* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download CBS_Nov_22_05
Major histocompatibility complex wikipedia , lookup
Psychoneuroimmunology wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Marburg virus disease wikipedia , lookup
Infection control wikipedia , lookup
Herd immunity wikipedia , lookup
Eradication of infectious diseases wikipedia , lookup
Childhood immunizations in the United States wikipedia , lookup
Vaccination policy wikipedia , lookup
Human leukocyte antigen wikipedia , lookup
Hepatitis B wikipedia , lookup
DNA vaccination wikipedia , lookup
Molecular mimicry wikipedia , lookup
Transmission (medicine) wikipedia , lookup
Germ theory of disease wikipedia , lookup
Immunocontraception wikipedia , lookup
Sociality and disease transmission wikipedia , lookup
Globalization and disease wikipedia , lookup
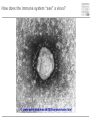
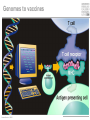
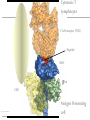
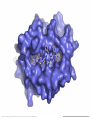
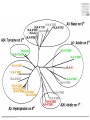
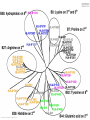
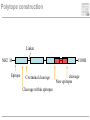
Immunological Bioinformatics Ole Lund Center for Biological Sequence Analysis BioCentrum-DTU Technical University of Denmark [email protected] Challenges and failures of the immune system Outside Infection with microbe A Infection with microbe B Allergen -> allergy Peptide drugs Time Creation Creation of self of an immune system/ Tolerance to self Autoimmunity (break of tolerance to self) Inside Cancer How does the immune system “see” a virus? Immune system overview •Innate – fast, unspecific •Addaptive – specific, remembers… •Cellular •Cytotoxic T lymphocytes (CTL) •Helper T lymphocytes (HTL) •Humoral •B lymphocytes Figures by Eric A.J. Reits MHC Class I pathway Response to 1:(5x20x200) = 1:2000 9mers 1:5 1:200 Figure by Eric A.J. Reits 1:2 Genomes to vaccines Lauemøller et al., 2000 Cytotoxic T Lymphocyte T Cell receptor (TCR) Peptide MHC b2m CD8 Fugure by Thomas Blicher Antigen Presenting cell Figure by Anne Mølgaard, peptide (KVDDTFYYV) used as vaccine by Snyder et al. J Virol 78, 7052-60 (2004). Influenza A virus (A/Goose/Guangdong/1/96(H5N1)) Genome >Segment 1 agcaaaagcaggtcaattatattcaatatggaaagaataaaagaactaagagatctaatg tcgcagtcccgcactcgcgagatactaacaaaaaccactgtggatcatatggccataatc aagaaatacacatcaggaagacaagagaagaaccctgctctcagaatgaaatggatgatg gcaatgaaatatccaatcacagcagacaagagaataatggagatgattcctgaaaggaat and 13350 other nucleotides on 8 segments Proteins 9mer peptides >polymerase“ MERIKELRD MERIKELRDLMSQSRTREILTKTTVDHMAIIKKYTSGRQEKNPALRMKWMMAMKYPITAD ERIKELRDL KRIMEMIPERNEQGQTLWSKTNDAGSDRVMVSPLAVTWWNRNGPTTSTVHYPKVYKTYFE RIKELRDLM KVERLKHGTFGPVHFRNQVKIRRRVDINPGHADLSAKEAQDVIMEVVFPNEVGARILTSE IKELRDLMS SQLTITKEKKEELQDCKIAPLMVAYMLERELVRKTRFLPVAGGTSSVYIEVLHLTQGTCW KELRDLMSQ EQMYTPGGEVRNDDVDQSLIIAARNIVRRATVSADPLASLLEMCHSTQIGGIRMVDILRQ ELRDLMSQS NPTEEQAVDICKAAMGLRISSSFSFGGFTFKRTNGSSVKKEEEVLTGNLQTLKIKVHEGY LRDLMSQSR EEFTMVGRRATAILRKATRRLIQLIVSGRDEQSIAEAIIVAMVFSQEDCMIKAVRGDLNF RDLMSQSRT ... DLMSQSRTR LMSQSRTRE and 9 other proteins and 4376 other 9mers Weight matrices (Hidden Markov models) YMNGTMSQV GILGFVFTL ALWGFFPVV ILKEPVHGV ILGFVFTLT LLFGYPVYV GLSPTVWLS WLSLLVPFV FLPSDFFPS CVGGLLTMV FIAGNSAYE A2 Logo G F C A Lauemøller et al., 2000 Human Leukocyte antigen (HLA=MHC in humans) polymorphism - alleles http://www.anthonynolan.com/HIG/index.html HLA polymorphism - supertypes •Each HLA molecule within a supertype essentially binds the same peptides •Nine major HLA class I supertypes have been defined •HLA-A1, A2, A3, A24,B7, B27, B44, B58, B62 Sette et al, Immunogenetics (1999) 50:201-212 HLA polymorphism - frequencies Supertypes Phenotype frequencies Caucasian Black Japanese A2,A3, B7 83 % 86 % 88 % 88 % 86 % 86% +A1, A24, B44 100 % 98 % 100 % 100 % 99 % 99 % +B27, B58, B62 100 % 100 % 100 % 100 % 100 % 100 % A Sette et al, Immunogenetics (1999) 50:201-212 Chinese Hispanic Average O Lund et al., Immunogenetics. 2004 55:797-810 O Lund et al., Immunogenetics. 2004 55:797-810 O Lund et al., Immunogenetics. 2004 55:797-810 Combined method •Combining predicted MHC-I affinity with prediction of Cterminal proteasomal cleavage and TAP transport efficiency improves the ability to identify known CTL epitopes MV Larsen et al., Accepted for publication in European Journal of Immunology Infectious Diseases •More than 400 microbial agents are associated with disease in healthy adult humans •There are only licensed vaccines in the United states for 22 microbial agents (vaccines for 34 pathogens have been developed) •Immunological Bioinformatics may be used to •Identify immunogenic regions in pathogen •These regions may be used as in rational vaccine design •Which pathogens to focus on? Infectious diseases may be ranked based on •Impact on health •Dangerousness •Economic impact Infectious Diseases in the World •11 million (19%) of the 57 million people who died in the world in 2002 were killed by infectious or parasitic infection [WHO, 2004] •The three main single infectious diseases are HIV/AIDS, tuberculosis, and malaria, each of which causes more than 1 million deaths Deaths from infectious diseases in the world in 2002 www.who.int/entity/whr/2004/annex/topic/en/annex_2_en.pdf Dodo Pathogenic Viruses 1st column: log10 of the number of deaths caused by the pathogen per year 2nd column: DNA Advisory Committee (RAC) classification DNA Advisory Committee guidelines [RAC, 2002] which includes those biological agents known to infect humans, as well as selected animal agents that may pose theoretical risks if inoculated into humans. RAC divides pathogens into four classes. Risk group 1 (RG1). Agents that are not associated with disease in healthy adult humans Risk group 2 (RG2). Agents that are associated with human disease which is rarely serious and for which preventive or therapeutic interventions are often available Risk group 3 (RG3). Agents that are associated with serious or lethal human disease for which preventive or therapeutic interventions may be available (high individual risk but low community risk) Risk group 4 (RG4). Agents that are likely to cause serious or lethal human disease for which preventive or therapeutic interventions are not usually available (high individual risk and high community risk) 3rd column: CDC/NIAID bioterror classification classification of the pathogens according to the Centers for Disease Control and Prevention (CDC) bioterror categories A–C, where category A pathogens are considered the worst bioterror threats 4th column: Vaccines available A letter indicating the type of vaccine if one is available (A: acellular/adsorbet; C: conjugate; I: inactivated; L: live; P: polysaccharide; R: recombinant; S staphage lysate; T: toxoid). Lower case indicates that the vaccine is released as an investigational new drug (IND)). 5th column: G: Complete genome is sequenced Data derived from /www.cbs.dtu.dk/databases/Dodo. Biodefence Targets www2.niaid.nih.gov/Biodefense/ bandc_priority.htm NIH project Pathogen HLA binding Elispot Influenza X X Variola major (smallpox) vaccine strain X X Yersinia pestis X Francisella tularensis (tularemia) X LCM X Lassa Fever X Hantaan virus (Korean hemorrhagic fever virus) X Rift Valley Fever X Dengue X Ebola X Marburg X Multi-drug resistant TB (BCG vaccine) X Yellow fever X Typhus fever (Rickettsia prowazekii) X West Nile Virus X CBS/panum X Strategy for determination of peptide-HLA binding Step I: Folding of MHC class I molecules in solution b2m Heavy chain peptide Incubation Peptide-MHC complex Step II: Detection of de novo folded MHC class I molecules by ELISA Development C Sylvester-Hvid et al., Tissue Antigens. 2002 59:251-8 ELISPOT assay •Measure number of white blood cells that in vitro produce interferon-g in response to a peptide •A positive result means that the immune system have earlier reacted to the peptide (during a response ot a vaccine/natural infection) SLFNTVATL SLFNTVATL SLFNTVATL SLFNTVATL SLFNTVATL SLFNTVATL Two spots Preliminary results •167 peptides have so far been tested for binding to a HLA molecule •113 of these (67%) have been shown to bind to the relevant HLA allele with a affinity better than 500nM •180 predicted epitopes from influenza A virus were tested in an ELISPOT assay •12 were so far found to be epitopes (recognized by donors previously exposed to Influenza) •14% of peptides binding with an affinity better than 500nM were found to be epitopes •1:2000 randomly chosen peptides are epitopes Vaccination •Vaccination •Administration of a substance to a person with the purpose of preventing a disease •Traditionally composed of a killed or weakened microorganism •Vaccination works by creating a type of immune response that enables the memory cells to later respond to a similar organism before it can cause disease Early History of Vaccination •Pioneered India and China in the 17th century •The tradition of vaccination may have originated in India in AD 1000 •Powdered scabs from people infected with smallpox was used to protect against the disease •Smallpox was responsible for 8 to 20% of all deaths in several European countries in the 18th century •In 1721 Lady Mary Wortley Montagu brought the knowledge of these techniques from Constantinople (now Istanbul) to England •Two to three percent of the smallpox vaccinees, however, died from the vaccination itself •Benjamin Jesty and, later, Edward Jenner could show that vaccination with the less dangerous cowpox could protect against infection with smallpox •The word vaccination, which is derived from vacca, the Latin word for cow. Vaccination Today •Vaccines have been made for only 34 of the more than 400 known pathogens that are harmful to man. •Immunization saves the lives of 3 million children each year, but that 2 million more lives could be saved if existing vaccines were applied on a full-scale worldwide Human Vaccines against pathogens Immunological Bioinformatics, The MIT press. Categories of Vaccines •Live vaccines •Are able to replicate in the host •Attenuated (weakened) so they do not cause disease •Subunit vaccines •Part of organism •Genetic Vaccines •Part of genes from organism Polytope construction Linker NH2 M Epitope COOH C-terminal cleavage Cleavage within epitopes cleavage New epitopes Helper responses Figures by Eric A.J. Reits Figure by Anne Mølgaard MHC class II prediction Complexity of problem – Peptides of different length – Weak motif signal Alignment crucial Gibbs Monte Carlo sampler M Nielsen et al., Bioinformatics. 2004 20:1388-97 RFFGGDRGAPKRG YLDPLIRGLLARPAKLQV KPGQPPRLLIYDASNRATGIPA GSLFVYNITTNKYKAFLDKQ SALLSSDITASVNCAK PKYVHQNTLKLAT GFKGEQGPKGEP DVFKELKVHHANENI SRYWAIRTRSGGI TYSTNEIDLQLSQEDGQTIE Class II binding motif Alignment by Gibbs sampler RFFGGDRGAPKRG YLDPLIRGLLARPAKLQV KPGQPPRLLIYDASNRATGIPA GSLFVYNITTNKYKAFLDKQ SALLSSDITASVNCAK PKYVHQNTLKLAT GFKGEQGPKGEP DVFKELKVHHANENI SRYWAIRTRSGGI TYSTNEIDLQLSQEDGQTI M Nielsen et al., Bioinformatics. 2004 20:1388-97 Random ClustalW Gibbs sampler Antibody responses Figures by Eric A.J. Reits Prediction of Antibody epitopes Linear – Hydrophilicity scales (average in ~7 window) • Hoop and Woods (1981) • Kyte and Doolittle (1982) • Parker et al. (1986) – Other scales & combinations • Pellequer and van Regenmortel • Alix – New improved method (Pontoppidan et al. in preparation) • http://www.cbs.dtu.dk/services/BepiPred/ Discontinuous – Protrusion (Novotny, Thornton, 1986) • Pernille Haste Andersen, 2005, in preparation Immunological bioinformatics Classical experimental research – Few data points – Data recorded by pencil and paper/spreadsheet New experimental methods – Sequencing – DNA arrays – Proteomics Need to develop new methods for handling these large data sets • Immunological Bioinformatics/Immunoinformatics Sune Frankild Jens Pontoppidan Morten Nielsen Pernille Haste Andersen Thomas Blicher Claus Lundegaard Anne Mølgaard Xiuxiu Ye