* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Diapositiva 1 - Progetto Onev
Gene regulatory network wikipedia , lookup
Gene expression profiling wikipedia , lookup
Expression vector wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Molecular evolution wikipedia , lookup
Gene expression wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Point mutation wikipedia , lookup
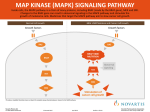
IRCCS Istituto Tumori Giovanni Paolo II, Bari, Italy Omica e nanotecnologie negli ESSERI VIVENTI Il Progetto ONEV Biomarkers for therapeutic response in metastatic melanoma S. Tommasi Bari, 1 Dicembre 2014 APPROVED AGENTS USED FOR THE TREATMENT OF METASTATIC MELANOMA Treatment type Approved drugs Mechanism of action Mechanism of resistance Chemotherapy DTIC, TMZ DNA methylation High expression MGMT MMR deficiency BER deficiency Inhibition of BRAFV600 mutant MAPK pathway reactivation PI3K/AKT/mTOR reactivation Trametinib Blockade of MEK in presence of BRAFV600 mut MAPK pathway reactivation IL2 Stimulation of T and NK cell-mediated acitivities Tumor escape from immune surveillance mechanisms Ipilimumab Tremelimumab CTLA4 blockade with Low melanoma antigen enhacement of effector expression T cell function and inhibition of Treg activity Pembrolizumab Nivolumab Binding to PD-1 resulting in enhancement of the immune response against tumor Targeted therapy Vemurafenib Dabrafenib Immunotherapy - Base excision repair (BER) key factors : predictive biomarkers - new promising targets for therapies in detail Liu L , and Gerson S L Clin Cancer Res 2006;12:328-331 PROGNOSTIC AND PREDICTIVE ROLE OF MGMT METHYLATION Tuominen et al. 2014 Base excision repair (BER) key factors : new promising targets for therapies in detail Liu L , and Gerson S L Clin Cancer Res 2006;12:328-331 Prognostic role in pts BRAF V600 untreated by Vemurafenib APE1 inhibition induces a synthetic lethal response in PTEN-deficient cells by causing accumulation of abasic sites and subsequent strand breaks, and ultimately the induction of apoptosis Abbotts R – Madhusudan 2014 BRAF C A U S E S o f R A F k i n a s e I N H I B I TO R S R E S I TA N C E NRAS & MEK mutations BRAF amplification Stromal secretion of HGF-like COT or EGFR activation NF1 loss ↓ BIM expression via PTEN loss Disrupted feedback regulation MAPK pathway 44% Downstreem effectors: NRAS, BRAF, MAP2K1, MAP2K2, MITF, NF1 Alternative splicing of BRAF mRNA PI3K pathway: PIK3CA, PTEN, PIK3R1 HOXD8 RAC1 Van Allen EM 2014 NRAS as predictive biomarker PI3K i MEK i IL2 Ipilimumab Next Generation Sequencing and Custom Ampliseq Panels: a new opportunity to predict response to chemotherapy, targeted therapy, immunotherapy………. SENSITIVITY TO ………. Project PON01_01297 “Virtualab” Next Generation Sequencing and Custom Ampliseq Panels: a new opportunity to predict response to targeted therapy The aim was to identify mutations able to predict therapy response in a quick, accurate and cost effective method. The panel was developed to analyze the coding region of 11 genes with a coverage of 93.85% . The melanoma custom panel size was 39.08Kb, contains 303 amplicons and for the analysis is required an input of 20ng of FFPE DNA (2 pools). Pinto confidential 2014 Patients in which at least one gene results mutated 90 % of alterations 80 70 60 50 40 30 20 10 0 NRAS CTLA4 MITF PIK3CA KIT BRAF MGMT PTEN CDK4 RB1 MC1R Pinto R confidential 2014 ION PGM, ARMS AND SANGER METHODS ION PGM Sanger Method BRAF V600 mut n.11 1 false negative BRAF V600 wt n.6 2 false positive NRAS Glu61 mut n.2 ARMS Method 1 false negative Pinto R confidential 2014 U S I N G c f D N A A D V A N TA G E S Analysis in the lack of tumor material in patients whose tumors are difficult to biopsy The small amount of archival tumor tissue mixed with normal stroma tissue Degraded DNA by formalin fixation – cfDNA presented less non specific amplifications Potentially mutant clones could be missed in biopsy from small part of tumor Shorten turnaround timefor testing PRE-ANALYTICAL AND ANALYTICAL PHASES SERUM PLASMA Sensitivity Specificity Sensitivity Specificity 44% 96% 52% 96% 95%CI 35%53% 95%CI 90%99% 95%CI 43%61% 95%CI 90%99% More BRAF mutations were detected in plasma than in serum although differences in assay sensitivity and specificity are not statistical different Plasma contains less wt DNA than serum but higher mutation fraction CONCORDANCE SERUM PLASMA 64% 70% Discordance between tumor and cfDNA could depend: on time of sampling Archival tumor biopsy did not fully represent the tumor heterogeneity Different mutation status in primary and metastatic sites Sensitivity of assays: ARMS 2% Aung 2014 Biomarker analysis in Phase II Dabrafenib study Ascierto JCO 2013 CORRELATION tumor tissue - cfDNA SPECIFICITY SENSITIVITY V600E 84% 100% 79% V600K 97% 99% 89% P=0.0134 Significant association between baseline cfDNA mut fraction and OS (HR:0,83; 95%CI, 0,72-0,96) P=0.0006 Significant association between baseline cfDNA mut fraction and PFS (HR:1,09; n=46) STUDY DESIGN AIM: to individuate novel candidate peptides useful as biomarkers to predict the response or resistance to treatment in two sets of patients treated with TMZ/FM and vemurafenib 39 patients - serum charge range 1,5 – 50 D IMAC 30 19 BRAF V600E/K Vemurafenib 17 BRAF wt 3 BRAF V600E Temozolomide/FM Garrisi, Strippoli Plos One 2014 M/Z P ROC Intensity Intensity Area BRAF mut BRAF wt Regulation 9446 0,0148 down 0,715 6,182 8,43 9295 0,0217 up 0,715 16,709 8,296 1883 0,023 up 0,296 14,284 7,596 9176 0,025 down 0,704 6,9 8,22 4652 0,027 down 0,696 7,4 11,351 BRAF mut vs BRAF wt Sequence Acrostic name Description coverage Score (%) SLAIN1 Invasiveness GLIS2 ABCC12 Resistance SLAIN motif-containing protein 1 20 38 Zinc finger protein 21 36 Multidrug resistance-associated protein 9 6 27 12 18 Cyclic AMP-dependent transcription ATF6 factor Garrisi, Strippoli Plos One 2014 VEMURAFENIB RESPONDERS VS NON RESPONDERS Regulation in M/Z p-value Shorter Resp 5900 0.019 Acrostic Sequence Description name Score coverage (%) up 12544 0.033 up 49124 0.048 up 11724 0.022 up KLF17 Krueppel-like factor 31 40 RBM10 RNA-binding protein 12 26 20 22 TOX high mobility group box TOX3 family member 3 PFS P=0.0011 Regulatory activity of gene transcription Basal Level RNA alternative splicing Overexpressed Garrisi, Strippoli Plos One 2014 TMZ / FM RESPONDERS VS NON RESPONDERS BRAF wild-type patients Regulation in M/Z p-value Resp 6411 0.022 Acrostic name NQO1 up Description Sequence coverage (%) Score NAD(P)H dehydrogenase [quinone] 1 21 28 25 28 COMM domain-containing protein - HypertensionCOMD5 4075 0.020 Related Calcium-Regulated Gene Protein up CA4 Carbonic anhydrase 4 18 26 VATL V-type proton ATPase 16 kDa proteolipid subunit 41 26 TM50A Transmembrane protein 50A 37 26 PFS P=0.0024 Pathway of detoxification Overexpressed Cell acidification Basal level Garrisi, Strippoli Plos One 2014 miRNAs NEW PREDICTIVE BIOMARKERS? Greemberg 2014 miRNA expression in Metastatic Melanoma * Our cohort included 43 patients (treatment naïve and with histologically confirmed stage IV of metastatic melanoma), 30 cases were BRAF mutated at the codon 600, while 13 were wild type; * We have selected 15 miRNAs that scientific reports and informatics tools have established to target the crucial genes involved in melanoma biology.; * We analyzed miRNAs expression and the expression of the correspondent target genes by TaqMan probes; * We correlated miRNAs and gene expression data to time to progression of 20 patients treated with Vemurafenib. Pinto R confidential 2014 Non -invasive biomarkers : circulating miRNAS Proteic complexes Tumor miRNA biogenesis Tailored therapy Microvescicles Clinical decision miRNA release Clinical biomarkers Prognosis miRNA extraction (blood) Apoptotic bodies Passive delivery into bodily fluids Quantification qRT-PCR EXPERIMENTAL WORKFLOW BLOOD BASED miRNA STUDY AND ADVANTAGES Non invasive technique Possibility to multiple time point sampling Tissue specific dysregulation Stability to RNAase digestion, harsh conditions. Extended storage and multiple freeze-thaw cycles. Circulating mir125b expression in response to TMZ/FE treatment: preliminary results TP53 DNA repair and damage prevention De Summa confidential 2014 Cellular senescence CONCLUSIONS Deregulated miRNAs and their respective targets may serve as molecular targets/tools for future platform of molecular directed therapy or as predictive biomarkers in melanoma patients. Only integrated studies on genetics and epigenetics are needed to evidence a complete pattern of pharmacological sensitivity THANKS to: Molecular Biology Laboratory Pinto R. De Summa S. Garrisi V.M. Medical Oncology Unit Guida M. Strippoli S. Anatomopathology Unit Simone G. Popescu O. In vitro Pharmacology Laboratory Azzariti A. Porcelli L. Quatrale A.E. Biology & Biochemistry Lab Univ Bari Guida G Maida I. Cocco T. Grieco C. PON01_01297 Virtualab PONa_00034 ONEV