* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Highly Conserved Region 141–168 of the NS1 Protein Is a New
Survey
Document related concepts
Paracrine signalling wikipedia , lookup
Expression vector wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Biochemistry wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Protein structure prediction wikipedia , lookup
Point mutation wikipedia , lookup
Proteolysis wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Transcript
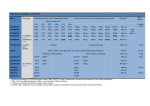
Jpn. J. Infect. Dis., 64, 109-115, 2011 Original Article Highly Conserved Region 141–168 of the NS1 Protein Is a New Common Epitope Region of Dengue Virus Promsin Masrinoul1, Magot Omokoko Diata1, Sabar Pambudi1, Kriengsak Limkittikul3, Kazuyoshi Ikuta1,2, and Takeshi Kurosu1,2* 1 Department of Virology, Research Institute for Microbial Diseases, Osaka University, Osaka 565-0871, Japan; and 2 Mahidol-Osaka Center for Infectious Diseases and 3 Department of Tropical Pediatrics, Faculty of Tropical Medicine, Mahidol University, Bangkok 10400, Thailand (Received December 15, 2010. Accepted January 12, 2011) SUMMARY: Dengue virus (DENV) nonstructural protein 1 (NS1) is a major target of humoral immunity in patients and is believed to be involved in DENV pathogenesis. In addition, NS1 is a diagnostic target as it is secreted, and circulates, in patients’ plasma at an early stage of viral infection. In this study, we aimed to identify common epitope regions for all serotypes by preparation of mouse monoclonal antibodies (MAbs) against NS1. A total of 10 out of the 20 hybridoma clones which were specific to DENV produced MAbs that recognized NS1. These MAbs mapped to three regions of DENV-2 NS1, namely amino acids 1–40 (epitope region 1), 141–168 (epitope region 2), and 267–312 (epitope region 3). Epitope region 2 was recognized by both complex-specific (2H11 and 3C4) and subcomplex-specific MAbs (4E5 and 5G12), whereas epitope regions 1 and 3 were recognized by subcomplex-specific MAbs (5E2, 1A5, and 3F10) only. These epitope regions were found to be highly conserved among all four serotypes of DENV by sequence analysis and database comparison. The MAbs against these epitope regions, especially 2H11 and 3C4, could therefore be valuable diagnostic tools. reported that immunization with NS1 provides protection to mice against lethal DENV, Japanese encephalitis virus (JEV), and West Nile virus (WNV) infections, thus suggesting that NS1 could be a target for therapy or a subunit vaccine against flaviviruses (11–14). The identification of antigenic regions of NS1 proteins conserved among all four DENV serotypes will be useful for the development of subunit vaccines (15), and monoclonal antibodies (MAbs) against these common epitope regions will be valuable diagnostic tools. In this study, we generated DENV NS1-specific MAbs and identified three cross-reactive epitope regions. Epitope region 1 (amino acids [aa] 1–40) and epitope region 3 (267– 312) were identified by subcomplex-specific MAbs (5E2, 1A5, and 3F10), which reacted with more than one serotype of DENV but not all serotypes, and epitope region 2 was newly identified by complex-specific MAbs (2H11 and 3C4), which reacted with all serotypes of DENV and certain subcomplex-specific MAbs (4E5 and 5G12). These epitope regions, especially epitope region 2, were found to be highly conserved among all DENV serotypes, thereby suggesting that MAbs 2H11 and 3C4 could prove to be useful diagnostic tools. INTRODUCTION Dengue fever/dengue hemorrhagic fever (DHF) is the most important human arthropod-borne viral disease, and the incidence of severe and life-threatening forms of this disease, namely DHF or dengue shock syndrome, is increasing (1,2). Dengue virus (DENV) has four serotypes (DENV 1–4), all of which are responsible for severe forms of the disease. The World Health Organization currently estimates that there are 50 to 100 million dengue infections globally each year, more than 500,000 of which are DHF (3). However, although DENV is a major public health problem, an effective vaccine or treatment for DENV infection has yet to be developed. The DENV genome contains three structural proteins (capsid, pre-membrane [prM], and envelope) and seven nonstructural proteins (NS1, NS2a, NS2b, NS3, NS4a, NS4b, NS5). NS1 is a 46–50-kDa glycoprotein expressed in infected mammalian cells in both membrane-associated (mNS1) and secreted (sNS1) forms (4,5). Furthermore, circulating sNS1 proteins have been detected in patients’ plasma with DENV infections. sNS1 could therefore be a target for early diagnosis. NS1 is also known to be a major target of humoral immunity in DENV infection (6–8). Likewise, there is accumulating indirect evidence for an involvement of NS1 in DENV pathogenesis because high levels of NS1 have been detected in patients’ plasma and correlated with a severe form of the disease (9,10). In addition, several groups have MATERIALS AND METHODS Cells and viruses: DENV serotypes 1 (Mochizuki strain), 2 (16681 strain), 3 (H87 strain), 4 (H241 strain), and JEV (JaGAr strain) were propagated in C6/36 cells for 7–9 days. The culture medium was stored at –80°C before use. Myeloma PAI cells were cultured in Roswell Park Memorial Institute 1640 medium (RPMI 1640; NACALAI TESQUE, Kyoto, Japan) supplemented with 10% fetal calf serum *Corresponding author: Mailing address: Department of Virology, Research Institute for Microbial Diseases, Osaka University, Suita, Osaka 565-0871, Japan. Tel: +81 6 6879 8308, Fax: +81 6 6879 8310, E-mail: [email protected] 109 (FCS). Human renal epithelial cell line 293T cells were cultured in Dulbecco’s modified Eagle medium (DMEM; Gibco, Gland Island, N.Y., USA) supplemented with 10% FCS. Baby hamster kidney (BHK) cells were grown in Eagle’s minimum essential medium (MEM; NACALAI TESQUE) supplemented with 10% FCS. All cell lines were cultured at 37°C with 5% CO2. The C6/36 mosquito cells were grown in Leibovitz 15 medium (Gibco) supplemented with 0.3% BactoTM Tryptose Phosphate Broth (Becton Dickinson, Sparks Glencoe, Md., USA) and 10% FCS at 28°C. DENV, BHK cells, and C6/36 cells were kindly provided by Dr. Yoshinobu Okuno, Kanonji Institute, Research Foundation for Microbial Diseases of Osaka University, Kanonji, Kagawa, Japan. Human DENV patient plasma: Plasma from patients infected with DENV 1–4 was kindly provided by the Faculty of Tropical Medicine, Mahidol University, Bangkok, Thailand. This study was approved by the local Ethics Committee, and all patients provided informed consent to participate. Generation of dengue-specific MAbs: For the preparation of DENV 1–4 particles, the supernatants from BHK cells individually infected with DENV 1–4 were ultracentrifuged at 100,000 × g for 2 h. The pelleted DENV particles were resuspended in PBS and kept at –80°C prior to injection. The recombinant NS1 protein (rNS1) derived from DENV-2 New Guinea C strain (NGC) was expressed in Escherichia coli BL21 strain and purified under denaturing conditions, as described previously (16). Four-week-old female BALB/ c mice (JAPAN SLC, Shizuoka, Japan) were immunized intraperitoneally with a mixture of DENV 1–4 particles with/ without inactivation by 4% paraformaldehyde (PFA) in PBS, or recombinant NS1 from DENV-2 NGC with complete Freund’s adjuvant (Wako, Osaka, Japan). The immunized mice were boosted 3–4 times with BHK cells infected with a mixture of DENV 1–4. Three days after the final booster, splenocytes were prepared and fused to myeloma PAI cells using polyethylene glycol #1500 (Roche Diagnostics, Mannheim, Germany). The fused cells were cultured in DMEM supplemented with 15% FCS in the presence of hypoxanthine-aminopterin-thymidine (Gibco). MAbs produced from the hybridomas were screened in an immunofluorescent assay using BHK cells infected with DENV 1– 4. Hybridomas were cloned twice by limiting dilution. Indirect immunofluorescence assay (IFA): For the preparation of virus antigen, BHK cells infected with DENV 1–4 or mock-infected were collected 2 days postinfection. Cells infected separately with DENV 1–4 were mixed and seeded onto 96-well plates for hybridoma screening. The cells were fixed with 4% PFA in PBS, permeabilized with 1% Triton X-100 in PBS for 5 min, and then incubated with hybridoma culture fluid or anti-DENV-specific MAbs (2H2) (HB114, ATCC) as control for 1 h. They were then washed three times with PBS and further treated with Alexa 488 goat anti-mouse IgG antibody (Invitrogen, Paisley, UK) at a dilution of 1:300 for 30 min. Finally, they were washed three times with PBS prior to observation by fluorescence microscopy. Determination of serotype-specificity and isotyping of MAbs: The serotype specificities of MAbs were determined by IFA and immunoblotting using DENV-infected BHK cells. The isotypes of antibodies were examined by immunochromatrography using an IsoQuickTM kit (Sigma, St. Louis, Mo., USA) SDS-PAGE and Western blotting (WB) analysis: The DENV-infected BHK cells were dissolved in sodium dodecyl Table 1. Primers used for the construction of cDNA clones Recombinant gene Primer NS1-NS2A (Full) Fw (EcoRV) Rv (BamH1) 5´ GCCGATATCAGATAGTGGTTGCGTTGTGAGC 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (41~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGAAACTAGCTTCAGCTATCCAGAAA 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (81~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGGAAAATGAGGTGAAGTTAACTATT3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (121~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGGCAAAAATGCTCTCTACAGAGTC 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (141~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGGCAGAATGCCCCAACACAAATAG 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (169~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGCTAAAATTGAAAGAAAAACAGGAT 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (197~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGGATATGGGTTATTGGATAGAAAGTG 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (221~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGAAAAACTGCCACTGGCCAAAATC 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (267~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGCCATGGCATCTAGGTAAGCTTGAG 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ NS1-NS2A (313~) Fw (EcoRV) Rv (BamH1) 5´ GGCGATATCGTGCTGCCGATCTTGCACATTACCA 3´ 5´ GCGGGATCCTCATCGGGTCCTAAGCATTTCCTCC 3´ The underlines indicate the restriction enzyme recognition sites. 110 sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) sample buffer without β-mercaptomethanol and heated at 100°C for 5 min. The resulting samples were separated on a 15% SDS-PAGE gel and transferred to polyvinylidene fluoride (PVDF) membranes (Millipore Corp., Bedford, Mass., USA). These membranes were incubated overnight with antibodies produced by hybridoma clones and then with horseradish peroxidase-conjugated anti-mouse IgG (Jackson ImmunoResearch Laboratories, West Grove, Pa., USA) for 1 h. The reactive viral proteins were visualized using ECL WB detection reagent (GE Healthcare, Buckinghamshire, UK). Plasmid construction: For the construction of plasmids expressing NS1 protein, viral RNA was extracted using a QIAmp Viral RNA Mini kit (Qiagen, Valencia, Calif., USA), and reverse transcription was performed using Superscript III (Invitrogen) with random primers. The resultant cDNAs were amplified by PCR with the set of primers (Table 1). The PCR products were inserted into the EcoRV/BamHI sites of pFLAG-CMVTM-3 expression vector (Sigma), then the plasmids were transformed into E. coli XL1-blue cells and subjected to sequence analysis. The expression of these mutant NS1 proteins was confirmed by WB analysis using an anti-Flag M2 antibody. Transfection: For epitope mapping, a series of truncated mutant NS1 plasmids was transfected into 293 T cells using Lipofectamine 2000 (Invitrogen). At 2 days after transfection, cells were detached by treatment with 0.5 mM EDTA in PBS and washed once with PBS. The cell precipitates were resuspended with PBS, coated onto glass slides, and then subjected to IFA with MAbs, human plasma infected with DENV (HPD), or anti-Flag M2 antibody (Sigma). The residual proteins were detected by immunoblotting. Sequence analysis: The amino acid sequences of NS1 proteins were aligned using ClustalW, and percent identities were calculated using Bioedit version 7.0.5. Analysis of amino acid sequence variation within epitope regions: A total of 1,228 DENV-1 sequences, 815 DENV-2 sequences, 572 DENV-3 sequences, and 91 DENV4 sequences were collected from the NCBI protein database. The amino acid sequences within epitope regions were analyzed by using Bioedit version 7.0.5. Variations for each amino acid residue were displayed in an Entropy H(x) plot. Identification of antigenic DENV proteins: Immunoblotting after SDS-PAGE was carried out with MAbs using DENV-infected BHK cell lysate under non-reducing conditions to identify antigenic target DENV proteins. A total of 10 out of the 20 MAbs (50%) reacted with a DENV protein at 46 kDa (NS1) (Fig. 1), whereas two MAbs (10%) reacted with a DENV protein at 18–20 kDa (prM) (data not shown). The target viral proteins for eight MAbs could not be identified, thus suggesting that these MAbs might recognize conformational epitopes. Of the 10 anti-NS1 MAbs described above, 2H11 and 3C4 were found to be complex-specific, 4G1 and 4C2 were found to be serotype-specific, and the others (1A5, 3F10, 4E5, 5E2, 4G11, and 5G12) were found to be subcomplex-specific (Table 2). The MAb isotypes were determined using an immunochromatography kit, and all were found to be of the IgG isotype. MAbs 4C2, 1A5, 3F10, 4E5, and 5E2 were identified as IgG1,MAbs 4G1, 5G12, 4G11, and 2H11 as IgG2, and MAb 3C4 as IgG3. Mapping of cross-reactive MAbs to recombinant truncated forms of NS1 revealed three predominant epitope regions: The majority of subcomplex anti-NS1 MAbs (1A5, 3F10, 4E5, 5E2, 5G12, 2H11, and 3C4) reacted with DENV2. To identify the epitope regions of these MAbs, a series of recombinant DENV-2 NS1 truncated mutants was expressed in 293T cells. The plasma from DENV-infected patients reacted with all mutant NS1s except one (313–352), thus indicating that B-cell epitopes of NS1 are spread across the whole region of NS1. Epitope mapping using these MAbs revealed three predominant epitope regions, with the 5E2 MAb mapping to an N-terminal region corresponding to aa 1–40 (epitope region 1). Four MAbs (2H11, 3C4, 4E5, and 5G12) mapped to the region located at aa 141–168 (epitope region 2), and MAbs 1A5 and 3F10 mapped to a C-terminal region located at aa 267–312 (epitope region 3) (Fig. 2). As it has not been identified previously, region 2 can be considered to be a new B-cell epitope region. The identified epitope regions exhibit high conservation among the four DENV serotypes: Since most of the antibodies obtained were serotype cross-reactive antibodies (complex- and subcomplex-specific), the epitope regions for each of the four serotypes were aligned (Fig. 3) and the percent identities of their amino acid sequences examined (Table 3). Epitope region 2 was found to be highly conserved among all DENV serotypes, ranging in percent identity from 75.0 to 82.1%. Epitope regions 1 and 3 were also highly conserved among DENV-1, -2, -3 and DENV-2, -3, -4, respectively, ranging in percent identities from 72.5 to 92.5% and 71.1 to 78.2%, respectively. Furthermore, these epitope regions were found to be DENV-specific as none of them exhibited high percent identities with JEV (values ranging from 42.5 to 65.2%). The MAbs against these regions, especially 2H11 and 3C4, could therefore prove to be valuable diagnostic tools as they cross-react with all DENV serotypes. Highly conserved epitope regions among prevalent DENV: To confirm the conservation of each epitope region, their amino acid sequences were analyzed using 2,706 reference sequences for DENV 1–4 from the NCBI database (Fig. 4). For further analysis at the amino acid level, the consensus sequences corresponding to the epitope regions of each serotype were obtained and their peptide sequences compared. Highly conserved regions were found at aa 12– 20 and 25–35 of epitope region 1, aa 154–161 of epitope RESULTS Generation and specificity of MAb against DENV: In order to generate mouse MAb, BALB/c mice were immunized intraperitoneally with a mixture of DENV 1–4 particles with/without inactivation or recombinant DENV2 NS1 protein, and boosted with DENV-infected BHK cell lysate. A total of 20 hybridomas were screened using BHK cells infected with the DENV 1–4 mixture. To determine the cross-reactivity to each serotype, the MAb culture supernatants were examined by IFA using DENV-infected BHK cells. Eight MAbs (3A1, 2B8, 3D8, 3A4, 3H12, 4C2, 4G1, and 6A5) reacted specifically with only one serotype (serotype-specific), whereas six MAbs (1A5, 3F10, 4E5, 5E2, 4G11, and 5G12) reacted with more than one DENV serotype, although not all serotypes (subcomplex-specific), and four MAbs (2H11, 3C4, 3D2, and 6D6) reacted with all DENV serotypes (complex-specific). Furthermore, two MAbs (2F12 and 2H2) reacted with all DENV serotypes as well as JEV (group-specific) (Table 2). 111 Table 2. Characterization of serotype-specific and cross-reactive specific MAbs MAb Reactivity by IFA Ig isotype (κ, λ) DENV-1 DENV-2 DENV-3 DENV-4 JEV IgM κ IgG2b κ IgG3 κ IgM κ IgG2a κ IgG1 κ IgG2a κ IgM κ IgG1 λ IgG2a κ IgG1 κ IgG2b κ IgG1 κ IgG1 κ IgG2a κ IgG3 κ IgG2b κ IgG3 κ IgM κ IgM κ – – – – – – – – + + + + – – + + + + + + + – – – – – – – + – + + + + + + + + + + – – – – – – – – – + + + + + + + + + + + – + + + + + + + – – – – + + + + + + + + – – – – – – – – – – – – – – – – – – + + 3A1 2B8 3A4 3D8 3H12 4C2 4G1 6A5 4E5 4G11 5E2 5G12 1A5 3F10 2H11 3C4 3D2 6D6 2F12 2H2 Protein Immunization2) U1) U U U U NS1 NS1 U NS1 NS1 NS1 NS1 NS1 NS1 NS1 NS1 prM prM U U A B C C C C C C A C B C A A B B C C B C 1) : U indicates unidentified. : Different immunizations were primarily used. A, B, and C indicate immunizations with an inactivated mixture of DENV, a live mixture of DENV, and recombinant NS1, respectively. All of mice were boosted with DENV-infected BHK cell lysates. DENV-4 DENV-3 DENV-2 DENV-1 Mock 2) region 2, and aa 267–274 and 294–302 of epitope region 3 (Fig. 4). HPD DISCUSSION 4G1 Three B-cell epitope regions of DENV NS1 have been identified by using cross-reactive MAbs. These regions were defined as epitope regions 1 (1–40), 2 (141–168), and 3 (267– 312) and were found to stretch along the whole region of NS1. Plasma from dengue patients reacted with almost the whole region (aa 1–312) of NS1, except for the C-terminal region (aa 313–352). Epitope regions 1 and 3 are partly consistent with those identified in previous reports (17,18), whereas epitope region 2 is identified for the first time in this study. It should be noted that our immunization and boosting schemes differ from those used in previous papers. Thus, in this study, three different antigens, namely a mixture of DENV 1–4 particles with/without inactivation and rNS1 protein, were used for immunization, and cell lysates derived from DENV 1–4-infected BHK cells were used for boosting. In contrast, in other studies, NS1 purified by affinity column (19), rNS1 expressed in E. coli (17,20,21), and DENV-infected mouse brain homogenate (22) were used for immunization and for boosting. We believe that these differences in the types of immunogen utilized may explain why the epitope regions identified by each study do not match completely. The newly identified epitope region 2 is likely to be a common region for all DENV serotypes as two MAbs (2H11 and 3C4) reacted with all such serotypes and a further two (4E5 and 5G12) reacted with DENV-1, -2, and DENV-1, -2, and -3, respectively. Sequence analysis using references obtained from the NCBI database confirmed that this region was highly conserved among DENVs, with the eight 4C2 4G11 1A5 3F10 4E5 5E2 5G12 2H11 3C4 Fig. 1. Characterization of serotype-specific and cross-reactive MAbs. The cell lysates of C6/36 cells infected with DENV 1–4 individually were subjected to SDS-PAGE and transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were stained with each MAb separately. The NS1 proteins exhibited the mobility at 46 kDa. 112 A (41-352) (81-352) (121-352) (141-352) (169-352) (197-352) (221-352) (267-352) (313-352) (+) (+) (+) (+) (+) (+) (+) (+) (+) (+) (-) (+) (+) (+) (+) (+) (+) (+) (+) (+) (-) (-) (+) (+) (+) (+) (+) (+) (+) (+) (+) (-) (-) (+) (+) (+) (+) (+) (+) (+) (+) (+) (-) (-) (+) (+) (+) (+) (+) (-) (-) (-) (-) (-) (-) (+) (+) (+) (+) (+) (-) (-) (-) (-) (-) (-) (+) (+) (+) (+) (+) (-) (-) (-) (-) (-) (-) (+) (+) (+) (+) (+) (-) (-) (-) (-) (-) (-) (+) (-) (-) (-) (-) (-) (-) (-) (-) (-) Mock αFlag (1-352) (-) M2 HPD 1A5 3F10 2H11 3C4 4E5 5G12 5E2 B 1 NS1 Epitope region 1 (1-40) 5E2 352 Epitope region 2 (141-168) Epitope region 3 (267-312) 2H11 1A5 3C4 3F10 4E5 5G12 Fig. 2. Epitope mapping of cross-reactive MAbs. (A) The NS1 truncated mutants were expressed in 293T cells. The cells were stained with anti-Flag M2 antibody, HPD, and MAbs, and followed by Alexa 488 goat anti-mouse IgG secondary antibody. The MAbs mapped to three main regions, 1–40, 141–168, and 267–312. (B) Schematic figure of three main epitope regions of each MAb on NS1 protein. amino acids positioned at aa 154–161 of epitope region 2 being particularly highly conserved among the reference sequences, even in different DENV serotypes (Fig. 3). This region may therefore be part of a recognition site for the complex-specific MAbs 2H11 and 3C4. We expected to obtain cross-reactive (complex- and subcomplex-specific) anti-NS1 MAbs more easily upon immunization of mice with rNS1. However, this was not the case, and MAbs generation actually improved when mice were immunized with a live/inactivated mixture of DENVs. Curiously, more than half of the MAbs produced by immunization with DENV-2 rNS1 did not react with DENV-2 NS1, although they did react with other DENV serotypes. This suggests that the conformation of rNS1 may not fully reflect the native form of NS1. Since DENV NS1 easily forms inclusion bodies in E. coli (23), rNS1 was purified under denaturing conditions (16), which may have changed the actual conformation of NS1. It is presumably 113 Flavivirus strain Peptide sequence Epitope region 1 (1-40) DENV-1-Mochizuki DENV-2-16681 DENV-3-H87 DENV-4-H241 JEV-JaGAr Epitope Region 2 (141-168) DENV-1-Mochizuki DENV-2-16681 DENV-3-H87 DENV-4-H241 JEV-JaGAr 141 PECPDDQRAWNIWEVEDYGFGIFTTNIW 168 A...NTN....SL........V...... ....SAS....V.........V...... S...NER....FL........M...... K....EH....SMQI..F....TS.RV. DENV-1-Mochizuki DENV-2-16681 DENV-3-H87 DENV-4-H241 JEV-JaGAr 267 PWHLGKLELDFDLCEGTTVVVDEHCGNRGPSLRTTTVTGKVIHEWC 312 31 ........M...F.D......T.D............AS..L.T... ...........NY.......IS.N..T..........S..L..... ........I..GE.P....TIQ.D.DH.........AS..LVTQ.. ..DENGIV....Y.P..K.TIT.D..K....V....DS..L.TD.. Epitope region 3 (267-312) 1 DSGCVVNWKGRELKCGSGIFVTNEVHTWTEQYKFQADSPK 40 ......S..NK.........I.DN...........PE..S .M...I....K............................. .T..A.S.S.K..........IDN...........PE..A .T..AIDITRK.MR.......H.D.EA.VDR..YLPET.R Fig. 3. Amino acid sequence alignment of epitope regions among flaviviruses. Table 3. Amino acid homology of epitope regions among DENV1–4 and JEV Percent identity (%) of amino acid sequence Epitope Flavivirus Epitope region 1 (1–40) DENV-1 DENV-2 DENV-3 DENV-4 JEV Epitope region 2 (141–168) Epitope region 3 (267–312) DENV-1 (Mochizuki) DENV-2 (16681) DENV-3 (H87) DENV-4 (H241) JEV (JaGAr) 1D1) 77.5 ID 92.5 75.0 ID 72.5 82.5 72.5 ID 42.5 45.0 47.5 52.5 ID DENV-1 DENV-2 DENV-3 DENV-4 JEV ID 75.0 ID 82.1 78.9 ID 75.0 82.1 75.0 ID 57.1 53.5 50.0 53.5 ID DENV-1 DENV-2 DENV-3 DENV-4 JEV ID 80.4 ID 82.6 78.2 ID 65.2 76.0 71.1 ID 56.5 65.2 63.0 63.0 ID 1) : ID indicates identical virus strain. not necessary to prepare purified NS1 to obtain anti-NS1 antibodies; instead, it is likely to be more efficient and convenient to immunize with virus particles and boost with cell lysates derived from DENV-infected cells. In summary, MAbs 2H11 and 3C4, which recognize a newly identified epitope region (aa 141–168) of the NS1 protein, could be useful tools for early diagnosis as they react with all DENV serotypes. Conflict of interest None to declare. REFERENCES 1. Gubler, D.J. (1998): Dengue and dengue hemorrhagic fever. Clin. Microbiol. Rev., 11, 480–496. 2. Malavige, G.N., Fernando, S., Fernando, D.J., et al. (2004): Dengue viral infections. Postgrad. Med. J., 80, 588–601. 3. World Health Organization (2009): Dengue and dengue haemorrhagic fever. Fact sheet N°117. Online at <http://www.who.int/mediacentre/ factsheets/fs117/en/>. 4. Flamand, M., Megret, F., Mathieu, M., et at. (1999): Dengue virus type 1 nonstructural glycoprotein NS1 is secreted from mammalian cells as a soluble hexamer in a glycosylation-dependent fashion. J. Virol., 73, 6104–6110. 5. Winkler, G., Maxwell, S.E., Ruemmler, C., et al. (1989): Newly synthesized dengue-2 virus nonstructural protein NS1 is a soluble protein but becomes partially hydrophobic and membrane-associated after dimer- Acknowledgments This work was supported in part by SYSMEX Corporation (Osaka, Japan). P.M. was supported by the Ministry of Education, Culture, Sports, Science and Technology of Japan. The manuscript was proofread by Medical English Service (Kyoto, Japan). We thank Dr. Arias J. Fernando for the critical reading of the manuscript. 114 Entropy (Hx) Epitope region 1 (1-40) 1.4 1.2 1 0.8 0.6 0.4 0.2 33 34 35 36 37 301 302 303 304 40 32 300 39 31 299 38 30 297 298 28 29 296 27 26 25 24 23 22 21 20 19 18 17 16 15 14 13 12 11 9 10 8 7 6 5 4 3 2 1 0 Entropy (Hx) Epitope region 2 (141-168) 1.4 1.2 1 0.8 0.6 0.4 0.2 168 167 166 165 164 163 162 161 160 159 158 157 156 155 154 153 152 151 150 149 148 147 146 145 144 143 141 142 0 Entropy (Hx) Epitope region 3 (267-312) 1.4 1.2 1 0.8 0.6 0.4 0.2 311 312 310 309 308 307 306 305 295 294 293 292 291 290 289 288 287 286 285 284 283 282 281 280 279 278 277 276 274 275 273 272 271 270 269 268 267 0 Amino acid position Fig. 4. Entropy plot for amino acid variations of the main epitope regions among DENV1–4 database sequences. ization. Virology, 171, 302–305. 6. Falkler, W.A., Jr., Diwan, A.R. and Halstead, S.B. (1973): Human antibody to dengue soluble complement-fixing (SCF) antigens. J. Immunol., 111, 1804–1809. 7. Huang, J.H., Wey, J.J., Sun, Y.C., et al. (1999): Antibody responses to an immunodominant nonstructural 1 synthetic peptide in patients with dengue fever and dengue hemorrhagic fever. J. Med. Virol., 57, 1–8. 8. Kuno, G., Vorndam, A.V., Gubler, D.J., et al. (1990): Study of antidengue NS1 antibody by Western blot. J. Med. Virol., 32, 102–108. 9. Alcon, S., Talarmin, A., Debruyne, M., et al. (2002): Enzyme-linked immunosorbent assay specific to dengue virus type 1 nonstructural protein NS1 reveals circulation of the antigen in the blood during the acute phase of disease in patients experiencing primary or secondary infections. J. Clin. Microbiol., 40, 376–381. 10. Young, P.R., Hilditch, P.A., Bletchly C., et al. (2000): An antigen capture enzyme-linked immunosorbent assay reveals high levels of the dengue virus protein NS1 in the sera of infected patients. J. Clin. Microbiol., 38, 1053–1057. 11. Chen, Z., Liu, L.M., Gao N., et al. (2009): Passive protection assay of monoclonal antibodies against dengue virus in suckling mice. Curr. Microbiol., 58, 326–331. 12. Chung, K.M., Nybakken, G.E., Thompson, B.S., et al. (2006): Antibodies against West Nile virus nonstructural protein NS1 prevent lethal infection through Fc gamma receptor-dependent and -independent mechanisms. J. Virol., 80, 1340–1351. 13. Henchal, E.A., Henchal, L.S. and Schlesinger, J.J. (1988): Synergistic interactions of anti-NS1 monoclonal antibodies protect passively immunized mice from lethal challenge with dengue 2 virus. J. Gen. Virol., 69, 2101–2107. 14. Schlesinger, J.J., Brandriss, M.W., Cropp, C.B., et al. (1986): Protection against yellow fever in monkeys by immunization with yellow fever virus nonstructural protein NS1. J. Virol., 60, 1153–1155. 15. Gibson, C.A., Schlesinger, J.J. and Barrett, A.D. (1988): Prospects for a virus non-structural protein as a subunit vaccine. Vaccine, 6, 7–9. 16. Kurosu, T., Chaichana, P., Yamate, M., et al. (2007): Secreted complement regulatory protein clusterin interacts with dengue virus nonstructural protein 1. Biochem. Biophys. Res. Commun., 362, 1051– 1056. 17. Chen, Y., Pan, Y., Guo, Y., et al. (2010): Comprehensive mapping of immunodominant and conserved serotype- and group-specific B-cell epitopes of nonstructural protein 1 from dengue virus type 1. Virology, 398, 290–298. 18. Falconar, A.K., Young, P.R. and Miles, M.A. (1994): Precise location of sequential dengue virus subcomplex and complex B cell epitopes on the nonstructural-1 glycoprotein. Arch. Virol., 137, 315–326. 19. Falconar, A.K. (1997): The dengue virus nonstructural-1 protein (NS1) generates antibodies to common epitopes on human blood clotting, integrin/adhesin proteins and binds to human endothelial cells: potential implications in haemorrhagic fever pathogenesis. Arch. Virol., 142, 897–916. 20. Burke, D.S., Nisalak, A., Johnson, D.E., et al. (1988): A prospective study of dengue infections in Bangkok. Am. J. Trop. Med. Hyg., 38, 172–180. 21. Mason, P.W., Zugel, M.U., Semproni, A.R., et al. (1990): The antigenic structure of dengue type 1 virus envelope and NS1 proteins expressed in Escherichia coli. J. Gen. Virol., 71, 2107–2114. 22. Henchal, E.A., Henchal, L.S. and Thaisomboonsuk, B.K. (1987): Topological mapping of unique epitopes on the dengue-2 virus NS1 protein using monoclonal antibodies. J. Gen. Virol., 68 (Pt 3), 845– 851. 23. Das, D., Mongkolaungkoon, S. and Suresh, M.R. (2009): Super induction of dengue virus NS1 protein in E. coli. Protein Expr. Purif., 66, 66–72. 115