* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download PowerPoint Slides
Rheumatic fever wikipedia , lookup
Immune system wikipedia , lookup
Immunocontraception wikipedia , lookup
Psychoneuroimmunology wikipedia , lookup
Sociality and disease transmission wikipedia , lookup
Globalization and disease wikipedia , lookup
Vaccination wikipedia , lookup
Innate immune system wikipedia , lookup
Hygiene hypothesis wikipedia , lookup
Childhood immunizations in the United States wikipedia , lookup
Gluten immunochemistry wikipedia , lookup
Neonatal infection wikipedia , lookup
Schistosomiasis wikipedia , lookup
Multiple sclerosis research wikipedia , lookup
Hospital-acquired infection wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
Tuberculosis wikipedia , lookup
DNA vaccination wikipedia , lookup
Hepatitis B wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Antimicrobial peptides wikipedia , lookup
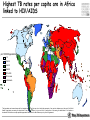
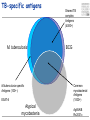
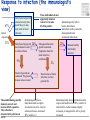
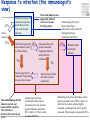
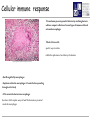
TB vaccines and diagnostics Introduction Global burden of tuberculosis (The economist’s view) An estimated 15 million active cases, leading to….. An estimated 9 million new infections Approx 2 million deaths Approx 2 Billion USD in direct control costs And an uncounted indirect cost in lost lives and productivity Tuberculosis: Transmission Primary Infection 10% Death ~2 mill infection (2 bill, TB TB ~ 9 mill/yr) TB Exposure/Infection 30% 90% Latent TB Reactivation 5-15% First 2yrs highest chance of developing TB disease Clearance 70% Treatment with several drugs for 6 months or more can cure more than 95% of patients If not treated 60 % dies TB • At present tuberculosis kills more people than any other infectious disease about 3 million people a year, including almost 300,000 children under 15, and is producing over 7,000 deaths and over 24,000 new cases every day. • No new drugs have been added to the first-line treatment regimen for TB for >30 yrs. • There is a clear synergy between M. tuberculosis and HIV, and active TB increases HIV-related immunodeficiency and mortality. • TB remains the largest attributable cause of death in HIVinfected individuals and is responsible for 32% of the deaths of HIV-infected individuals in Africa. • The neediest populations, in countries where TB incidence is highest, do not have access to treatment and, furthermore, in many cases, anti-TB drugs are ineffective. Highest TB rates per capita are in Africa linked to HIV/AIDS per 100 000 population < 10 10 to 24 25 to 49 50 to 99 100 to 299 300 or more No Estimate The boundaries and names shown and the designations used on this map do not imply the expression of any opinion whatsoever on the part of the World Health Organization concerning the legal status of any country, territory, city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. Dotted lines on maps represent approximate border lines for which there may not yet be full agreement. © WHO 2002 Stop TB Department TB cases have been rising in Africa and E Europe Incidence rate (/100K/yr) 500 rise in incidence slowing 400 Africa - high HIV 300 Africa - low HIV 200 Eastern Europe 100 incidence falling 0 1990 1992 1994 1996 1998 2000 2002 2004 Stop TB Department Development of Tuberculosis (the clinician’s view) Exposure TB (5%) TB (3%) TB (1%) TB (less than 0.1%/year) Health y (95%) Health y (92%) Health y (91%) Healthy (approx. 90%) Year 1 Year 2 Year 3 thereafter Response to infection (the immunologist’s view) 67% Initial exposure 33% These individuals do not apparently skin-test convert Early bacterial growth arrested at early time point. May (or may not) result in latent infection Early bacterial growth not contained. Leads to clinical illness 9% Bacterial growth not contained. Progressive disease unless treated These individuals generally skin-test convert. They often have characteristic patterns on X-ray. 24% Subsequent bacterial growth contained. Symptoms abate but latent infection established. 2% Reactivation of latent infection at a later point in life 22% Remain healthy but latently infected TB vaccines (BCG) • A 60-year follow-up study of American Indians reported the longterm efficacy of BCG to be 52%. The reasons for the low efficacy of the BCG vaccine may be generic differences in the BCG strains, differences in immunological properties of study populations or exposure to environmental factors such as mycobacteria. • Today, most of the world's population is vaccinated with BCG. It is generally accepted that BCG protects against childhood TB but this immunity wanes with age, resulting in no or insufficient protection against TB. • Among new vaccine candidates are live attenuated Mycobacterium tuberculosis vaccines, recombinant BCG, DNA vaccines, subunit vaccines and fusion proteins with novel adjuvants and delivery systems. Some of these vaccines are now in clinical trials. Reasons for failure: Treatment outcomes are worst in Africa and Europe W Pacific Died Failed SE Asia Defaulted Transfered Europe Not Evaluated E Med Americas Africa 0 10 20 30 Percent of cohort 40 Stop TB Department TB-specific antigens M. tuberculosis M.tuberculosis specific Antigens (100+): ESAT-6 Atypical mycobacteria Shared TB complex Antigens (4000+) BCG Common mycobacterial Antigens (1000+) Ag85A/B Rv2031c TB diagnostics • Left untreated, each person with active TB disease will infect on average between 10 and 15 people every year. Risk of TB in ESAT+ healthy contacts from Ethiopia Doherty et al., JCM , Feb. 2002 PPD: skin test (Purified Protein Derivative) High ESAT-6 immune reactivity reflects high levels of M. tuberculosis replication People who fail to control bacterial replication become ESAT+ and get TB CFU “Clinical disease” threshold “positivity” threshold People who fail to control initial bacterial replication become ESAT+, but if they control later infection, become latently infected Time after infection People who control initial bacterial replication remain ESAT-, and may or may not be latently infected Response to infection (the immunologist’s view) 67% Initial exposure 33% These individuals do not apparently skin-test convert or become ESAT-6 positive Early bacterial growth arrested at early time point. May (or may not) result in latent infection Early bacterial growth not contained. Leads to clinical illness 9% Subsequent bacterial growth contained. Symptoms abate but latent infection established. 22% Remain healthy but latently infected 2% Bacterial growth not contained. Progressive disease unless treated These individuals generally skin-test convert and become ESAT-6 positive. They often have characteristic patterns on X-ray. 24% Immunologically, little is known about these individuals as they cannot be distinguished from uninfected individuals Reactivation of latent infection at a later point in life Immunologically these individuals tend to express elevated levels of IL-4 and in advanced disease, decreased IFN-g and IL-12 Immunologically, these individuals tend to express elevated levels of IFN-g and IL-12, and while IL-4 often remains slightly increased, its antagonist IL-4d2 is greatly increased Bacterial response to infection Acute infection infection Latent Expression of early phase genes genes such as Ag85 and ESAT-6 Expression of late phase such as a-crystallin and the DosR regulon Acute Disease CFU Reactivation of infection Immune conversion Latency? Latent infection Elimination? 1-3 4-50 Years after exposure Alteration of antigen recognition as disease progresses (ET) 10000 1000 100 IFN-g (pg/ml) 10 TB HHC 6000 LTBI RV2031c response ESAT-6 response in clinical groups p<0.001 p<0.001 4000 R2 = -0.097 R2 = 0.3688 2000 10000 R2 = -1.0634 0 0 1000 2000 3000 ESAT-6 response 1000 100 10 TB HHC LTBI Rv2031c response in clinical groups 4000 5000 CC HHC TB Linear (CC) Linear (HHC) Linear (TB) Alteration of antigen recognition as disease progresses (Ga and NL) Slope of linear regression no. of spots from ESAT-6 stimulation vs Rv2031c 0.35 3.0 2.5 2.0 1.5 1.0 0.5 0.0 TB HHC CC Clinical status of participants from Ethiopia 0.30 0.25 0.20 0.15 0.10 0.05 0.00 TB HHC 0.0005 0.0000 TB CC Clinical status of participants from The Gambia 0.0010 Slope of linear regression IFN-g from ESAT-6 stimulation vs Rv2031c Slope of linear regression no. of spots from ESAT-6 stimulation vs Rv2031c 3.5 TST+ Clinical status of participants from the Netherlands A lowered ratio of ESAT-6 immune reactivity to Rv2031c reactivity reflects a shift from acute to latent TB People who fail to control bacterial replication become ESAT+ and get TB Time after infection Rv2031c ESAT-6 Rv2031c ESAT-6 Rv2031c ESAT-6 CFU “Clinical disease” threshold “positivity” threshold People who fail to control initial bacterial replication become ESAT+, but if they control later infection, become latently infected People who control initial bacterial replication remain ESAT-, and may or may not be latently infected Response to infection (the immunologist’s view) 67% Initial exposure 33% These individuals do not apparently skin-test convert or become ESAT-6 positive Early bacterial growth arrested at early time point. May (or may not) result in latent infection Early bacterial growth not contained. Leads to clinical illness 9% Subsequent bacterial growth contained. Symptoms abate but latent infection established. 22% Remain healthy but latently infected 2% Bacterial growth not contained. Progressive disease unless treated These individuals generally skin-test convert and become ESAT-6 positive. They often have characteristic patterns on X-ray. 24% Immunologically, little is known about these individuals as they cannot be distinguished from uninfected individuals Reactivation of latent infection at a later point in life Immunologically these individuals tend to express elevated levels of IL-4 and in advanced disease, decreased IFN-g and IL-12. They weakly recognise Rv2031c Immunologically, these individuals tend to express elevated levels of IFN-g and IL-12, and while IL-4 often remains slightly increased, its antagonist IL-4d2 is greatly increased. They strongly recognise Rv2031c Summary • Immunity to M. tuberculosis is dependent on the generation of Th1 immunity, particularly IL-12, IFN-g and TNF-a • As the bacteria persists in the face of this Th1 response, it begins to alter its proteome towards a pattern characteristic of latency, downregulating some antigens, upregulating others • At the same time, a Th2 response seems to develop • Susceptibility to infection therefore appears to correlate not so much with inability to generate a Th1 response, as with inability to maintain it long term, or perhaps inability to direct it to relevant antigens • We are starting to see evidence that M. tuberculosis-derived antigens are driving some of this Th2 response Identification of CD8+ epitopes • Vaccines4TB • Vaccines against tuberculosis are urgently needed. CD4 T cell responses play a major role in the generation of acquired immunity against M. tuberculosis. However, it is increasingly recognised that CD8 cytotoxic T cells (CTL) also contribute to optimal host defence against mycobacteria. • Unfortunately, relatively few CTL responses against TB have been identified. • http://ec.europa.eu/research/health/povertydiseases/projects/110_en.htm Sheila Tang Cellular immune response Tubercle bacilli enter aveoli IFN-g Tubercle bacilli TB peptide Infect* macrophages TCR Within few weeks Lysosome Th1 immune response +TB ER CD4+/CD8+ T cells Recruit to lung Cytokines: IL-2, TNFa and IFN-y CD8 T cells MACROPHAGE Cellular immune response Granulomas prevent spread of infection by confining bacteria within a compact collection of several types of immune cells and activated macrophages Role of these cells: specific ways to isolate inhibit the replication of, and destroy the bacteria http://www.granuloma.homestead.com/tb_microscopic.html • Bacilli engulfed by macrophages • Replicate within the macrophages 2-3 weeks before spreading throughout the body • 95% contain the bacteria in macrophages But due to Mtb. complex waxy cell wall the bacteria are protected inside the macrophages TB genome. Where to look? Epitope Prediction TBVAC epitopes(14) Proteins with CD8 epitopes(25) Proteins from vaccine trials Selected in proteins -previously described by other groups to have CTL epitopes (Michel Klein WP3) (Michel Klein WP3) 3 epitopes/protein used in vaccine trials (21) Additional epitopes from proteins with CD8 epitopes (43) Peptides TBVAC peptides: Ag85A/B, ESAT6, PPE, HBHA TB-CD8 peptides: Mycobacteria tuberculosis H37Rv strain 1 2 3 4 5 6 7 8 9 10 11 1 2 3 4 5 6 7 8 9 10 11 12 13 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Peptides (A2)TB-VAC #108-50 (A2)TB-VAC #108-51 (A2)TB-VAC #108-52 (A2)CD8 #108-63 (A2)CD8 #108-64 (A2)CD8 #108-65 (A2)CD8 #108-66 (A2)CD8 #108-67 (A2)CD8 #108-68 (A2)CD8 #108-69 (A2)CD8 #108-70 (A3)TB-VAC #108-53 (A3)TB-VAC #108-54 (A3)TB-VAC #108-55 (A3)TB-VAC #108-56 (A3)CD8 #108-71 (A3)CD8 #108-72 (A3)CD8 #108-73 (A3)CD8 #108-74 (A3)CD8 #108-75 (A3)CD8 #108-76 (A3)CD8 #108-77 (A3)CD8 #108-78 (A3)CD8 #108-79 (B7)TB-VAC #108-57 (B7)TB-VAC #108-58 (B7)TB-VAC #108-59 (B7)TB-VAC #108-60 (B7)TB-VAC #108-61 (B7)TB-VAC #108-62 (B7)CD8 #108-80 (B7)CD8 #108-81 (B7)CD8 #108-82 (B7)CD8 #108-83 (B7)CD8 #108-84 (B7)CD8 #108-85 (B7)CD8 #108-86 (B7)CD8 #108-87 ESAT6 Ag85B PPE H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv Ag85B HBHA Ag85A HBHA H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv Ag85A HBHA Ag85B HBHA Ag85A HBHA H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv Sequence LLDEGKQSL GLAGGAATA LLGQNTPAI VLMGGVPGV GLLDVTDNV SMLPPGYPV YLAEGHACL HLSGPLAGV YIMKLHHLV LLHDIGKPV SLYEKSGSZ AVYLLDGLR KLVGIELPK ALYLLDGLR QSFEEVSAR TVGYMYIMK ATFEAVLAK RTEILGLVK ATIEAVLAK KIMDYGKYK QINELHHSK KYFVRSTEK GTFKSVAVK SVFPFDGTR RVRGAVTGM APAKKAAPA RAWGRRLMI RVEESRARL MPVGGQSSF APAKKAAAK RARKRGITM RARKRGITL RPKPDYSAM KPIPHRTVL RVRQAWDTL TPVEHGLVL KVRGRLLAL LPAQLTATA Antigen SECRETED L-ALANINE DEHYDROGENASE ALD PROBABLE NAD-DEPENDENT GLUTAMATE DEHYDROGENASE GDH (NAD-GDH) PROBABLE CONSERVED TRANSMEMBRANE PROTEIN hypothetical protein Rv1461 ISONIAZID INDUCTIBLE GENE PROTEIN INIB DNA-DIRECTED RNA POLYMERASE hypothetical protein Rv2823c hypothetical protein Rv1461 DNA-DIRECTED RNA POLYMERASE hypothetical protein Rv0094c PROBABLE NAD-DEPENDENT GLUTAMATE DEHYDROGENASE GDH (NAD-GDH) hypothetical protein Rv1148c PROBABLE INITIATION FACTOR IF-3 INFC SUPEROXIDE DISMUTASE [FE] SODA hypothetical protein Rv1461 60 KDA CHAPERONIN 2 GROEL2, HEAT SHOCK PROTEIN 65 hypothetical protein Rv3378c hypothetical protein Rv1148c hypothetical protein Rv0094c hypothetical protein Rv3378c hypothetical protein Rv1461 hypothetical protein Rv1073 ISONIAZID INDUCTIBLE GENE PROTEIN INIB PROBABLE CONSERVED TRANSMEMBRANE PROTEIN hypothetical protein Rv1148c Analysing peptide screening CD8+ antigen Specific cells R4 CD8 100 CD3 101 102 FL 3-CD3 Pcp 103 CD4+CD8 cells 104 peptides donor 45 4 0 peptides #108-65 #108-64 #108-63 #108-70 0 #108-70 2 #108-69 2 #108-69 4 #108-68 6 #108-68 8 #108-67 8 #108-67 Donor 33 #108-66 peptides #108-66 10 #108-65 peptides #108-64 10 #108-52 Donor 40 #108-63 0 #108-52 2 #108-51 8 #108-51 8 #108-50 4 PPD 6 % CFSE 10 #108-50 6 % CFSE #108-70 #108-69 #108-68 #108-67 #108-66 #108-65 #108-64 #108-63 #108-52 #108-51 #108-50 PPD % CFSE 10 PPD #108-70 #108-69 #108-68 #108-67 #108-66 #108-65 #108-64 #108-63 #108-52 #108-51 #108-50 PPD % CFSE CD8 T cell proliferation to A2 motif bearing peptides Donor 42 6 4 2 0 Peptide Name TB.o42.epitopes A2 11613 MHC A2 TB.o42.epitopes A2 11630 TB.o42.epitopes A2 11633 A2 A2 11630 11633 TB.o42.epitopes A2 11623 TB.o42.cons A2 11699 TB.o42.cons A2 11687 TB.O42.CONS A2 11690 A2 A2 A2 A2 11623 11699 11687 11690 TB-CD8(A2) 10864 TB-CD8(A2) 10865 A2 A2 10864 10865 TB-CD8(A2) 10868 TB.o42.cons A2 11691 TB.o42.cons A2 11697 TB-CD8(A2) 10866 TB.o42.cons A2 11693 TB-VAC(A2) 10850 TB.o42.cons A2 11689 TB-VAC(A2) 10852 TB-VAC(A2) 10851 TB-CD8(A2) 10863 TB-CD8(A2) 10867 A2 A2 A2 A2 A2 A2 A2 A2 A2 A2 A2 10868 11691 11697 10866 11693 10850 11689 10852 10851 10863 10867 Pep.no. 11613 Sequence 10/10-05(nM) YMLDMTFPV 1 FLQGAKWYL 2 YLAENTFVV 3 WMYEGKHVL 5 LLDEPTNHL 17 FLFGDDDAL 20 VLDEPSIGL 20 GLLDVTDNV 22 SMLPPGYPV 25 YIMKLHHLV 33 LLDEPTNNL 33 DMWEHAFYL 34 YLAEGHACL 46 RMWEFLDRL 56 LLDEGKQSL 86 LLLGGTSEI 105 LLGQNTPAI 116 GLAGGAATA 853 VLMGGVPGV non HLSGPLAGV non peptides 0 peptides peptides Donor 50 10 peptides #108-79 #108-78 #108-77 #108-76 #108-75 #108-74 #108-73 #108-72 Donor 53 #108-71 #108-56 #108-55 15 #108-54 % CFSE 15 #108-53 PPD #108-79 #108-78 #108-77 #108-76 #108-75 #108-74 15 #108-79 0 #108-73 #108-72 #108-71 20 #108-78 0 #108-56 20 #108-77 0 #108-55 20 #108-76 5 #108-75 5 #108-54 10 #108-74 5 #108-73 15 #108-72 Donor 59 #108-71 15 #108-56 20 #108-55 20 #108-54 peptides #108-53 10 % CFSE 5 #108-53 % CFSE Donor 47 PPD #108-79 #108-78 #108-77 PPD 0 #108-76 5 #108-75 #108-79 #108-78 #108-77 #108-76 #108-75 #108-74 #108-73 #108-72 #108-71 10 #108-74 #108-73 #108-72 #108-71 #108-56 #108-55 #108-54 #108-53 PPD % CFSE #108-56 #108-55 #108-54 #108-53 PPD % CFSE CD8 T cell proliferation to A3 motif bearing peptides Donor 49 10 A3-peptides. Binding versus peptide immunogenicity T B.o42.c ons A3 11705 A3 11705 T B.o42.c ons A3 11706 A3 11706 T B- CD8(A3) 10878 A3 10878 T B.o42.epitopes A3 11643 A3 11643 T B- CD8(A3) 10875 A3 10875 T B.o42.epitopes A3 11640 A3 11640 T B.o42.epitopes A3 11653 A3 11653 T B.o42.c ons A3 11719 A3 11719 T B- CD8(A3) 10874 A3 10874 T B.o42.c ons A3 11700 A3 11700 T B.o42.c ons A3 11707 A3 11707 T B.o42.c ons A3 11708 A3 11708 T B- CD8(A3) 10871 A3 10871 T B- CD8(A3) 10876 A3 10876 T B- CB8(A3) 10877 A3 10877 T B.o42.epitopes A3 11648 A3 11648 T B.o42.epitopes A3 11645 A3 11645 T B.o42.epitopes A3 11644 A3 11644 T B.o42epitopes A3 11639 A3 11639 T B.o42.c ons A3 11711 A3 11711 T B.o42.c ons A3 11716 A3 11716 T B.o42.epitopes A3 11642 A3 11642 T B.o42.epitopes A3 11634 A3 11634 T B- VAC(A3) 10855 A3 10855 T B- VAC(A3) 10856 A3 10856 T B.o42.c ons A3 11720 A3 11720 RVYLQGHGY TLLESFLFY GTFKSVAVK RVFGFRTAK KIMDYGKYK RVMPVFAFK RVYLNGIGK ALFDRPAFK ATIEAVLAK AVHGYYIGY KLMALELFK KLYPNVDFY TVGYMYIMK QINELHHSK KYFVRSTEK ATFEVFLAK AVFPRYHPR AVFDSFVER AVMLVHTYY KIGEVIGPK QVFKGVVIR YVYPDNLPR AVFLSYIGY ALYLLDGLR QSFEEVSAR NIMEFCKAY 3 4 29 37 41 53 66 78 104 109 109 178 318 335 750 913 930 1003 1328 1500 3258 5618 >5000 non non non 3/5 donor recognition 0 peptides peptides donor 60 25 20 15 10 5 peptides #108-87 #108-86 #108-85 #108-84 #108-83 #108-82 #108-81 #108-80 #108-62 #108-61 #108-60 #108-59 #108-58 donor 53 #108-87 #108-86 0 #108-85 0 #108-84 5 #108-57 20 #108-83 20 PPD 25 #108-82 10 % CFSE 25 #108-81 #108-87 #108-86 #108-85 #108-84 #108-83 #108-82 #108-81 #108-80 #108-62 #108-61 #108-60 #108-59 #108-58 #108-57 15 #108-80 #108-62 #108-61 #108-60 #108-59 #108-58 #108-57 PPD % CFSE 5 PPD % CFSE CD8 T cell proliferation to B7 motif bearing peptides donor 59 15 10 B7-peptides Binding versus peptide immunogenicity TB.o42.epitopes B7 11669 TB-CD8(B7) 10886 TB.o42.epitopes B7 11658 TB-CD8(B7) 10881 TB.o42epitopes B7 11666 TB-VAC(B7) 10859 TB.o42.epitopes B7 11660 TB.o42.cons B7 11735 TB.o42.cons B7 11729 TB.o42.epitopes B7 11667 TB-CD8(B7) 10883 TB.o42.cons B7 11724 TB-VAC(B7) 10861 TB.o42.epitopes B7 11661 TB.o42.epitopes B7 11656 TB.o42.epitopes B7 11665 TB-CD8(B7) 10884 TB.o42.epitopes B7 11662 TB.o42.epitopes B7 11668 TB-CD8(B7) 10882 TB-VAC(B7) 10858 TB-CD8(B7) 10885 TB.o42.cons B7 11746 TB-CD8(B7) 10880 TB.o42.epitopes B7 11659 TB.o42.epitopes B7 11657 TB.o42.cons B7 11731 TB-VAC(B7) 10857 TB.o42.cons B7 11741 TB-VAC(B7) 10860 TB.o42.cons B7 11733 TB-VAC(B7) 10862 TB.o42.cons B7 11738 TB-CD8(B7) 10887-2 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 B7 11669 10886 11658 10881 11666 10859 11660 11735 11729 11667 10883 11724 10861 11661 11656 11665 10884 11662 11668 10882 10858 10885 11746 10880 11659 11657 11731 10857 11741 10860 11733 10862 11738 10887-2 SPRSRNRSF KVRGRLLAL RPRQRGIPF RARKRGITL IPRLGGMAF RAWGRRLMI RPVFARLPF GPAFVRTKL GPRGRHVVL RPRVAQLTF KPIPHRTVL RIRSERPAF MPVGGQSSF VPADHRLAF RPAGARAAF VPRENATAF RVRQAWDTL VPRDRNGTF LPAEVRAAF RPKPDYSAM APAKKAAPA TPVEHGLVL TPRIANRLL RARKRGITM APRGFRAAF APRARTAAF IPAPGLGAL RVRGAVTGM MPRLSRNAA RVEESRARL TPALATRGF APAKKAAAK YPACEAIGL LPAQLTATA 0,3 0,4 0,4 0,4 0,4 0,4 0,6 0,9 1,0 1,2 1,4 1,5 1,8 1,9 2,0 2,0 2,1 2,6 2,8 2,8 3,0 3,2 3,9 4,5 4,6 5,6 5,9 6,5 10,3 38,0 318,0 346,5 436,0 >5000 B7 peptides #108-87 #108-86 #108-85 #108-84 2 2 0 0 10 8 4 2 0 Y 6 • No responses to peptides #108-79 #108-78 #108-77 #108-76 8 #108-75 #108-74 BCPP2 #108-73 #108-72 #108-71 donor 37 #108-56 4 #108-55 A2 donors #108-54 6 % CFSE 10 #108-53 PPD #108-70 #108-69 #108-68 #108-67 #108-66 8 #108-83 #108-82 #108-65 BCPP1 #108-81 #108-80 #108-64 #108-63 #108-52 #108-51 #108-50 PPD % CFSE Donor 43 #108-62 #108-61 #108-60 #108-59 #108-58 #108-57 PPD % CFSE PPD-ve individuals do not respond to peptides 10 Donor 56 A3 donors 6 4 A2 peptides A3 peptides donor 46 B7 donors • PPD responses < 1% cfse+ve Peptides Recognised by CD8 T cells B7 peptides A3 peptides A2 peptides 1 2 3 4 5 6 7 8 9 10 11 (A2)TB-VAC #108-50 (A2)TB-VAC #108-51 (A2)TB-VAC #108-52 (A2)CD8 #108-63 (A2)CD8 #108-64 (A2)CD8 #108-65 (A2)CD8 #108-66 (A2)CD8 #108-67 (A2)CD8 #108-68 (A2)CD8 #108-69 (A2)CD8 #108-70 4/11 (36%) ESAT6 Ag85B PPE H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv 1 2 3 4 5 6 7 8 9 10 11 12 13 (A3)TB-VAC #108-53 (A3)TB-VAC #108-54 (A3)TB-VAC #108-55 (A3)TB-VAC #108-56 (A3)CD8 #108-71 (A3)CD8 #108-72 (A3)CD8 #108-73 (A3)CD8 #108-74 (A3)CD8 #108-75 (A3)CD8 #108-76 (A3)CD8 #108-77 (A3)CD8 #108-78 (A3)CD8 #108-79 Ag85B HBHA Ag85A HBHA H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv 9/13 (70%) 1 2 3 4 5 6 7 8 9 10 11 12 13 14 (B7)TB-VAC #108-57 (B7)TB-VAC #108-58 (B7)TB-VAC #108-59 (B7)TB-VAC #108-60 (B7)TB-VAC #108-61 (B7)TB-VAC #108-62 (B7)CD8 #108-80 (B7)CD8 #108-81 (B7)CD8 #108-82 (B7)CD8 #108-83 (B7)CD8 #108-84 (B7)CD8 #108-85 (B7)CD8 #108-86 (B7)CD8 #108-87 Ag85A HBHA Ag85B HBHA Ag85A HBHA H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv H37Rv 6/14 (43%) SUMMARY • 19/38 predicted peptides induced a CD8 proliferative response • The frequency of proliferating CD8 T cell response to peptides varied between individuals • Heterogeneous response to peptides • For A3-peptide responses, 3/5 donors recognised the same peptide: QINELHHSK (CD8-#108-76), suggesting it may be immunodominant peptide Acknowledgements. Vaccines4TB Prof. Dr. Tom Ottenhoff Michel Klein Tuberculosis group Immunological bioinformatic group Immunohematology and Blood Transfusion CBS-BioCentrum, DTU Leiden University Medical Center Techinical University of Denmark Leiden, Netherlands In silico peptide prediction, NetCTL Proliferation assays, FACS analysis and IFN-g-ELISA Leucosep Isolation of PBMC Tom Søren Buus, MD, Ph.D Prof. IMMI, University of Copenhagen MHC binding Ugur Sahin Fatima Kazi Ganymed Pascale van Weeren Genetic library And the rest of the Ottenhoff’s group