiregional Admixture with Archaic Humans. Science. http://www.sciencemag.org/content/334/6052/89.full Introgression with other Homo lineages in Africa? Contemporary African populations contain a small proportion of genetic material (~2%) that introgressed ~35,000 years ago from an “archaic” (sister) species of Homo that split from the ancestors of anatomically modern humans ~700,000 years ago Migrations and Distribution of Homo erectus (green), Neanderthals (olive), Denosovans (blue), Homo sapiens (red) Homo erectus in Europe gave rise to Neanderthals Homo erectus in Asia gave rise to Denisovans Homo sapiens Homo neanderthalensis Homo erectus 4 Denisovans Genetic exchange between different species of the genus Homo Studying the genomes of our sister species relative to our genome reveals genes that are uniquely Homo sapiens and that have undergone selection during our recent history (last few hundred thousand years) Origins of Homo sapiens (1) “Out of Africa” is mostly correct, as anatomically modern Homo sapiens emerged out of Africa ~120,000 years ago (2) Such that genetic divergences among modern human populations are small (2) Most of the Genetic Diversity is in Africa and a clade nested within the African clade is found outside of Africa, indicating that ancestral populations originated in Africa, followed by recent founder effects and genetic drift as populations moved out of Africa Origins of Homo sapiens (3) Mitochondrial and partial genome sequence data suggested that all other Homo lineages went extinct without contributing to Homo sapiens (4) HOWEVER, whole genome sequencing of other species of Homo indicate that H. sapiens did mate with other species, at least with Homo neanderthalensis and Denisovans and statistical genomic analyses suggest intermating with African lineages of archaic Homo (5) Thus, the “multiregional hypothesis” is partially correct, in that modern populations of Homo did receive genetic contributions from archaic Homo species (sister species) in different geographic regions Origins of Homo sapiens So, H. sapiens did emerge recently Out of Africa recently, only ~100,000 years ago (most of the genome reflects this genomic signature) But then, intermated with local sister species of Homo in different regions throughout Africa and Eurasia (1-4% of the genome reflects this introgression) How long will we last? Neanderthals abd Denisovans were successful lineages of Homo for 300-400,000 years. We have only been around for ~120,000 years… Will we survive as long as our sister taxa, or go extinct sooner? Migrations of H. sapiens Figure taken from: John K. Wiencke. 2004. Impact of race/ethnicity on molecular pathways in human cancer. Nature Reviews Cancer 4:79-84 Native American Frequency of allele Pacific Islander Asian European Genetic Drift during Human Migrations Loss of Alleles Middle Eastern NE African Sub-Saharan African Alleles at a locus Tishkoff et al. 1996 Lots of reticulation among populations “races” do not separate out into neat groups How will we evolve next? Evolution in Contemporary Human Populations Much of the genome in human populations is under selection 304 (9.0%) out of 3,377 potentially informative loci show evidence of rapid amino acid evolution (Bustamante 2005) Currently, signatures of negative selection (selection against alleles) are used to find alleles that cause disease, such as alleles that cause diabetes, obesity, etc. Evolution in Contemporary Human Populations Modern lifestyle imposes selection (environment has shifted recently) Selection in response to diseases Microbial disease agents are evolving more quickly now (antibiotics), and impose selection on human populations For example, AIDS is the leading cause of deaths worldwide, and is applying intense selection on human populations Genocide and war Example: more than 90% Native Americans killed off Many other examples of Genocide leading to decimation of ethnic groups (loss of genetic diversity) Recombination among previously isolated populations New gene combinations (novel genotypes) Increases in heterozygosity We are products of our Evolutionary History Hunter-gather lifestyle: 95% of our evolutionary history Diets (typically High fiber) Exercise (walk 7-10 miles/day) Lower calorie diet resulting in later menses, fewer children (no more than 4, spacing of children due to lactation) Smaller Groups (~25) Our environment has shifted recently… The Onion: Human Feet Originally Used For Walking, Anthropologists Report July 22, 1998 | ISSUE 48•17 ISSUE 33•25 OXFORD, ENGLAND—A new report in the Journal Of The Anthropological Society Of Oxford reveals that human feet were likely once used as a means of extravehicular locomotion. "Apparently, as recently as 20 years ago, the foot was used in a process called 'walking,' by which the human body actually propelled itself," the report read. "Starting sometime in the late 1970s, these crude early feet gradually evolved into their present function of operating the gas and brake pedals on automobiles." The same team of researchers discovered in 1994 that the human brain was once used for various problem-solving applications before evolving into an absorption/storage unit for lyrics to TV-show Evolution in Future Human Populations The Future: Genetic Engineering We are already genetically screening embryos following invitro fertilization (see film Gattaca 1997) We should start thinking about the ethics involved in genetic engineering humans (we are already engineering agricultural species) Summary (1) Genetic differences between human and chimps are small; differences are mostly regulatory (development), especially trans-regulatory… some cis-regulatory changes (2) There was an adaptive radiation of hominid species ~3 mya, such that several species coexisted (3) Overall pattern toward larger brains, smaller teeth and jaws, longer legs, less sexual dimorphism… (4) Evolution is not perfect: jaw and tooth evolution was not that well-coordinated (orthodontics); knee, ankle, hip problems associated with bipedalism (8) Evolution occurs in a jagged and bushy manner; i.e., we did not always descend from the more robust or bigger brained species, even though on average brain size was increasing Summary (6) Genetic data of human populations support that Modern humans evolved in African relative recently, and then migrated out of Africa ~100,000 years ago (7) There is evidence of interbreeding between Homo sapiens and our closest relatives Homo neanderthalensis, even though the species are quite divergent and split ~500,000 years ago (8) Genetic drift occurred as humans migrated out of Africa, with loss of alleles from Africa, to Europe, to Asia, to Native North America (9) Large variance in male reproductive success in evident in human populations (e.g. Genghis Khan) (10) Human populations are under strong selection, especially in response to diseases 1. Which statement is FALSE regarding genetic differences between humans and other apes? (a) There are structural evolutionary differences in the alleles of FOXP2 that are unique to Homo sapiens and Homo neanderthalensis versus those of chimpanzees (b) The genome sequences of humans and other great apes are mostly identical, with less than 2% differences in coding sequences (c) Some key differences between human and chimpanzee gene expression appear to be due to differences in expression of transcription factors (d) The transcription factors that differ in expression between humans and chimpanzees form networks of coordinately-regulated genes (all up or down in expression as a unit) (e) Most genetic differences between humans and chimps appear to be due to cis-regulatory changes 2. Which of the following is FALSE regarding human brain evolution? (a) Rapid brain evolution in humans is likely due to evolutionary changes in developmental genes (b) Rapid brain evolution in humans might involve rapid evolution of expression of trans-acting factors, such as microRNAs (c) Percentage of genes that are differentially expressed between humans and chimpanzees are much greater in the brain than in other tissues, such as the testis (male sex organs) (d) Rapid brain evolution in humans might involve rapid evolution of expression of trans-acting factors, such as transcription factors (e) A gene deletion of a tumor suppressor gene that allowed growth in the brain might also allow the proliferation of cancer, leading to an evolutionary tradeoff 3. What are some anatomical features that do NOT distinguish humans from other great apes? (a) Bipedalism (b) Low brain to body size ratio in Homo (c) Shape of the pelvis (d) Language (e) Reduction in tooth and jaw size in Homo 4. Which of the following is FALSE regarding genetic differences between humans and their closest relatives (chimpanzees)? (A) Most of the DNA sequences are identical between humans and chimps; only about 1.6% of total sequence, and only about 1% of the coding region differs (B) Physical differences between humans and chimps are largely caused by differences in regulatory genes, such as those involved in gene expression (such as in the brain) (C) Humans do not have unique genes relative to chimpanzees. All the differences are due to the regulation (turning off and on) of shared genes (D) A few mutations in a few genes could affect biochemical or cellular processes on a global scale (across all cells or causing profound effects on biochemical pathways) 5. Which of the following is false regarding Hominin evolution? (a) All hominin fossil species prior to Homo erectus are found only in Africa (b) Evolution of Homo body parts was not synchronous, such that the brain, organ tissues, jaw, teeth, etc. diverged at different rates (c) Australopithecines are less likely to have needed braces to straighten their teeth relative to species of Homo (d) The lineage leading to Homo sapiens was generally derived from the larger bodied and larger brained ancestral species (e) During the course of hominin (Australopithecines and Homo) evolution several different hominin species co-existed at the same time. Answers: 1E 2C 3B 4C 5D 1. Which of the following is TRUE regarding patterns of human evolution? (A) Homo neanderthalensis shares no alleles with Homo sapiens (us), suggesting that interbreeding never occurred (B) The lineage leading to Homo sapiens generally evolved from the largest bodied and largest brained Hominid species from any given time period (C) Homo sapiens is the only Hominid species to leave Africa (D) On average, brain size grew dramatically relative to body size during the course of Hominid evolution 2. Based on what evidence do we now think that Homo sapiens mated with other species of Homo, such as Denisovans and Neanderthals? (a) Sequences of mitochondrial genomes show that there has been some introgression of genes from other species of Homo into Homo sapiens (b) Partial genome sequences from Homo neanderthalensis and Denisovans (~15% of the genome sequence) began to show evidence of introgression (c) Full genome sequences from bone fragments of "archaic" Homo species from Africa show evidence of introgression (d) Alleles unique to Homo show evidence of introgression (e) It was not until we obtained the whole genome sequences of our sister species of Homo that we were able to determine that a small proportion of human genomes (~1-5%) was derived from other species of Homo 3. Current molecular genetic data in general support the hypotheses that modern humans emerged out of Africa recently and then spread all over the world (followed by some admixture with local Homo species). Which line of evidence would support this hypothesis? (A) Low genetic divergences among modern human populations, and loss of alleles during migrations with greater distance from Africa (B) Low genetic diversity among human populations, and increases in genetic diversity with increasing distance from Africa (C) Genetic Drift and genetic exchange with Homo neanderthalensis (D) Large genetic divergences among modern populations, and loss of alleles during migrations with greater distance from Africa 4. Which of the following statements is NOT supported by phylogenies of modern humans? (A) Movement of modern humans out of Africa was characterized by founder effects with increasing distance from Africa. (B) A very large proportion of the human genetic diversity exists among Africans. (C) So far, there is no concrete evidence that Homo erectus or Homo neanderthalensis contributed genetically to modern human populations. (D) Ethnic groups of modern Homo sapiens do not separate out cleanly into distinct clades, probably as a result of recent separation (incomplete lineage sorting) and admixture 5. Which of the following would be a good strategy for studying traits that are unique to Homo neanderthalensis? (a) Examine genes that are divergent between Homo and other primates (b) Examine genes that are shared between Homo sapiens, Denisovans, and Neanderthals (c) Examine genes that have introgressed into the Homo sapiens genome from Homo neanderthalensis (d) Examine genes that are present in the Homo neanderthalensis genome, but not present in the Homo sapiens genome (e) Examine genes that are polymorphic in the Homo neanderthalensis genome 6. Which types of alleles show an overrepresentation of introgression from Neanderthals and Denisovans into Homo sapiens populations? (a) Paralogs (b) hox genes (c) MHC loci (d) trans-acting factors (e) Kernels of GRN 7. Which of the following is the MOST CORRECT regarding the origins of modern Homo sapiens? (a) Homo sapiens migrated out of Africa ~120,000 years ago, and migrated across Europe and Asia without genetic exchange with other species of Homo in Eurasia or Africa (b) Lineages Homo each evolved independently in multiple geographic regions for ~1 million years, leading to modern Homo sapiens populations (c) Homo sapiens migrated out of Africa ~120,000 years ago, but there is now evidence that Homo neanderthalensis, Denisovans, and other species of Homo contributed roughly 1-5% to the genomes of modern human populations (d) Homo sapiens migrated out of Africa ~200,000 years ago and intermated with Homo neanderthalensis, Denisovans, while populations within Africa remained genetically the same (e) The genus Homo experienced an adaptive radiation followed by hybridization, leading to complete fusing of the species and loss of genetic differentiation among species of Homo Answers 1D 2E 3A 4C 5D 6C 7C Optional Slides Evolution of HIV resistance in Human Populations CCR5-Δ32 (or CCR5-D32 or CCR5 delta 32) is a mutant allele of the receptor CCR5, where the deletion of a 32 base pair segment makes the receptor nonfunctional The allele has a negative effect upon T cell function, but appears to protect against smallpox and HIV HIV has no receptor to bind to and cannot enter the cell This allele is found in 14% of Europeans HIV can impose selective pressure for CCR5-Δ32, increasing the frequency of this allele in human populations (Sullivan et al. 2001) Amy D. Sullivan et al. 2001. The coreceptor mutation CCR5Δ32 influences the dynamics of HIV epidemics and is selected for by HIV. Proc Natl Acad Sci USA. 98: 10214–10219. Evolution of HIV resistance in Human Populations CCL3L1 Some individuals have resistance against HIV-1 due to high number of copies of the CCL3L1 gene (Chemokine (C-C motif) ligand 3-like 1). This protein binds to chemokine receptor CCR5, and competes with HIV for binding Copy number of this gene varies among individuals; most individuals have 1-6 copies in the diploid genome. With increased copy number, there is more CCL3L1 expressed, and so competition for the CCR5 binding site is increased. This leads to a decrease in the ability of HIV to infect the cell. Gonzales et al. 2005. The Influence of CCL3L1 Gene-Containing Segmental Duplications on HIV-1/AIDS Susceptibility. Science. 307(5714):1434-1440. Genetic Diagnostics Genetic testing for Ancestry and disease alleles has become very common For example, “23andMe”: https://www.23andme.com/