* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The Pharmacogenetics Clinical Decision Support System is a
Survey
Document related concepts
Drug interaction wikipedia , lookup
Pharmacognosy wikipedia , lookup
Polysubstance dependence wikipedia , lookup
Pharmaceutical industry wikipedia , lookup
Clinical trial wikipedia , lookup
Drug discovery wikipedia , lookup
Prescription costs wikipedia , lookup
Prescription drug prices in the United States wikipedia , lookup
Neuropsychopharmacology wikipedia , lookup
Pharmacokinetics wikipedia , lookup
Adherence (medicine) wikipedia , lookup
Theralizumab wikipedia , lookup
Transcript
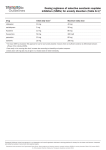
Pharmacogenetics Clinical Decision Support System Med_Inf 406 Eric Chavez, Sean Heffernan, David W. West March 16, 2014 2 Chavez, Heffernan, West Introduction Background The Pharmacogenetic Clinical Decision Support System aims to advance personalized medicine by utilizing preemptive genotyping to help providers make safe and appropriate medication orders. An underlying assumption is that as pharmacogenetic testing becomes more and more common in the future, providers will have a patient’s genotype available before they order medications. The results of pharmacogenetic testing will be stored in a clinical data repository of the electronic health record (EHR). This data will either be automatically uploaded to the clinical data repository from laboratory information systems or from genetic testing vendors, or it will be manually entered into the EHR by staff in a standardized, coded format. Unlike other lab data which change over time, results of genetic testing are stable over a patient’s lifetime. The same data can be reused repeatedly to help providers make decisions about future medication orders. As the field of pharmacogenetics advances, more actionable information can be determined from the results of these tests. Genetic testing is becoming less expensive and more common in clinical practice. This type of testing results in pharmacogenetic data. Pharmacogenetics is the study of how a patient’s genotype and phenotype affect his individual response to medications. Genetic testing can provide information about an individual patient’s ability to metabolize medications through genotyping the cytochrome P450 system. Patients can be classified as rapid metabolizers, normal metabolizers, or slow metabolizers. When multiple choices of medications exist to treat a given medical condition, this information can be used to help providers make decisions about medication selection and dosing. Rapid metabolizers would be expected to show a lower clinical response to a given medication or to require higher than standard dosing strategies. Slow metabolizers would be expected to suffer more toxicity and adverse effects and would need lower than standard doses. Some genetic testing can inform clinicians of the variability in efficacy of certain medications for a patient based on his genotype. Pharmacogenetics is believed to be on the verge of revolutionizing personalized medicine by targeting therapies for individual patients, reducing trial-and-error treatments, and greatly reducing toxicity and adverse effects. This will help to reduce overall healthcare costs. Patients respond to drugs in different ways. There are some groups of patients that will experience severe adverse effects and there are some groups of patients that will not respond at all to certain drugs. Factors that may affect response to drugs include age, gender, disease state, smoking, drug interactions, patient adherence, and genetic factors. It has been shown that genetic factors may account for 20-95% of individual variation in drug response (Kalow, 1998). The amount of variation depends on the class of drug and genetic phenotype of the patient. Pharmacogenetics can provide some explanation of how genetic variants change drug response (Weinshilboum, 2003). 3 Chavez, Heffernan, West Examples of the benefits of personalized medicine using genetic testing include individualized drug dosing, individualized drug selection, and economic savings. These will be illustrated below using the examples of warfarin, hepatitis C, and HIV. Warfarin has a narrow therapeutic index. Inadequate or excessive anticoagulation can lead to increased risk of adverse cardiovascular event or bleeding complication. Warfarin dosage is complicated by individual variability and requires regular monitoring to achieve proper anticoagulation effects. Initial therapy is usually by fixed dosage which is then adjusted based on the patient’s anticoagulation effect, as measured by international normalized ratio (INR). Because genetic factors account for 30-35% of the variability of warfarin response, having genetic test results could lead to a faster time to achieve stable therapeutic dose. In one study, 46% of patients benefitted from genetically informed drug dosing because they required lower or higher doses of warfarin than the standard dosage (Klein, 2009). Another study showed that patients treated with genetically informed dosing had 28% less hospitalizations after 6 months (Epstein, 2010). The ability to more accurately predict warfarin dosing by avoiding under-dosing (risk of coagulation) or overdosing (risk of bleeding) can lead to improved drug efficacy, improved patient safety, and organizational cost savings. Genetic testing can help guide decision making when the clinical effect of a drug is expected to vary according to patient genotype and phenotype. Adverse clinical effects may differ according to phenotype when slow metabolizers accumulate toxic metabolites of a drug. Genetic testing can help guide decision making by steering toward alternative drug treatments or dosing regimens. Patients who are slow metabolizers would benefit from reduced dosing of the standard drug therapy or from taking a different drug altogether. Fast metabolizers may benefit from higher than standard dosing regimens. They would be expected to show less benefit from standard doses. This could delay the efficacy of treatment which means longer time with the illness. Longer time with the illness means more patient suffering and dissatisfaction, potentially more time lost due to a decreased functional state, and more cost to the healthcare system and the overall economy. An example is chronic Hepatitis C which is treated with PegINF-α-2a or PegINF-α-2b and ribavirin. Less than half of patients achieve sustained viral repression with the medications. There is a genetic test that can predict those who will have sustained viral repression based on a polymorphism in the IL28B gene (Ge, 2009). Test such as these help providers know who will respond to a given drug therapy. This can save time, save money, and save patients from adverse drug effects when it is predicted that the benefit is very low. Avoiding inappropriate drug selection can save money. For example if a more expensive drug is shown to be less effective in patients with certain genotypes, the cost of trying that drug may be avoided and alternative medications can be selected. Avoiding ineffective drug regimens can save money by reducing patient suffering and dysfunction and by speeding time to recovery. Avoiding serious adverse drug reactions can save money by reducing hospitalizations or office visits and by reducing patient morbidity. It may be more costeffective to use generic drugs that are associated with adverse effects in some genotypes 4 Chavez, Heffernan, West than to use a more expensive drug without the potential adverse effects for everyone, if genetic testing is used to help select appropriate patients for the generic drug. An example is in the treatment of HIV with abacavir. It is estimated that 5-8% of patients will develop a hypersensitivity reaction to the medication in the first 6 weeks of treatment. Rechallenging with the medication after initial treatment may result in worsening of the reaction or even death. Genotyping can reduce this risk by predicting who will have the hypersensitivity reaction. In one study, screening patients for the genotype associated with the reaction and not giving those patients abacavir lead to complete elimination of the reaction in the study population (Gazzard, 2008). In an analysis of cost-effectiveness, it was found that genetic testing and treatment with abacavir when appropriate was less costly than completely avoiding abacavir and using a more expensive alternative medication, tenofovir (Kauf, 2010). The field of pharmacogenetics is rapidly developing. The majority of practicing physicians lack any formal training in the field, and most do not know how to incorporate this information into their practices to improve safety and outcomes for patients (Baars, 2005). It would be impossible for providers to memorize all of this information as more and more clinically relevant information and actionable guidelines are added. The pharmacogenetics clinical decision support system can help to solve this problem by using a just-in-time tool that brings actionable recommendations to providers at the point of care. This system will integrate into the computerized provider order entry (CPOE) module of the EHR and make recommendations to alter drug choice or alter drug dose when providers attempt to write medication orders. In order to reduce alert fatigue the system will issue alerts only when strong clinical guidelines exist in the pharmacogenetics literature. There are few studies that report on the development and implementation of pharmacogenetic clinical decision support systems. In 2009 Deshmukh, et al. studied two different methods of presenting genetic test results to a Cerner clinical decision support system. They used the CYP2C9 gene and used a data model of either single nucleotide polymorphisms (SNP) or alleles as the discreet test reports. Both models worked for the clinical decision support system (CDSS). The allele-based system was easier to write decision rules for, and the SNP-based system allowed for retention of more discreet data that could possibly be used and represented in different ways in the future. Overby (2010) reported on the feasibility of collecting and standardizing pharmacogenetic data from various sources, turning that data into computable form, and writing decision rules based on the data. In 2012 Overby, et al. expanded on their previous work and described a prototype of semiactive and active CDSS using translated pharmacogenetic data to drive rules-based decisions in a Cerner EHR environment. The semi-active component allowed providers to look up genetic testing information about a specific drug while the active component offered recommendations for therapy change or dose adjustments when providers attempted to order a medication. Bell (2014) et al. describe an active CDSS in which genetic test results are captured and encoded in the EHR problem list. Alerts were generated when providers attempted to prescribe a medication that matched a high risk phenotype in the problem list. In a review article, Welch and Kawamoto (2013) report on six different studies that use 5 Chavez, Heffernan, West genomic data as part of CDSS to guide medication management for warfarin and antiretroviral medications. Stakeholders, Goals, and Objectives For the purposes of this project we will assume that our target healthcare organization is a large multispecialty medical center with both inpatient and outpatient services. The stakeholders for the project are visualized in “onion-ring” format in figure 1. The innermost circle contains the product which is the pharmacogenetics clinical decision support system. The next circle contains the business system which are those stakeholders who do the work in developing the product and who have operational roles. The next circle contains the business who are the stakeholders who will benefit directly from the product. The outermost circle contains the wider environment of stakeholders who will be indirectly affected by the product. The Clinical Decision Support Committee is the main driving force for the CDS initiative. The committee may or may not include several of the other key stakeholders listed below. The committee is responsible for developing the goals and objectives for the CDS system and for evaluating the science behind the system and the evidence that it will provide benefit for patients and for the organization. The committee solicits feedback from key stakeholders to ensure that the project is feasible. Overall, the committee guides the development, implementation, evaluation, and monitoring of the CDSS. It is important for the CDS committee to engage all stakeholders through every phase of the CDS lifecycle. Clinical leadership is necessary to evaluate the need for the CDSS and the medical evidence to support the system. Clinical leaders need to be involved in every stage of the project development and implementation to give feedback on how provider workflows are effected and how patient care is impacted. Notable among the clinical leaders is the Physician Champion who plays a key role in the development and implementation of the CDSS. Usually the champion is well respected among his/her peers, has excellent communication skills, remains active clinically, and helps to grow support by spreading enthusiasm. Technical leadership is necessary to translate the clinical ideas into a functional operating plan in the computer information system. It will be necessary to involve members of the EHR and CPOE committees in the technical leadership since this CDSS uses the EHR and CPOE system as its point of interaction with providers. A vendor representative will be included to share ideas about problems and solutions that other organizations have faced when implementing similar CDS systems. The administrative leadership is ultimately responsible for approving the CDS project. They are responsible for determining whether or not the CDSS meets the goals of the organization, for evaluating the return on investment, and for fundraising or budgeting for the project. Administrative leadership will be indirectly affected as it is hoped that outcomes will improve and cost savings can be realized for the organization. 6 Chavez, Heffernan, West Genetic testing labs will need to ensure that results of genetic testing can be automatically integrated into the EHR and CPOE systems End users are the front line clinicians who will be using the CDSS on a daily basis. They should be active in all phases of the project to ensure buy-in and adoption. End users can give valuable feedback during various stages of the project to make adjustments for work flow in order to help the CDSS achieve success. Resistors and detractors among the end user group should be included in the planning process so that potential problems can be addressed early. Patients may be involved during the project design and evaluation to give feedback and to help spread support for the system. They should be made aware of the potential benefits and risks of the new CDSS. It is hoped that ultimately patients will be the primary beneficiaries of the new system with improved outcomes and safety. Figure 1. Stakeholders 7 Chavez, Heffernan, West Organizational goals and clinical goals of the project have to do with improving patient lives, improving efficiency of physician workflows, and saving healthcare dollars. These goals are displayed in figure 2. More specific objectives for the project are shown in figure 3. These objectives may be used as the metrics for a CDSS evaluation program. Figure 2. Organizational and clinical goals. Improve patient safety, outcomes, and satisfaction • Minimize harmful drug effects • Optimize treatment for individual patients • Reduce length of patient morbidity and loss of function by selecting effective drug regimens more rapidly Improve physician effectiveness and efficiency • Make more effective decisions in regards to drug selection and dosing • Predict optimal dosing for drugs with a narrow therapeutic index • Avoid trial-and-error treatments Reduce healthcare costs • Develop cost-effective treatment by selecting the most effective and appropriate drug for each individual patient 8 Chavez, Heffernan, West Figure 3. Specific objectives. Improve patient safety reduce the number of adverse drug events Improve dosing regimens Decrease from standard dosage or select alternative medication for patients with slow metabolizer phenotypes Select most effective therapy Increase the number of orders written for drugs shown to be effective for an individual patient’s genotype Improve treatment response times Improve the time to achieve stable therapeutic dosing for drugs with narrow therapeutic windows (i.e. warfarin) reduce the number of hospitalizations and office visits due to adverse drug events Increase from standard dosage or select alternative medication for patients with rapid metabolizer phenotypes Reduce the number of orders written for drugs shown not to be effective for an individual patient’s genotype Improve time to achieve reduction or resolution of patient symptoms by more rapidly selecting the most effective treatment regimen Information System Inventory The pharmacogenetics clinical decision support model shown below in figure 4 represents current and future system states, which will easily generalize to other clinical environments, allow providers to enter metabolizer status when ordering a drug, and support reporting and interpreting genetic test results. The CDS will 1) integrate genetic testing results into the Clinical Data Repository for use in the existing CPOE system and 2) store the results from genetic labs in the CDS Knowledge Base where rules will be applied for specifying results and preparing them for presentation to users via the Clinical System Interface. 9 Chavez, Heffernan, West Figure 4. Clinical Decision Support Model The EHR Application Environment supports the functional capabilities of the decision support system. Discrete data are stored within the Clinical Data Repository and the Clinical User can interact with the Clinical System Interface to view data and order medication (CPOE), which may trigger decision support implemented in the repository. Interventions and offered medication choices display to the user through the Clinical System Interface. The Clinical Decision Support Module implements the user interface features, providing methods for transforming inputs (genetic test results or medications being ordered) into patient-specific output (pharmacogenomics links to e-resources or alert messages). Imbedding the decision support tool in the EHR will allow the use of pharmacogenomics knowledge in making prescribing decisions. System Inventory Needed Acquisitions One File System for the Result Specification XML Repository Two Databases (one each) for the Pharmacogenomic Knowledge Base and CDS Knowledge Base Two application servers for deploying the new databases One Rules Engine for retrieval and calculation of genetic tests and related medication options One web server for linking the CDS to the vendor’s e-resources Workstations o 10 new workstations to be deployed across Ambulatory, Inpatient, Behavioral Health, Laboratory, and Pharmacy Services o Note: Although the setting for deploying the CDS is a large urban hospital, the intended user base of the software is limited, so the number of new workstations needed is minimal 10 Chavez, Heffernan, West Personnel o Physicians o Lab Technicians o Pharmacists o Nurse Practitioners o Software Administrators o IT Technicians o Vendor Liaison Software o System Interface Software o Database Deployment Software o Rules Design/Maintenance Software o Web Services Software Intervention Selection and Workflow Opportunities Medication dosing has drawn the attention of clinical decision support systems for decades, especially after the Institute of Medicine (IOM) published its report, “To Err is Human” in 1999 in which medication errors were most prominent among all types of errors which occur in the health care setting (Kohn, 1999). The implementation of computerized physician order entry (CPOE), even in its simplest form, circumvents errors related to legibility, but as a change in workflow, there has been concern in the literature that other types of errors not previously seen are being introduced. Since the IOM published its report, clinical decision support around dosing has become more and more sophisticated. This support is predicated on the enhancement in CPOE to require granular elements of the order to be entered. These elements include dose, dose unit, frequency, and route at a minimum. Third party expert content providers have emerged to provide dosing support for virtually any FDA approved medications where granular CPOE has been implemented. Dose range checking is the most basic type of decision support, essentially alerting the prescriber if the single or daily dosage falls outside of a standard range. Condition-based defaults and weight-based dosing are other forms of clinical decision support that start to take other clinical factors into account when providing support. Lab studies such as creatinine can be incorporated to provide further dosing support. Genetic studies that help predict metabolism and/or effectiveness of certain drugs is a new and growing field that shows great promise for being integrated into clinical decision support systems. Many physicians are now ordering these tests to assist their dosing decisions. The proposed CDSS will integrate genetic testing results into the clinical data repository in a manner that will effectively provide support at the point of order entry with the goal of achieving a therapeutic, but non-toxic level of medication. Genetic tests are an excellent example of “patient-level” data in that the results do not change over time. Family history and birth history are other examples of patient-centered data. Once the data is captured, it 11 Chavez, Heffernan, West does not need to be recaptured. It does need to be readily accessible at the point of care. Therefore, avoiding duplication of genetic testing would be an additional benefit of the proposed CDSS, though not its primary objective. A number of system features will be important to integrate this information into medication dosing decisions. The results originating in the genetic labs need to be normalized into a value that can guide and alert dosing. The medication repositories supplied by third party domain expert systems need to include this relationship with genetic testing in a manner similar to age, weight, and renal function based dosing elements. Finally, the health care organization’s EHR needs to support and integrate these various elements into a useful presentation to the prescriber on pharmacogenetic-based clinical guidelines. Change Management Plan Clinical decision support systems ultimately strive to modify and align behaviors to best practices that will achieve optimal outcomes. Attempts at behavior change often encounter resistance. The likelihood that behavior change will occur tends to correlate positively to the perceived value of the change and negatively to the pain of adoption. The change management plan will address both of these principles. To boost the perceived value proposition prior to implementation, data will be gathered that helps define the opportunity to improve care. This will entail statistics on the frequency of medication orders for which genetic data could alter management. One can then apply knowledge about the frequency of genetic phenotypes in the population to infer the frequency with which medication dosing could be enhanced to avoid lack of efficacy due to under dosing or toxicity due to overdosing. To mitigate the pain of adoption, the effort required to incorporate the genetic information into the dosing decision needs to be easy to find and presented in a manner that can be quickly converted into a dosing modification. Reports that are composed of free text, even if presented in tabular form, need to be stored in relevant granular data elements. The correct genetic elements need to be connected to the drugs for which a given gene will have an impact. The connection must classify the effect of the gene, and then offer to alter the dosing accordingly. In cases where the genetic data suggests a pure lack of efficacy, the connection between the genetic phenotype and the drug should lead to an offering of alternatives that would have greater efficacy. Post-implementation efforts can continue to try to reinforce the value proposition. This can take the form of queries that report the frequency of adoption of modified dosing in accordance with genetic data. This information can be trended and shared as an organization quality improvement objective, with more detailed “drill-down” data to demonstrate inter-provider or inter-department variability. Areas of success can be examined closely to see if practices exist that could be replicated to areas of lower success. The opportunity to offer rationales as to why suggested dosing was not accepted will serve 12 Chavez, Heffernan, West to enhance clinical decision support if other variables can be introduced that improves accuracy. A continued iterative process that focuses on enhancing one or both of these principles (increased value proposition, decreased pain of adoption) should serve to solidify the change in the organization as a whole. System Design The pharmacogenetic CDSS enables the use of pharmacogenetic data each time a relevant medication is prescribed as part of pre-emptive testing and stores the results in the EHR before a drug is prescribed. Accordingly, the pharmacogenetic CDSS advances personalized medicine by addressing the inherited variation in drug responses by maximizing the drug efficacy and minimizing toxicity for individual patients. The CDSS can also reduce costs associated with inappropriate drug treatments or serious adverse drug reactions. Included in the System Design is a discussion of the: 1. Components of the System Architecture, which specifies the functional requirements of the CDSS 2. Pharmacogenetic Use Case for Clinical Assessment and Genetic Testing 3. Conceptual Model for the Pharmacogenetic CDS in terms of Client/Server and WebBased implementations Components of System Architecture The components of the CDSS architecture are best represented by a specification of its functional requirements. Below in figure 5 is a depiction of the system’s functions and the interaction between processes and outputs. 13 Chavez, Heffernan, West Figure 5. System architecture. Pharmacogenomics CDS Functional Requirements Triggers Data Elements Interventions Offered Choices PGx eResources Events that cause a decision support tule to be invoked Used by a rule to make inferences Actions a decision support module can take Order Entered Lab Result Stored Lab Result Drug List Diagnosis Age Gender History Show Guidelines Pick Lists Data Entry Template Cancel Current Order Defer Warning Override Rule/Keep Order Edit Problem List Notify Logs Deriving CDS Rules FDA Info on Genomic Biomarkers Evidence Based Synopses CPIC Guidelines Article Searches 14 Chavez, Heffernan, West Pharmacogenetics Use Case The use case below shows how personal health and family health history information is exchanged with genetic testing information between consumers and clinicians based on two scenarios: 1. Clinical Assessment 2. Genetic Testing PGx CDS Use Case Perspective / Rule Clinician Clinical Assessment Testing Lab 1. Construct Medical History 2. Evaluate Relevant Genetic Test Scenarios 3. Order Test 1. Receive Test Orders 2. Prepare for Test Genetic Testing 3. Perform Test 4. Develop / Transmit Lab Result 5. Provide Supplemental Information 4. Receive Lab Tests 5. Perform Interpretation 6. Care Planning These scenarios demonstrate how genetic test results are represented in EHR components of a pharmacogenetic CDSS and the three phases of performing genomic tests: 15 Chavez, Heffernan, West 1. Pre-Analytic Phase to determine which genetic test is appropriate 2. Analytic Phase where samples are analyzed 3. Post-Analytic Phase where results are reported and interpreted Intervention Specification The pharmacogenetic CDSS is an alert-type intervention. Triggering occurs when a provider uses a CPOE system to prescribe a medication for a patient. The trigger is data-driven since a query will be performed in the clinical data repository to determine if genetic testing results exist for the patient. If such results are available, then a query will be performed to match the drug to a known drug-gene pair in the pharmacogenomics knowledge base. Decision rules in the CDSS will be based on clinical practice guidelines that are published by the Clinical Pharmacogenetics Implementation Consortium (CPIC). Founded in 2009, the CPIC is a group of organizations that develops and publishes clinical practice guidelines for actionable prescribing decisions based on the results of genetic testing (Caudle, 2014). CPIC guidelines are written for drug-gene pairs where evidence clearly shows that genetic variations affect efficacy or risk and where prescribing recommendations such as changing dose or changing drug can be made. CPIC guidelines are written in accordance with Institute of Medicine standards for the development of clinical practice guidelines. The CDSS will use the following logic (see figure 6). If genetic test results are available for the patient and a guideline exists for a drug-gene pair, then a modal alert box will immediately be displayed for the prescriber in the CPOE user interface. The alert will inform the prescriber of the patient’s genotype or phenotype and the expected variation in efficacy or risk of adverse effect for the drug. Based on data from the guideline contained in the pharmacogenomics knowledge base, the alert will offer specific advice for dosage adjustment or alternative drug treatment as appropriate. An infobutton will be included in the alert box which will link to more detailed information and evidence for the recommended changes in treatment. The prescriber will then have an option of canceling the order, modifying the order, or overriding the alert and proceeding with the order. When overriding an order the provider will need to justify the override by selecting a reason from a dropdown list. If no genetic test results exist for the patient, if there are no drug-gene matches, or if there are no important actionable clinical practice guidelines, then no alert will be triggered and workflow will proceed as normal. This logic step will help to prevent alert fatigue by presenting providers with an alert only when actionable guidelines exist for a drug-gene pair. Because the field of pharmacogenetics is rapidly changing, the pharmacogenomics knowledge base of the CDSS will need to be updated with new practice guidelines from the CPIC at frequent intervals. We will begin by updating these data sources every three months. The CDSS will help the organization to meet the stated goals and objectives by helping providers avoid prescribing medications for patients when a known risk exists based on 16 Chavez, Heffernan, West genotype. Adverse drug reactions will be avoided by lowering doses or by choosing alternative medications. Cost savings can be realized by avoiding adverse drug reactions, by avoiding the prescription of drugs known not to be effective for a particular genotype, and by selecting medications known to be more effective for a particular genotype. Figure 6. Pharmacogenetic Clinical Decision Support System Logic 17 Chavez, Heffernan, West User Interface According to a report from the HIMSS Usability Task Force (2009) user interface design can be a key component for ensuring that clinical decision support systems and other health information technology products meet their intended goals of improving patient care and safety. This is because a well-designed user interface will enhance provider adoption and lead to routine clinical use. The design of a CDSS must be satisfying to the user, must be effective in helping the user do his work, and must be effective at guiding the user towards making the recommended decision. It is wise to include end users in the planning stages of interface design so that changes can be made in an iterative fashion in order to maximize acceptance. The HIMSS Task Force notes several key principles for interface design. The interface should be simple and concise and present information in a way that is natural and familiar to the target end user. The design should aim to minimize the cognitive load of the user who is usually a provider under time pressure with multiple demands on his attention. Efficient interactions with a minimum number of steps to complete tasks will enhance user satisfaction. The design should provide feedback along the way in order to allow a user to learn the system, especially since opportunities for formal training can often be limited. In a review on the topic of interface design, Horsky et al. (2012) recommend that consistency between the CDSS and the larger EHR be maintained. This includes visual formats, terminology, nomenclature, and processes. They recommend presenting advice with short explanations and links to more detailed evidence. This way, users will develop trust in the decision support system over time since they will know where recommendations are coming from rather than being confronted with a “black box” that issues advice. Clinical decision support systems should offer guidance and recommendations rather than commands. Physician are much more likely to follow recommendations from decision support systems when they are offered alternatives to carry out rather than a hard stop or a recommendation against a particular action. Recommendations should be written in succinct and unambiguous language that can be understood after reading it in one pass. Alerts should be contained on one page with the shortest amount of text possible followed by an actionable item. The interface design of the pharmacogenetics CDSS will follow the principles outlined above as well as the four A’s recommended by Kanstrup (2011): “Attention, All in one, At a glance, and At hand.” As mentioned in the intervention specification section above, the CDSS will engage when a provider uses the CPOE system to write an order for a medication. Input to the CDSS from the provider will come directly from the CPOE. If an alert is triggered based on the decision support system logic, a modal alert box will be immediately displayed. This alert box will have a visual design that is consistent with the rest of the EHR with the exception that it will be of a different color in order to draw “attention.” 18 Chavez, Heffernan, West The alert box will contain all relevant information about the drug-gene pair and clinical guideline recommendations “all in one” box “at a glance.” Verbiage will be minimized to present the provider with a succinct explanation. Hyperlinks and infobuttons will be included in the alert box so that providers can immediately have access to more detailed information regarding the drug-gene pair and the clinical guidelines if they wish. Immediately following the recommendations, providers will have “at hand” the ability to override, modify, or cancel the order. If providers chose to override the order, they will be asked to justify the override using a drop down list which will include the following reasons: “patient previously tolerates this therapy,” “dose or therapy adjustment already made,” “guideline not relevant for this patient,” “genetic testing is erroneous,” “risks and benefits have been considered/patient will be carefully monitored,” and “other.” Providers will also be able to enter free text into the override justification box. This information can be used to later fine-tune the sensitivity of alerts. If providers choose to modify the order, they will be taken back to the CPOE screen with the original drug order information previously entered intact. Providers may then change the dose or change the drug. If they choose to cancel the order, they will be taken back to the CPOE screen with a blank medication order entry form. The following are two illustrations of the user interface. Example 1: The provider attempts to order abacavir for a patient via the CPOE. The CDSS searches the clinical data repository and finds that genetic test results exists for the patient. The CDSS then searches the pharmacogenetics knowledge base for abacavir and finds there is a related drug-gene pair, abacavir and HLA-B. It then looks for HLA-B test results for the patient. If those test results exist, it then looks for a clinical guideline for abacavir/HLA-B. It finds that clinical guidelines do exists. If the patient has the HLAB*57:01 allele then the clinical guideline from CPIC recommends alternative therapy due to high risk for hypersensitivity reaction. 19 Chavez, Heffernan, West The clinical decision support system will then display an alert box shown in figure 7. Figure 7. Example clinical decision support alert box for abacavir. PGx Medication Alert This patient has genetic test results indicating the presence of the HLA-B*57:01 genotype. Patients with this genotype are at significantly increased risk of hypersensitivity reaction with Abacavir. There are strong recommendations to avoid Abacavir for this patient. Cancel Order Modify Order Override Recommendation More Information OK The hyperlink labeled “more information” in the alert box will link to the clinical guidelines describing the relationship between abacavir and HLA-B and the evidence regarding risk of hypersensitivity reaction and recommendation to avoid using the medication in patients with this genotype. The provider may then chose to override (for example if the patient has already been taking abacavir with no problems). If he doesn't override, then he may cancel the order or modify it. Example 2: The provider attempts to order codeine for a patient. The CDSS searches for genetic test results. It then searches for codeine and finds the drug-gene pair codeine and CYP2D6. It then looks for CYP2D6 results for the patient and looks for clinical guidelines for Codeine/CYP2D6. The system then reports the clinical guideline recommendations based on the patient's phenotype as shown in figure 8. 20 Chavez, Heffernan, West Figure 8. Example clinical decision support alert box for codeine. PGx Medication Alert This patient has genetic test results indicating a poor metabolizer phenotype for CYP2D6. The patient is likely to have greatly reduced morphine formation after administration of Codeine leading to insufficient pain relief. Avoid using Codeine due to lack of efficacy. Consider alternative analgesics such as morphine or a nonopioid. Tramadol should also be avoided for this patient. Cancel Order Modify Order Override Recommendation More Information OK Conceptual Model for the Pharmacogenetics CDSS Based on the System Inventory, the summary below of each model component is organized by method of implementation: Client/Server or Web-based. Client/Server Based The CDSS EHR Application Environment supports the functional capabilities that were evaluated by the group and shown in the Components of System Architecture diagram. Discrete data elements are captured within the Clinical Data Repository. The clinical user can interact with the Clinical System Interface to view data elements and perform clinical tasks, e.g., ordering a medication. Such events will then trigger decision support to fire. Triggers are implemented in the Clinical Data Repository. Interventions and Offered Choices are displayed to the clinical user within the CDSS Clinical System Interface. Web-Based The Clinical Decision Support (CDS) Module implements the IF-THEN rules that are defined in the Rules/Execution Engine and based on each patient’s genotype data combined with phenotype data stored in the CDS Knowledge Base. The database and rules together relate actionable genotype and phenotype pairs to genome-informed advice messages. If defined rules are met, CDS is delivered in real-time at the point of care through the EHR. 21 Chavez, Heffernan, West The CDS Module also implements the User Interface features (information, warnings, recommendations) and provides the methods for transforming input parameters (i.e., data about genetic test results and the medication being ordered) into a specific output (i.e., Pharmacogenetic links to e-Resources or an Alert message). Glossary of Terms Clinical Data Repository Consolidates data from multiple clinical sources and contains lab results, patient demographics, pharmacy information, radiology reports, admission / discharge summaries code vocabularies, progress notes, etc. Pharmacogenetic Knowledge Base Sponsored by NIH (pharmaKB.org), this knowledge base is the curator of research information on genetic variations in drug responses, e.g., how genetic factors influence a patient’s response to drugs Genomics Vendor Knowledge Base Contains genotyping and genetic testing results for patients CDS Knowledge Base Uses an inference engine to facilitate presentations of drug interactions, dosing suggestions, alerts, and diagnoses by applying rules to patient data Knowledge Engineering Knowledge acquisition While the interaction relationship within a drug-gene pair is likely to remain stable once established, the quantity of relevant genotypes is growing rapidly. It would be beyond the scope of our healthcare organization to try and directly acquire and maintain a compendium of these pairs from primary pharmacogenetic literature. Fortunately, as previously noted, the Clinical Pharmacogenetics Implementation Consortium (CPIC) represents a collaborative effort of significant academic breadth which composes clinical guidelines for drug-gene pairs in a format that complies with Institute of Medicine (IOM) standards. The “acquisition” process in this case is actually a selection process where the organization chooses the guidelines that are considered relevant for its needs. It may be tempting at first to think about a wholesale adoption of guidelines, but in the long run, a process of “taking ownership” by selecting and applying the guideline is an important one to aid in the adoption of these new guidelines. The Clinical Decision Support Committee (CDSC) for the organization is central to have the organization take ownership of these guidelines. They would be able to assess the needs of the organization and leverage the resources necessary to approve and oversee a robust 22 Chavez, Heffernan, West process of acquisition and adoption. There would be merit in considering a subcommittee for this activity to handle the anticipated volume of new guidelines emanating from CPIC. Knowledge representation Taking “ownership” of a drug-gene pair guideline from the CPIC will require more than a simple title/description of the relationship. These relationships need to be parsed into a consistent set of elements which will facilitate their incorporation in to the organization’s CDSS. The proposed structure can be seen in Appendix A using a table of drug gene pairs where three layers are inferred. The top layer represented by the first two columns in green are the drug-gene pair descriptions. The last three columns in white (Phenotype, Guideline Recommendation, Recommendation Type, etc.) would have a potential many-to-one relationship with the pair as indicated by many pairs being seen more than once. The Genotype column in yellow has a many-to-one relationship with the Phenotype recommendations since a given phenotype could derive from multiple genotypes. Ultimately these columns would correlate with data entry points for the clinical decision support system to allow the elements to present in the user interface in the method previously described. Other meta-data will also be helpful to capture including the date approved by the CDSC and the primary organizational researcher. A future review data could also be entered to facilitate reporting and queuing of guidelines needing review and renewal. These future reviews would occur at a pre-determined interval to determine if new literature has presented any refinements or recommendations, perhaps around lab monitoring. Evaluation The Pharmacogenetics Clinical Decision Support System is a relatively new model and requires both validation and verification. The validation phase will be a robust testing program before implementation. This testing will be handled in three segments called unit testing, scenario testing and integrated testing. 1. Unit Testing: During this phase of testing, each drug-gene pair result is tested by having the particular genetic result entered on a test patient in a test domain. The corresponding drug component would then be entered to validate that the clinical decision support that fires works as designed. It should offer accurate information about how to respond. The cycle here is relatively simple, but the volume of cycles is directly tied to the volume of drug-gene pairs being implemented. 2. Scenario Testing: During this phase of testing, a testing script is used to elicit specific scenarios that may challenge the robustness of the CDSS. A sample testing script is attached in Appendix B and would likely be enhanced by the project team to add additional scenarios desired by stakeholders. 23 Chavez, Heffernan, West 3. Integrated Testing: Unit testing will occur within the confines of the CDSS and the EHR. Integrated testing will engage all of the necessary systems to test the entire flow of the CDSS. This will include the addition of relevant drug-gene pairs to the relevant knowledgebase, interfacing of gene results from the laboratory information system to the EHR and completion of medication order including transmittal via e-prescribing. The goal is to detect unintended consequences to related downstream systems. Verification will commence after implementation and will utilize a defined set of metrics to assess impact. There will be pre-verification analysis to determine a baseline for metrics of interest: Medication ordering frequencies for each medication where a drug-gene pair exists Dosing variability for each medication where a drug-gene pair exists Adverse events for each medication where a drug-gene pair exists Frequency and count of dose changes, including discontinuation and starting, for each medication where a drug-gene pair exists The baseline metrics above will be continued after implementation, and once the CDSS is implemented, the following metrics will be added: Alert frequency for absence of relevant gene testing Percentage of corresponding medications prescribed without prior genotype information Alert frequency for therapeutic modification due to relevant drug-gene pair Response frequencies to alert (override, change medication, alter dosing) Percentage of medications prescribed where dosing incorporated suggested dose modification Specific drug-gene pairs may utilize metrics of enhanced efficacy. In the case of warfarin based products, median time to therapeutic effect as measured by Prothrombin Times (PT) and their related INR. Not all medications will have this type of a metric available. Discussion Several previous studies have reported on the feasibility of incorporating pharmacogenetic data into CDSS and some have reported on the deployment of such data in CDSS in pilot studies or in a limited scope with decision support for a particular drug or class of drugs (Bell 2014, Bielinski 2014, Deskhmukh 2009, Goldspell 2013, Overby 2010 and 2012). Here we have presented ideas for a wider scope of pharmacogenetic CDS. Given the underlying assumption that patients will already have the results of genetic testing available in the 24 Chavez, Heffernan, West clinical data repository (preemptive genotyping), this system will have limited value to providers and patients in organizations where the prevalence of such genetic testing is low. It is expected that in the future genetic testing will be less expensive and more ubiquitous. As this occurs, this CDSS will be at the ready to help make use of that data for the benefits of those patients who have had genetic testing done. The interpretation of genetic testing can be confusing, especially for providers with limited training and limited time. Evaluating the vast amount of research in pharmacogenetics is a daunting task. For this reason, our system employs guidelines that have already been vetted and approved by the CPIC. The pharmacogenetics CDSS has been designed keeping in mind the three pillars of the CDS Roadmap outlined by the American Medical Informatics Association as reviewed by Lyman (2010). “Best knowledge available when needed” means that this system presents only CPIC approved guidelines with actionable decision support to providers at the point of care when they attempt to prescribe a medication. This fits into the standard workflow and should acceptable for providers. “High adoption and effective use” means that alerts will be kept to a minimum in order to prevent alert fatigue. Scientific evidence of each recommendation will be available to help build trust and support for the system. “Continuous improvement of knowledge and CDS methods” means that the CDSS will be evaluated regularly and refined according to provider and organization needs. The knowledge base will be updated at frequent intervals using information from the CPIC guidelines. Alerts will be fine-tuned based on the number and reasons for overrides. Scalability is an important factor for the CDSS as pharmacogenetic knowledge is constantly evolving. Utilizing decision rules based on the CPIC guidelines will help to ensure scalability in the future because staff responsible for maintaining the CDS knowledge base and decision rules will be able to find this information in one place. New information can be taken from the CPIC and quickly integrated into the CDSS. Although this system was design to engage prospectively at the point of care when providers attempt to prescribe medications, it could be expanded. The system could potentially be used retrospectively to scan all patient data including active medications and genetic testing results. If the system finds patients who have been prescribed medications which have a drug-gene match and an actionable clinical guideline, it could inform the prescribing physician via email about the guideline. The provider could then make an adjustment to therapy or dosing if clinically relevant. The system could also be employed as a research tool. Providers could access the system to search for drug-gene matches for patients prior to accessing CPOE to prescribe. This may offer some benefit in that providers can discuss treatment options with patients, considering risks and benefits. As providers and healthcare organizations strive to offer ever more sophisticated treatment plans based on personalized medicine, the pharmacogenetics clinical decision support system will be an essential tool. It will help providers by gathering and presenting information that is relevant to an individual patient at the point of care. It will help 25 Chavez, Heffernan, West disseminate to providers the vast amount of pharmacogenetic data available and therefore aid in the more rapid adoption of clinical guidelines based on pharmacogenetic data. This system has the potential to greatly improve patient safety and reduce healthcare costs. 26 Chavez, Heffernan, West References Baars M. J. (2005). Deficiency of Knowledge of Genetics and Genetic Tests among General Practitioners, Gynecologists, and Pediatricians. Genetics in Medicine. 7(9), 605610. Bell G. C. (2014). Development and Use of Active Clinical Decision Support for Preemptive Pharmacogenetics. Journal of the American Medical Informatics Association. 21, e93-e99. Bielinski S. J. (2014). Preemptive Genotyping for Personalized Medicine: Design of the Right Drug, Right Dose, Right Time--Using Genomic Data to Individualize Treatment Protocol. Mayo Clinic Proceedings. 89(1), 25-33. Caudle K. E. (2014). Incorporation of Pharmacogenetics into Routine Clinical Practice: the Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline Development Process. Current Drug Metabolism. 15(1), 1-9. Deshmukh V. G. (2009). Efficiency of CYP2C9 Genetic Test Representation for Automated Pharmacogenetic Decision Support. Methods of Information in Medicine. 48(3), 282-290. Epstein R. S. (2010). Warfarin genotyping reduces hospitalization rates results from the MM-WES (Medco-May Warfarin Effectiveness study). Journal of the American College of Cardiology. 55, 2804-28-12. Gazzard B. G. (2008). British HIV Association Guidelines for the treatment of HIV-1infected adults with antiretroviral therapy. HIV Medicine. 9, 563-608. Ge D. (2009). Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 461, 399-401. Goldspeil B. R. (2013). Integrating Pharmacogenetic Information and Clinical Decision Support into the Electronic Health Record. Journal of the American Medical Informatics Association. 0, 1-7. HIMSS Usability Task Force (2009). Defining and Testing EMR Usability: Principles and Proposed Methods of EMR Usability Evaluation and Rating. Horsky J. (2012). Interface Design Principles for Usable Decision Support: A Targeted Review of Best Practices for Clinical Prescribing Interventions. Journal of Biomedical Informatics. 45(6), 1202-1216. Hulse M. (1997). User-Interface Design of a Web-Based Clinical Decision Support System. Proceedings of the American Medical Informatics Association Annual Fall Symposium. p. 951. 27 Chavez, Heffernan, West Kalow W. (1998). Hypothesis: comparisons of inter- and intra-individual variations can substitute for twin studies in drug research. Pharmacogenetics. 8, 283-289. Kanstrup A. M. (2011). Four Principles for User Interface Design of Computerised Clinical Decision Support Systems. Studies in Health Technology and Informatics. 166, 65-73. Kauf T. L. (2010). Economic efficiency of genetic screening to inform the use of abacavir sulfate in the treatment of HIV. Pharmocoeconomics. 28, 1025-1039. Klein T.E. (2009). Estimation of the warfarin dose with clinical and pharmacogenetic data. New England Journal of Medicine. 360, 753-764. Kohn L. T. (1999). To Err is Human: Building a Safer Health System. Washington, DC: National Academy Press, Institute of Medicine. Lyman J. A. (2010). Clinical Decision Support: Progress and Opportunities. Journal of the American Medical Informatics Association. 17(5), 487-492. Osheroff J. A. (2012). Improving Outcomes with Clinical Decision Support. Chicago, IL: Healthcare Information and Management Systems Society. Overby C. L. (2010). Feasibility of Incorporating Genomic Knowledge into Electronic Medical Records for Pharmacogenomic Clinical Decision Support. BMC Bioinformatics. 11(Suppl 9), s1-9. Overby C. L. (2012). Developing a Prototype System for Integrating Pharmacogenomics Findings into Clinical Practice. Journal of Personalized Medicine. 2(4), 241-256. Overby C.L. (2011) An Evaluation of Functional and User Interface Requirements for Pharmacogenomics Clinical Decision Support. IEEE International Conference on Healthcare Informatics. Ross S. (2012). Promises and Challenges of Pharmacogenetics: An Overview of Study Design, Methodological and Statistical Issues. JRSM Cardiovascular Disease. 1(2), 1-13. Weinshilboum R. (2003). Inheritance and drug response. New England Journal of Medicine. 348, 529-537. Welch B. M. (2013). Clinical Decision Support for Genetically Guided Personalized Medicine: a Systematic Review. Journal of the American Medical Informatics Association. 20(2), 388-400. 28 Chavez, Heffernan, West Appendix A. Pharmacogenetics Knowledge Base Table Drug Gene clopidogrel CYP2C19 clopidogrel CYP2C19 Genotype Phenotype Guideline Recommendation *1/*2, *1/*3, intermediate consider alternative therapy such as prasugrel or ticagrelor due to risk of reduced platelet inhibition and increased risk of bleeding and adverse cardiovascular *2/*17 metabolizer *2/*2, events consider alternative therapy such as prasugrel or ticagrelor due to significant risk of reduced platelet inhibition and severely increased risk of bleeding *2/*3, *3/*3 poor metabolizer adverse cardiovascular events Type of Modification Recommendation factor Lab Dose Type therapy change therapy change standard loading dose, reduce maintenance dose by 25% phenytoin CYP2C9 *1/*2, *1/*3 low metabolizer phenytoin CYP2C9 *2/*2, very low *2/*3, *3/*3 metabolizer *1/*1, *1/*2, warfarin CYP2C9 amitriptyline CYP2D6 amitriptyline CYP2D6 due to risk for adverse effects such as ataxia, nystagmus, dysarthria and sedation dose change standard loading dose, reduce maintenance dose by 50% due to risk for adverse effects such as ataxia, nystagmus, dysarthria and sedation dose change dose reduction of warfarin is recommended due to risk 0.75 Maintenance 0.5 Maintenance of bleeding and cardiovascular events, it is *1/*3, low metabolizer recommended to use a dosing algorithm *1/*1xN, ultrarapid avoid tricyclic antidepressant due to lack of efficacy, *1/*2xN *4/*10, metabolizer intermediate consider alternative drug not metabolized by CYP2D6 therapy change consider 25% reduction in dosing due to increased risk of *5/*41 metabolizer side effects avoid tricyclic antidepressant due to high risk of side effect, consider alternative drug not metabolized by *5/*5, *4/*6 poor metabolizer CYP2D6, if tricyclic is used consider 50% dose reduction dose change dose change 0.75 All 0.75 All *4/*4, *4/*5, amitriptyline CYP2D6 *1/*1xN, ultrarapid avoid codeine use due to potential for toxicity, consider alternative opiate avoid codeine use due to lack of efficacy, consider therapy change codeine CYP2D6 *1/*2xN *4/*4, metabolizer therapy change codeine CYP2D6 *4/*5, poor metabolizer alternative opiate capecitabine DPYD DPYD*2A/*2 complete DPD A deficiency decreased DPD avoid capecitabine due to risk of severe or fatal drug toxicity therapy change start with 50% dose reduction then titrate dose based on capecitabine DPYD DPYD*1/*2A activity toxicity fluorouracil DPYD DPYD*2A/*2 complete DPD A deficiency fluorouracil DPYD decreased DPD DPYD*1/*2A activity avoid fluorouracil due to risk of severe or fatal drug toxicity therapy change start with 50% dose reduction then titrate dose based on toxicity dose change therapy change dose change 0.5 All 0.5 All DPYD*2A/*2 complete DPYD tegafur tegafur DPYD DPYD A deficiency avoid tegafur due to risk of severe or fatal drug toxicity decreased DPYD start with 50% dose reduction then titrate dose based on DPYD*1/*2A activity toxicity therapy change dose change 0.5 All Monitoring 29 Chavez, Heffernan, West avoid rasburicase, contraindicated due to risk of severe rasburicase G6PD abacavir HLA‐B *57:01 positive hemolysis avoid abacavir due to significantly increased risk of therapy change positive hypersensitivity therapy change avoid alluprinol due to significant risk of severe allopurinol HLA‐B HLA‐B*58:01 high risk cutaneous adverse reaction therapy change avoid carbamazepine if patient is naïve due to significant risk of cutaneous adverse reactions, if patient has been on carbamazepine therapy for longer than 3 months carbamazepine HLA‐B peginterferon alfa‐2a IFNL3 *15:02 CT or TT carrier unfavorable simvastatin SLCO1B1 TC consider dose reduction due to increased risk of intermediate risk myopathy simvastatin SLCO1B1 CC *1/*2, azathioprine TPMT high risk *1/*3A, *1/*3B, high risk *3A/*3A, *2/*3A, azathioprine TPMT without reaction may consider continued use consider alternative therapy due to lack of efficacy consider dose reduction due to increased risk of myopathy, consider routine creatine kinase monitoring consider initial dose reduction of 30‐70% or target dose and titrate based on tolerance consider alternative therapy due to risk of therapy change therapy change dose change 0.25 All dose change 0.5 All dose change 0.5 All myelosupression, if azathioprine is used drastically extremely high risk reduce dose by 10‐fold and consider three times a week dosing instead of daily dosing therapy change 0.1 All high risk consider initial dose reduction of 30‐70% or target dose and titrate based on tolerance dose change 0.5 All *2/*3A, extremely high used drastically reduce dose by 10‐fold and consider *3C/*3A, *1/*2, risk three times a week dosing instead of daily dosing dose change 0.1 All *1/*3A, *1/*3B, *3A/*3A, high risk consider initial dose reduction of 30‐50% or target dose and titrate based on tolerance dose change 0.5 All *3C/*3A, *3C/4, *1/*2, mercaptopurine TPMT *1/*3A, *1/*3B, *3A/*3A, mercaptopurine TPMT thioguanine TPMT *2/*3A, extremely high used drastically reduce dose by 10‐fold and consider thioguanine TPMT *3C/*3A, risk three times a week dosing instead of daily dosing dose change reduce initial dose by 30% for patients receiving dose of more than 250 mg/m^2 due to risk of myelosupression 0.1 All irinotecan UGT1A1 UGT1A1*28 high risk and arrhythmia 0.7 >250 mg/m^2 dose change dose reduction of warfarin is recommended due to risk of bleeding and cardiovascular events, it is warfarin VKORC1 GG, GA, AA low metabolizer recommended to use a dosing algorithm dose change 0.75 All CPK 30 Chavez, Heffernan, West Appendix B. Scenario Testing Script Step 1 Team User/ Dept prescriber Workflow Step order corresponding medication for drug/gene pair 2 prescriber order corresponding medication for drug/gene pair 3 prescriber order corresponding medication for drug/gene pair 4 prescriber 5 prescriber order corresponding medication for drug/gene pair where 2 genes relevant to the medication are gene result returns with pre-existing order for medication prescriber order corresponding medication for drug/gene pair prescriber order corresponding medication for drug/gene pair prescriber order corresponding medication for drug/gene pair prescriber order corresponding medication for drug/gene pair where 2 genes relevant to the medication are gene result returns with pre-existing order for medication prescriber Actual Outcome Testing Step/Process Expected Outcome Test Patient 1 (no pre-existing prescriber alerted to absence result) - order coumadin of any gene testing and OUTPATIENT advised to order as results could affect dosing and/or Test Patient 2 (pre-existing Prescriber completes order for gene result with genotype that medication without any alert does not affect dosing and/or efficacy) - order coumadin Test Patient 3 (pre-existing prescriber alerted to presence gene result with genotype that of genotype that affects does affect dosing and/or dosing and/or efficacy with efficacy) - order coumadin recommended modification Test Patient 4 (2 pre-existing prescriber sees both alerts gene results with genotype that affect dosing and/or efficacy of single medication) order coumadin Test Patient 3 (pre-existing prescriber alerted to presence gene result with genotype that of genotype that affects does affect dosing and/or dosing and/or efficacy with efficacy) - refill/reorder recommended modification Test Patient 1 (no pre-existing prescriber alerted to absence result) - order coumadin of any gene testing and INPATIENT advised to order as results could affect dosing and/or Test Patient 2 (pre-existing Prescriber completes order for gene result with genotype that medication without any alert does not affect dosing and/or efficacy) - order coumadin Test Patient 3 (pre-existing prescriber alerted to presence gene result with genotype that of genotype that affects does affect dosing and/or dosing and/or efficacy with efficacy) - order coumadin recommended modification Test Patient 4 (2 pre-existing prescriber sees both alerts gene results with genotype that affect dosing and/or efficacy of single medication) order coumadin - INPATIENT Test Patient 3 (pre-existing prescriber alerted to presence gene result with genotype that of genotype that affects does affect dosing and/or dosing and/or efficacy with efficacy) - refill/reorder recommended modification Pass/ Fail Tester Date Tested Comments or Issues pharmacist 6 7 dispense corresponding medication for drug/gene pair order corresponding medication for drug/gene pair Test Patient 1 (no pre-existing pharmacist alerted to absence result) - dispense coumadin - of any gene testing and INPATIENT advised to order as results could affect dosing and/or pharmacist Test Patient 2 (pre-existing pharmacist completes gene result with genotype that dispense for medication does not affect dosing and/or without any alert efficacy) - dispense coumadin pharmacist order corresponding Test Patient 3 (pre-existing pharmacist alerted to medication for gene result with genotype that presence of genotype that drug/gene pair does affect dosing and/or affects dosing and/or efficacy efficacy) - dispense coumadin - with recommended pharmacist order corresponding Test Patient 4 (2 pre-existing pharmacist sees both alerts medication for gene results with genotype drug/gene pair where that affect dosing and/or 2 genes relevant to efficacy of single medication) the medication are dispense coumadin pharmacist gene result returns Test Patient 3 (pre-existing pharmacist alerted to with pre-existing order gene result with genotype that presence of genotype that for medication does affect dosing and/or affects dosing and/or efficacy efficacy) - refill/redispense with recommended coumadin - INPATIENT modification result abstractor result entry enter genotype with incorrect result component will only string allow validated strings medical records historical Enter patient reported no alert fires assistant medication medication (Coumadin) on