* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download ISMB2006-Docking7
Survey
Document related concepts
Transcript
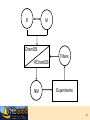
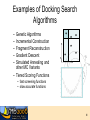
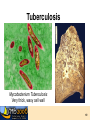
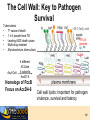
7. Molecular Docking and Drug Discovery R M ChemDB Filters RChemDB NM Experiments 2 The Docking Problem • • Given: receptor binding pocket and ligand. Task: quickly find correct binding pose. Two critical modules: 1. Search Algorithm 2. Scoring Function 3 Definitions • pKd = measures tightness of binding • pKi = measures ability to inhibit • Mechanisms of action—for instance: – Competitive inhibition (most typical docking case) – Allosteric inhibition (bind to different pocket) – Allosteric activation 4 Challenges • Search algorithm – Speed (5M compounds or more) – Local minima – High-dimensional search space • Scoring function – – – – – Strict control of false positives Good correlation with pKd Multiple terms No consensus Non-additive effects (solvation, hydrophobic interactions) • Note: pKd does not always correspond with activity • ADME concerns 5 Examples of Docking Search Algorithms – – – – – Genetic Algorithms Incremental Construction Fragment Reconstruction Gradient Descent Simulated Annealing and other MC Variants – Tiered Scoring Functions • fast screening functions • slow accurate functions 6 High Dimensionality: Flexibility • Most algorithms handle ligand flexibility but do NOT handle receptor flexibility. • Iterative Docking to find alternate conformations of the protein – Dock flexible ligand – Minimize receptor holding ligand rigid – Repeat 7 Scoring Function • Energy of Interaction (pKd) • • • • • • Electrostatics Van der Waal’s interactions Hydrogen bonds Solvation effects Loss of entropy Active site waters 8 ADME ADME concerns can be more important than bioactivity. Most of these properties are difficult to predict. • Absorption • Distribution • Metabolism • Excretion 9 Docking Programs • Dock (UCSF) • Autodock (Scripps) • Glide (Schrodinger) • ICM (Molsoft) • FRED (Open Eye) • Gold, FlexX, etc. 10 Evaluation of Docking Programs • Evaluation of library ranking efficacy in virtual screening. J Comput Chem. 2005 Jan 15;26(1):11-22. • Evaluation of docking performance: comparative data on docking algorithms. J Med Chem. 2004 Jan 29;47(3):558-65. • Impact of scoring functions on enrichment in dockingbased virtual screening: an application study on renin inhibitors. J Chem Inf Comput Sci. 2004 MayJun;44(3):1123-9. 11 Cluster Based Computing • Trivially parallelizable – Divide ligand input files – Some programs have specific parallel implementations (PVM or MPI implementations,…) • Commercial licenses are expensive 12 Consensus Scoring • Combining independent scoring functions and docking algorithms can improve results • Most common method: sort using the sum of the ranks of component scores • More sophisticated methods exist Consensus scoring criteria for improving enrichment in virtual screening. J Chem Inf Model. 2005 Jul-Aug;45(4):1134-46. 13 Adding Chemical Informatics Docking results can be improved by using chemical information about the hits. Chemicals which bind the same protein tend to have similar structure. Iterating back and forth between docking and searching large DB. Use other filters and predictive modules (e.g. Lipinski rules) ALGORITHM: 1. Dock and rank a chemical database 2. Create a bayesian model of the fingerprints of the top hits. 3. Re-rank the database based on their likelihood according to the bayesian model • Finding More Needles in the Haystack: A Simple and Efficient Method for Improving HighThroughput Docking Results J. Med. Chem., 47 (11), 2743 -2749, 2004. 14 Visualization • Viewers must be able to scroll through tens or hundreds of small molecule hits • Accessible viewers designed for this problem: – VIDA from OpenEye (free for academics) – ViewDock module of Chimera from UCSF (free, open source) 15 Long-term Goal of Drug Discovery • LTDD (Low Throughput Drug Design) instead of HTVS (High Throughput Virtual Screening) • Common ground: explore virtual space 16 Drug Discovery Case Study: Tuberculosis Tuberculosis Mycobacterium Tuberculosis Very thick, waxy cell wall 18 The Cell Wall: Key to Pathogen Survival Tuberculosis • 7th cause of death • 1 in 3 people have TB • Leading AIDS death cause • Multi-drug resistant • Mycobacterium tuberculosis >30 C fatty acid 10% of genome Sugar Acyl-CoA 6 different ACCase b subunits, AccD1-6 Homologs of PccB Focus on AccD4-6 Cell wall lipids: Important for pathogen virulence, survival and latency 19 Tuberculosis (TB): An old foe 20 The White Death Frederic Chopin 1810-1849 John Keats 1795-1821 21 TB: still a real threat, because….. Its ability to stay alive Multi-Drug Resistant (Super TB strain) 22 The Cell Wall: Key to Pathogen Survival >30 C fatty acid Tuberculosis • • • • • 7th cause of death worldwide 1 in 3 people have TB Leading cause AIDS death Multi-drug resistant Mycobacterium tuberculosis Acyl-CoA 10% of genome Sugar 6 different ACCase b subunits, AccD1-6 Homologs of PccB Focus on AccD4-6 Cell wall lipids: Important for pathogen virulence, survival and latency Substrate specificity for AccD4-6? 23 AccD5 Protein Structures AccD4 (3.3 Å) Solved AccD5 (2.9 Å) AccD6 (2.7 Å) 24 Structure of AccD5 25 Structure-Based Drug Design Enzyme assay AccD5-NCI65828 7 Crystals & Crystal structure 6 [I] = 0.00 [I] = 2.50 [I] = 5.00 [I] =10.00 1/Vo (min-1) 5 3. Combinatorial chemistry 4 3 2 1 0 -1 -0.006 -0.004 -0.002 0 0.002 0.004 0.006 0.008 0.010 0.012 0.014 1/[Malonyl-CoA]um-1 TB ACCase, AccD5 1. High throughput screening 2. Virtual Screening Lead compound 26 The Computational/Experimental Loop Similarity Search AccD5-NCI65828 7 6 [I] = 0.00 [I] = 2.50 [I] = 5.00 [I] =10.00 1/Vo (min-1) 5 4 3 2 1 0 -1 -0.006 -0.004 -0.002 0 0.002 0.004 0.006 1/[Malonyl-CoA]um-1 Assay 0.008 0.010 0.012 0.014 Docking 27 Docking Results • Diversity set (1990) from NCI NCI 65828 NCI 65537 NCI 294153 NCI 105348 NCI 150289 NH2 NH2 OMe OMe H N OMe N N N HO3S N N COCH2 Cl N O2N Me N BrHC CHBr N CO NO2 N H2NO2S O2N OH NCI 172033 NCI 143444 NCI 211736 H2N OH Cl O N Cl Cl Cl CO2H HN OH O NCI 299210 N N N Cl OH N N NH2 N N P Cl Cl Me H2NO2S Cl HO NCI 322921 N 3HCl N NH N H Br OH 28 NCI 65079 (IN2) NCI 622444 (IN1) NCI 65828 (Lead 1) O 300uM SO3H O NCI 4901 (IN3) HO NH2 HO OH N CH3 Cl HO3S HO Cl OH Cl HO3S NH2 N CH3 Cl OH N N NCI 65538 (IN4) NCI 65553 (IN5) SO3H SO3H N N NCI 65554 (IN6) NCI 65555 (IN7) SO3H SO3H 50uM N 300uM N N N 50-100uM N N OH N N N N O N N N NCI 172033 (Lead 2) O O O N HO SO3H OH HO OH SO3H HO3S Cl Cl Cl OH OH NCI 45188 (IN9) NCI 37050 (IN8) Cl SO3H NCI 45618 (IN10) Cl NH2 HO HO3S SO3H Cl Cl OH N NH2 Cl HO3S OH N N N H2N NH2 N N HO3S 29 Structure-Based Drug Design Identified AccD5 Inhibitors AccD5-NCI65828 7 6 [I] = 0.00 [I] = 2.50 [I] = 5.00 [I] =10.00 1/Vo (min-1) 5 4 3 2 1 0 -1 -0.006 -0.004 -0.002 0 0.002 0.004 0.006 0.008 0.010 0.012 0.014 1/[Malonyl-CoA]um-1 KI = 4.7 mM, KGI = ~50 mM New TB drug lead T. Lin, M. Melgar, S. J. Swamidass, J. Purdon, T. Tseng, G. Gago, D. Kurth, P. Baldi, H. Gramajo, and S. Tsai. PNAS, 103, 9, 3072-3077, (2006). US Patent pending. 30 Acknowledgements • Informatics – – – – – – – – – • Liva Ralaivola J. Chen S. J. Swamidass Yimeng Dou Peter Phung Jocelyne Bruand Chloe Azencott Alex Ksikes Ryan Allison Funding – – – – • Pharmacology – Daniele Piomelli • Chemistry – – – – – G. Weiss J. S. Nowick R. Chamberlin S. Tsai K. Shea NIH NSF Sun IGB 31 Two Strategies • Chemical similarity: • Docking: 32 AccD5 • Enzyme necessary for mycolic acid biosynthesis in M. tuberculosis. 33