* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Identification of Novel Starch Traits in Sorghum
Survey
Document related concepts
Non-coding DNA wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene desert wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene regulatory network wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Genome evolution wikipedia , lookup
Copy-number variation wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Community fingerprinting wikipedia , lookup
Point mutation wikipedia , lookup
Transcript
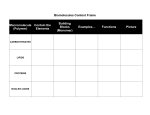
IAEA-CN-167-011P Identification of Novel Starch Traits in Sorghum (Sorghum bicolor. L) : A Reverse Genetics Approach Helen Hill, L.Slade Lee and Robert Henry Grain Foods CRC Limited, Centre for Plant Conservation Genetics, Southern Cross University P.O. Box 157, Military Road Lismore 2480 Australia. Introduction Methods Mature seeds of Sorghum bicolor L (cv. MR 43) were bombarded with gamma radiation to induce mutation. Following treatment, over 1000 irradiated seeds were grown in a glasshouse at 25-30°C with 16 hours day length. M2 leaf samples were collected and genomic DNA was extracted utilising a Qiagen 96-well MagAttract kit on the MWG Theonyx® robotic platform. PCR was performed to amplify our genes of interest and then directly sequenced. Following sequencing all individual data was manually visualised and aligned using SEQUENCHER™ 4.0 Software (Gene Codes Corporation, USA) and compared to published sequence in Genbank. Sorghum (Sorghum bicolor L.) is the fifth most important cereal grain crop in the world and reportedly feeds over 500 million people on a daily basis in the developing world providing dietary starch, dietary protein and some vitamins and minerals. Traditionally, sorghum grain is used for thin and thick porridges, noodles, tortillas, flatbread, biscuits, snacks, couscous, and traditional beer. In the West, it is predominantly used as an animal feed and is becoming important for ethanol production. More recently, specialised sorghum starch has been used for food and industrial uses. The aim of this study is to use a reverse genetics approach to detect DNA base changes in four important starch synthesis genes encoding; soluble starch synthase (SS I and SS IIa), granule bound starch synthase (GBSS) and starch branching enzyme (SBE IIb) in our gamma irradiated sorghum (cv. MR 43) population and to correlate sequence variation to altered starch phenotype which may subsequently by used for novel food applications. GBSS SS IIa Results We have surveyed over 3.3 kb of the GBSS gene, 2.5 kb region of SBE IIb, 1kb region of SS IIa gene and 400bp of the SS I gene and found numerous single nucleotide polymorphisms (SNP) and insertion/deletions (indels) in intronic regions and also some single nucleotide polymorphisms in exonic regions of the genes of interest. (Table 1) Table 1 summarises the natural variation found between the putative sorghum mutants and the published Genbank sequence for GBSS, SS IIa, SS I and SBE IIb respectively, which consisted of indels and base substitutions. Gene product Abbreviation Granule bound starch synthase GBSS Starch branching SBE II enzyme Figure 1 shows the difference in growth of the sorghum seedlings, after the seed was subjected to different gamma radiation doses. Conclusions Soluble starch synthase Soluble starch synthase Length Summary of findings studied 13 SNPs; including transitions being CgT, AgG or GgA base change and transversions being a 3.3kb AgC, TgA, TgG and GgT and a single dinucleotide AAgTC snp ; and 4 indels including an AT , T , TT and TATGTAGT indel 8 SNPs; including transitions AgG, TgC and GgA; transversions being TgA, TgG, CgG and a 2.5kb dinucleotide snp TTgAG; and 4 indels including an AA, ACG, AT and ATA indel. SS IIa 1kb 1 SNP being CgT change in exon SS1 400bp 1 SNP being GCgCA in exon Whereas mutations, in the form of single nucleotide polymorphisms (SNP) and insertion/deletion (indel) events, predominantly occurred in non-coding introns, some mutations were also found in exons in all genes studied. These SNP changes resulted in some differences in the amino acids in each of the starch synthesis genes of interest and which leads to alternative proteins encoded and may subsequently result in an altered starch phenotype in our mutant sorghum. Phenotypic analyses will be carried out to correlate sequence variation to altered starch phenotype. Screening of diverse sorghum germplasm by the EMAIL technique developed at CPCG is currently proceeding. SCU2554 Further Work