* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Infection T Cell Response during Chronic Viral +CD8 Exhaustion
Survey
Document related concepts
Immunocontraception wikipedia , lookup
Adaptive immune system wikipedia , lookup
Neonatal infection wikipedia , lookup
Psychoneuroimmunology wikipedia , lookup
Childhood immunizations in the United States wikipedia , lookup
Infection control wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Vaccination wikipedia , lookup
Innate immune system wikipedia , lookup
Henipavirus wikipedia , lookup
Human cytomegalovirus wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
DNA vaccination wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Transcript
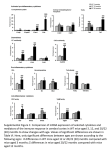
This information is current as of June 18, 2017. Single-Epitope DNA Vaccination Prevents Exhaustion and Facilitates a Broad Antiviral CD8+ T Cell Response during Chronic Viral Infection Christina Bartholdy, Anette Stryhn, Jan Pravsgaard Christensen and Allan Randrup Thomsen J Immunol 2004; 173:6284-6293; ; doi: 10.4049/jimmunol.173.10.6284 http://www.jimmunol.org/content/173/10/6284 Subscription Permissions Email Alerts This article cites 58 articles, 29 of which you can access for free at: http://www.jimmunol.org/content/173/10/6284.full#ref-list-1 Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2004 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 References The Journal of Immunology Single-Epitope DNA Vaccination Prevents Exhaustion and Facilitates a Broad Antiviral CD8ⴙ T Cell Response during Chronic Viral Infection1 Christina Bartholdy, Anette Stryhn,2 Jan Pravsgaard Christensen, and Allan Randrup Thomsen3 he CD8⫹ T lymphocytes play a central role in the defense against most viral infections. Via their TCR, they recognize foreign peptides bound to the MHC class I molecules. CD8⫹ T cell responses to foreign proteins are generally focused on a few or sometimes only a single immunogenic epitope (1–5), although CD8⫹ T cells specific for several subdominant epitopes often exist. These subdominant responses are under normal circumstances suppressed by the response to the dominant epitope(s), but may become highly relevant in the absence or silencing of the dominant response (6 –9). The fact that the CD8⫹ T cell response is normally quite focused has been exploited in the design of vaccines, in which only immunodominant parts of a given pathogen are used for immunization. Such vaccines have consisted of either synthetic peptides administered together with adjuvant or plasmid DNA (DNA vaccination). The latter offers several advantages, including simplicity, de novo synthesis of endogenous protein in the host, and adjuvancity due to CpG motifs T Institute of Medical Microbiology and Immunology, University of Copenhagen, Copenhagen, Denmark Received for publication June 8, 2004. Accepted for publication September 2, 2004. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. 1 This work was supported in part by the Danish Medical Research Council, the Biotechnology Center for Cellular Communication, the Haensch Foundation, and the Novo Nordisk Foundation. C.B. was the recipient of a Ph.D. scholarship from the Faculty of Health Sciences, University of Copenhagen (Copenhagen, Denmark). A.S. was supported by postdoctoral fellowships from the Faculty of Natural Science (Curie fellowship) and the Carlsberg Foundation. J.P.C. was supported by a senior research fellowship from the Benzon Foundation. 2 contained in the vector (10). DNA as well as peptide vaccines that prime CD8⫹ T cells specific for a single immunodominant epitope have been developed. Such single-epitope vaccines have to date primarily been evaluated in a rather simple context, with a focus on their ability to protect against primary infection. However, some risks may be associated with single-epitope vaccines due to the narrow immune response they induce. This includes selection of viral escape variants and/or increased susceptibility to reinfection with viral variants (11). A special characteristic of viruses, especially those belonging to the RNA subgroup, is their ability to mutate. As a consequence of that, natural variants, including immune escape variants, are easily generated (12). Viral escape from CD8⫹ T cell-mediated control has been observed for many viruses, including influenza and HIV (13–16). Inducing a monospecific immune response by single-epitope vaccination might decrease the selection pressure against these variants during subsequent infection. This would be particularly relevant during chronic viral infection, because the virus remains in the host for a longer period and thus has more time to mutate. It has been observed in several animal models that an initial monospecific, CD8⫹ T cell response increases the chance for selection of viral CD8⫹ T cell escape variants upon infection. Examples are vaccinated macaques infected with SIV (17, 18), TCR transgenic mice infected with influenza virus, and vaccinated mice as well as TCR transgenic mice infected with lymphocytic choriomeningitis virus (LCMV)4 (19 –21). A monospecific, CD8⫹ T cell response not only might select for viral variants in the individual host, but additionally increases the likelihood of such variants persisting and circulating in the host Current address: T-cellic, DK-2970 Hoersholm, Denmark. 3 Address correspondence and reprint requests to Dr. Allan Randrup Thomsen, Institute of Medical Microbiology and Immunology, The Panum Institute, 3C Blegdamsvej, Copenhagen DK-2200 N, Denmark. E-mail address: [email protected] Copyright © 2004 by The American Association of Immunologists, Inc. 4 Abbreviations used in this paper: LCMV, lymphocytic choriomeningitis virus; h2m, human 2-microlobulin; i.c., intracerebral; LCMV Arm, LCMV Armstrong clone 53b; NP, nucleoprotein; p.i., postinfection. 0022-1767/04/$02.00 Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 Induction of a monospecific antiviral CD8ⴙ T cell response may pose a risk to the host due to the narrow T cell response induced. At the individual level, this may result in selection of CD8ⴙ T cell escape variants, particularly during chronic viral infection. Second, prior immunization toward a single dominant epitope may suppress the response to other viral epitopes, and this may lead to increased susceptibility to reinfection with escape variants circulating in the host population. To address these issues, we induced a memory response consisting solely of monospecific, CD8ⴙ T cells by use of DNA vaccines encoding immunodominant epitopes of lymphocytic choriomeningitis virus (LCMV). We analyzed the spectrum of the CD8ⴙ T cell response and the susceptibility to infection in H-2b and H-2d mice. Priming for a monospecific, CD8ⴙ T cell response did not render mice susceptible to viral variants. Thus, vaccinated mice were protected against chronic infection with LCMV, and no evidence indicating biologically relevant viral escape was obtained. In parallel, a broad and sustained CD8ⴙ T cell response was generated upon infection, and in H-2d mice epitope spreading was observed. Even after acute LCMV infection, DNA vaccination did not significantly impair naturally induced immunity. Thus, the response to the other immunogenic epitopes was not dramatically suppressed in DNAimmunized mice undergoing normal immunizing infection, and the majority of mice were protected against rechallenge with escape variants. These findings underscore that a monospecific vaccine may induce efficient protective immunity given the right set of circumstances. The Journal of Immunology, 2004, 173: 6284 – 6293. The Journal of Immunology Materials and Methods zeo⫹. The OVA258 –265 peptide sequence was exchanged for either the gp33– 41, NP396 – 404, or NP118 –126 sequence using PCR. A PCR product covering the last 28 bases of the leader containing a HindIII site, the peptide, linker, and h2m was generated using as forward primer: for gp33, GCAGCCAAGCTTGACCGGCTTGTATGCTAAGGCCGTGTACAACT TCGCCACCTGCGGTG GTGGTAGTGGGGGA; for NP396, GCAGCC AAGCTTGACCGGCTTGTATGCTTTCCAGCCTCAGAACGGCCAGT TCATCGGTGGTGGTAGTGG GGGA; and for NP118, GCAGCCAA GCTTGACCGGCTTGTATGCTAGGCCCCAGGCTTCAGGGGTATAT ATGGGTGGTGGTAGTGGGGGA.Asbackwardprimer,GCAGCCGCGG CCGCTTACATGTCTCGATCCCACTTA (containing a NotI site) was used for all constructs. PCR products, containing some of the vector, the leader sequence, the peptide sequence, and the linkage to h2m terminated by a stop codon were subsequently cloned as a HindIII/NotI fragment into the template vector. The Escherichia coli strain XL1-blue (Stratagene, La Jolla, CA) was transformed with the constructs by electroporation. DNA sequencing using cycle sequencing, Big Dye Terminator, and ABI310 genetic analyzer (ABI PRISM; Applied Biosystems, Foster City, CA) identified positive clones. Primers were obtained from Hobolth DNA Syntese (Hilleroed, Denmark). Large-scale DNA preparations were produced using Maxi Prep (Qiagen, Valencia, CA). Gene gun immunization DNA was coated onto 1.6-nm gold particles in a concentration of 2 g of DNA/mg gold, and the DNA/gold complex was coated onto plastic tubes such that 0.5 mg of gold was delivered per shot (1 g of DNA/shot). These procedures were performed according to the manufacturer’s instruction (Bio-Rad, Hercules, CA). Mice were immunized on the abdominal skin using a handheld gene gun device using compressed helium (400 psi) as the particle motive force. Mice were inoculated twice with an interval of 4 wk and then allowed to rest for 3 wk before additional challenge. Virus For chronic infection, the vicerotropic LCMV Armstrong strain clone 13 (LCMV clone 13) was used (30, 31). Mice to be challenged systemically received 106 PFU in an i.v. injection of 0.3 ml. For acute transient infection, LCMV Armstrong clone 53b (LCMV Arm) was used (31). Mice to be infected received 104 PFU in an i.v. injection of 0.3 ml. For challenge with escape variants, the LCMV CD8⫹ T cell escape variants gp33-nil and NP396-nil, provided by M. B. A. Oldstone (The Scripps Research Institute, La Jolla, CA), were used (32, 33). The viral variants were derived from LCMV Arm; gp33-nil contains a single mutation in aa 38 (F3 L), and NP396-nil contains a single mutation in aa 403 (F3 L). These mutations result in escape from CD8⫹ T cell recognition (33). Mice to be challenged were given 200 PFU in an intracerebral (i.c.) injection of 0.03 ml. Mice Virus titration C57BL/6 mice and BALB/cJ mice were obtained from Taconic M&B (Ry, Denmark). Seven- to 10-wk-old female mice were used in all experiments, and animals were always allowed to acclimatize to the local environment for at least 1 wk before use. All animals were housed under specific pathogen-free conditions, as validated by screening of sentinels, and the experiments were conducted in compliance with national guidelines. To determine organ virus titers, the organs were first gently homogenized in PBS containing 1% FCS to yield a 10% (v/w) organ suspension. Organ suspensions were clarified by centrifugation, and serial 10-fold dilutions of the supernatants were prepared in PBS with 1% FCS; 0.2 ml of each dilution was then transferred in duplicate to flat-bottom, 24-well plates, and MC57G cells in MEM were added. Plates were incubated for 4 – 6 h at 37°C in 5% CO2 to allow cells to adhere. Subsequently, 0.3 ml of a 1/1 mixture of 2% methylcellulose in double-distilled water and doublestrength MEM with 10% FCS, antibiotics, and glutamine was added. After 48 h, cell monolayers were fixed with 4% formaldehyde in PBS for 20 –30 min at 20°C and permeabilized in 0.5% Triton X-100 in HBSS for 20 min. The next day, monolayers were labeled with a rat anti-LCMV mAb (VL-4) for 60 –90 min, intensively washed, incubated with peroxidase-labeled goat anti-rat Ab for 60 –90 min, and washed again. O-phenylendiamine (substrate) was added for10 –30 min, and the reaction was subsequently terminated by washing with water. The numbers of PFU were counted, and organ virus titers were expressed as PFU per gram of tissue (34). DNA vaccine construction The gp33, nucleoprotein 396 (NP396), and NP118 vaccine are the eukaryotic expression vectors pcDNA3.1/zeo⫹ (Invitrogen, Groningen, The Netherlands) containing the murine 2m leader, followed by either the gp33– 41, the NP396 – 404, or the NP118 –126 LCMV peptide epitope tethered to h2m through a 10-aa linker ((G3S)2GG; Fig. 1). The constructs were generated using as template a similar construct, murine 2m leader-OVA258 – 265-10-aa linker-h2m inserted as a NheI/NotI fragment in pcDNA3.1/ LCMV-induced meningitis studies FIGURE 1. Principal layout of the DNA vaccine constructs used. Only the promotor and the gene insert are shown. The vaccine contains the immediate/early promoter of CMV. The gene insert encodes the murine leader of 2m as a signal sequence (SS), an MHC class I-restricted immunodominant LCMV epitope covalently linked to h2m, and is terminated by a stop codon. The linker consists of 10 unbulky aa ((G3S)2GG). Mortality was used to evaluate the clinical severity of i.c. infection with LCMV variants. Mice were checked twice daily for a period of 14 days or until 100% mortality was reached. After that period, spleens and brains were removed from surviving mice and analyzed for CD8⫹ T cell responses or viral titers, respectively. Cell preparations Spleens were aseptically removed and transferred to HBSS. Single-cell suspensions were obtained by pressing the organs through a fine sterile Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 population. It is obvious that single-epitope vaccination does not protect against a primary infection with a viral variant mutated in the critical epitope. However, single-epitope-vaccinated individuals infected with a wild-type virus might subsequently be more susceptible to viral variants than unvaccinated, wild-type virus-infected individuals due to a more narrow memory response induced during wild-type infection. Thus, priming for a monospecific, CD8⫹ T cell response might influence the immunodominance hierarchy normally established during primary infection, and suppression of CD8⫹ T cells specific for other immunogenic epitopes of the virus is likely to be observed. If such individuals are later exposed to a virus variant mutated in the epitope used for vaccination, their ability to resist reinfection would be impaired relative to that of unvaccinated individuals. This is a highly undesirable effect for a vaccine, and it is therefore important to avoid such a situation. Previous studies have shown that prior immunization targeted to one immunodominant epitope can skew the immunodominance hierarchy upon subsequent viral infection (22, 23). However, it is not known whether such altered immune responses in effect increase the susceptibility to a secondary infection with viral variants. The present study was undertaken to investigate the reality of these potential risks associated with single-epitope vaccination, with focus on the possible impact on selection of viral variants as well as the susceptibility to challenge with such variants. As model system, the murine LCMV infection was used. LCMV is a natural mouse pathogen, and like many other RNA viruses, it easily mutates (20, 24). Depending on the strain and the infectious dose used, LCMV can cause either acute transient infection with establishment of solid memory CD8⫹ T cell responses (25, 26) or persistent/chronic infection with deletion or transient anergy of relevant CD8⫹ T cells (27, 28). Monospecific, CD8⫹ memory T cells were induced by use of DNA vaccines encoding an immunodominant MHC class I-restricted LCMV epitope covalently linked to human 2-microglobulin (h2m). Such constructs have previously been shown to efficiently induce long-lived antiviral CD8⫹ T cell responses that protect against acute systemic LCMV infection (29). 6285 6286 steel mesh. The cells were washed twice with HBSS, and the cell concentration was adjusted in RPMI 1640 containing 10% FCS supplemented with 2-ME, L-glutamine, and penicillin-streptomycin solution. mAb for flow cytometry The following mAbs were all purchased from BD Pharmingen (San Diego, CA) as rat anti-mouse Abs: FITC-conjugated anti-CD49d (common ␣-chain of LPAM-1 and VLA-4), CyChrome or PerCP-conjugated antiCD8a (Ly2), PE- or FITC-conjugated anti IFN-␥, and PE-conjugated anti-TNF-␣. Flow cytometric analysis Statistical analysis Because there was no evidence that our data were normally distributed, the nonparametric Mann-Whitney U rank-sum test was used to perform comparisons between groups. A value of p ⬍ 0.05 was considered statistically significant. Results Single-epitope DNA vaccination protects against chronic infection in H-2b mice The protective capacity of a monospecific, CD8⫹ T cell memory response induced by DNA vaccination was first investigated in H-2b (C57BL/6) mice. Constructs encoding either of the two H-2brestricted immunodominant LCMV epitopes, gp33– 41 and NP396 – 404 (35, 36), was used (gp33 vaccine and NP396 vaccine, respectively; Fig. 1). Mice were immunized twice, 4 wk apart, and then allowed to rest for 3 wk. Mice similarly immunized with empty vector and/or left untreated (unvaccinated mice) served as controls; in no case where both groups of controls were analyzed in parallel did we observe any differences between vector-vaccinated and unvaccinated mice. All mice were infected with 106 PFU of the rapidly replicating LCMV clone 13, and virus titers in spleen and lungs were followed for a period of 3 mo. As shown in Fig. 2, virus persisted for ⬎20 days in control mice, and in some cases virus was still detectable 2–3 mo after infection. In contrast, DNAvaccinated mice completely controlled the infection within 10 –20 days, and no reemergence of virus was detected for up to 3 mo postinfection (p.i.), indicating that selection for escape mutants does not constitute a relevant problem. Single-epitope DNA-vaccinated H-2b mice are rescued from CD8⫹ T cell failure during high dose infection with LCMV clone 13 The protection pattern shown in Fig. 2 primarily reflects CD8⫹ T cell-mediated virus control (30). Consequently, to understand the underlying mechanism, the CD8⫹ T cell effector response in single-epitope-vaccinated and control mice was analyzed by intracellular cytokine staining for IFN-␥ in the early (days 10 and 20 p.i.) as well as in the late (2–3 mo p.i.) phase of infection. To evaluate the breadth of the antiviral CD8⫹ T cell response, the responses to both immunodominant and one subdominant epitope (gp276 –286) were analyzed; together these populations make up the majority of LCMV-specific, CD8⫹ T cells in H-2b mice (35, 36). Infection of naive mice with high doses of rapidly replicating virus, such as LCMV clone 13, normally results in chronic infection due to failure of the immune response (28, 30, 31). At the CD8⫹ T cell level, this is characterized by either deletion of relevant cells or transient loss of effector function (28). Compared with control mice, single-epitope DNA-vaccinated mice generated substantially more LCMV-specific, CD8⫹ T cells in the early phase of the infection (Fig. 3). As expected, most of the LCMV-specific cells in these mice were directed toward the vaccine epitope. However, a tendency toward increased frequencies of cells specific for the other immunodominant as well as the subdominant gp276 –286 epitope was also observed; this was most clearly seen in NP396-vaccinated mice and primarily in the early phase of infection. Moreover, the absolute number of CD8⫹ T cells was 4 –5 times higher in single-epitope-vaccinated mice compared with control mice (not shown). Two or 3 mo after infection, FIGURE 2. Kinetics of organ virus titers in DNA-vaccinated, vector-vaccinated, or unvaccinated C57BL/6 mice after infection with LCMV clone 13. Mice were immunized twice 4 wk apart with gp33 vaccine (GP33), NP396 vaccine (NP396) or empty vector as a control vaccine (vector). In some experiments unvaccinated mice (unvacc) served as controls instead. Three weeks later, all mice were infected with 106 PFU of LCMV clone 13 i.v. Points represents individual mice. ⴱ, p ⬍ 0.05 relative to unvaccinated/vector-vaccinated mice (by Mann-Whitney rank-sum test). Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 For visualization of LCMV-specific (IFN-␥-producing) CD8⫹ T cells, 2 ⫻ 106 splenocytes were incubated for 5 h at 37°C in 0.2 ml of complete RPMI 1640 medium supplemented with 10 U of murine reIL-2 (R&D Systems Europe, Abingdon, U.K.), 3 M monensin (Sigma-Aldrich, St. Louis, MO), and 0.1 g/ml of the relevant peptide. The following peptides were used: MHC class I (H-2Db) restricted gp33– 41, NP396 – 404, or gp276 – 286 (35, 36); MHC class I (H-2Ld)-restricted NP118 –126 (37, 38); and MHC class I (H-2Kd)-restricted gp283–291 (6); cells cultured without peptide served as background controls. After incubation, cells were surfacestained, washed, permeabilized, and stained with IFN-␥-specific mAb as described previously (39, 40). In some experiments, cells were costained for IFN-␥ and TNF-␣ intracellularly. The frequency of IFN-␥⫹CD8⫹ T cells in unstimulated cultures was ⬍0.3%. Cells were analyzed using a FACSCalibur (BD Biosciences, San Jose, CA), and at least 104 cells were gated using a combination of low angle and side scatter to exclude dead cells and debris. Data analysis was conducted using CellQuest software (BD Biosciences). SINGLE EPITOPE VACCINATION AND IMMUNOSELECTION The Journal of Immunology 6287 LCMV-specific, IFN-␥-producing cells had decreased in singleepitope-vaccinated mice, whereas a distinct population of gp33– 41- and gp276 –286-specific cells had re-emerged in control mice (correlating with virus control in these mice; Fig. 2). Consequently, the quantitative differences between single-epitope-vaccinated mice and control mice regarding the T cell response toward nonimmunizing epitopes were less striking at this time point. However, although the frequencies of CD8⫹ T cells specific for these other epitopes were not always significantly increased in singleepitope-vaccinated mice compared with controls, there was a clear difference in the quality of the cells. Thus, in the early phase of infection, all LCMV-specific cells in control mice were impaired in their capacity to produce IFN-␥, as evidenced by relatively low mean fluorescence intensities (Fig. 4, A and B). In contrast, in DNA-vaccinated mice, not only cells specific for the vaccine epitope, but also cells specific for the other epitopes, produced high amounts of IFN-␥. In the late phase of infection, similar qualitative differences were observed between single-epitope-vaccinated mice and controls (Fig. 4, C and D). Even though gp33– 41-specific, CD8⫹ T cells in particular had re-emerged in control mice, the ability of these cells to produce IFN-␥ was still significantly impaired compared with that in their counterparts in single-epitope DNA-vaccinated mice. This difference in quality was particularly evident if we investigated the ability of LCMV-specific, CD8⫹ T cells to coproduce IFN-␥ and TNF-␣ in NP396 DNA-vaccinated mice and control mice 3 mo p.i. TNF-␣ production has previously been shown to be a good marker for fully functional memory CD8⫹ T cells (41, 42). As shown in Fig. 4D, a higher proportion of the IFN-␥-producing cells in NP396-vaccinated mice coproduced TNF-␣ compared with cells with similar specificity present in mice that had not been vaccinated before infection ( p ⬍ 0.02 for all epitopes). Thus, even at a time point when the infection is controlled in most mice, epitope-vaccinated as well as controls (Fig. 2), CD8⫹ memory T cells specific for the immunizing as well as other LCMV epitopes were functionally more differentiated in mice vaccinated with single-epitope constructs before viral challenge. Single-epitope DNA vaccination protects against chronic infection and favors epitope spreading in H-2d mice The above results suggest that rapid virus clearance combined with the broad CD8⫹ T cell response generated in single-epitope DNAvaccinated H-2b mice protects against viral recrudescence. It is possible, therefore, that the existence of several immunodominant epitopes may be a requirement for the DNA vaccine to be protective. To test this hypothesis, the outcome of high dose LCMV clone 13 infection in single-epitope DNA-vaccinated H-2d mice (BALB/c) was investigated. The antiviral CD8⫹ T cell response in these mice is focused toward a single immunodominant epitope, the Ld-restricted NP118 –126, and failure to generate an immune response to this epitope is known to severely impair virus control (43). In addition, a subdominant Kd-restricted epitope, gp283–291 exists, which during a transient immunizing infection only gives rise to a very small population of Ag-specific, CD8⫹ T cells (44). BALB/c mice were immunized with a vaccine encoding the NP118 –126 epitope covalently linked to h2m (NP118 vaccine) and were subsequently infected i.v. with 106 PFU of LCMV clone 13. As shown in Fig. 5, the protection pattern in single-epitopevaccinated and control (empty vector-vaccinated) BALB/c mice infected with LCMV clone 13 resembled that seen for similarly treated C57BL/6 mice (see Fig. 2 for comparison). In control mice, virus persisted for ⬎20 days and was in some cases still detectable 2–3 mo after infection. By contrast, NP118-vaccinated mice completely controlled the infection, and no re-emergence of virus was detected for up to 6 mo p.i. Again, this indicates that selection of viral escape mutants does not constitute a relevant problem. The single-epitope-vaccinated, virus-infected BALB/c mice generated substantially more LCMV-specific, CD8⫹ T cells compared with control mice (Fig. 6). Thus, high frequencies of vaccine-specific, CD8⫹ T cells could be detected in the early (days 10 and 20 p.i.) as well as the late (2–3 mo p.i.) phases of infection. Surprisingly, however, the CD8⫹ T cell response in DNA-vaccinated BALB/c mice was not particularly focused toward the vaccine epitope. Thus, between days 10 and 20 p.i., a high frequency of CD8⫹ T cells specific for the subdominant gp283–291 epitope Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 FIGURE 3. Kinetics of Ag-specific, CD8⫹ T cells in DNA-vaccinated, vector-vaccinated, and unvaccinated C57BL/6 mice after i.v. challenge with LCMV clone 13. Mice were immunized twice 4 wk apart with gp33 vaccine (GP33), NP396 vaccine (NP396), or empty vector (vector) for the control; in some experiments unvaccinated mice (unvacc) served as controls. Three weeks later, mice were infected with 106 PFU of LCMV clone 13 i.v. On the days indicated, splenocytes were stimulated in vitro with MHC class I-restricted LCMV peptides (gp33– 41, NP396 – 404, or gp276 –286) for 5 h, surface-stained, permeabilized, and stained for IFN-␥. Splenocytes incubated without peptide served as negative controls; VLA4⫹,IFN-␥⫹CD8⫹ T cells were in this case ⬍0.3%. The mean and SD are depicted. ⴱ, p ⬍ 0.05 relative to unvaccinated/vector-vaccinated mice (by Mann-Whitney rank-sum test). SINGLE EPITOPE VACCINATION AND IMMUNOSELECTION 6288 Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 FIGURE 4. The Journal of Immunology 6289 FIGURE 5. Kinetics of organ virus titers in DNA-vaccinated and vector-vaccinated BALB/c mice after infection with LCMV clone 13. Mice were immunized twice 4 wk apart with NP118 vaccine (NP118) or empty vector as control vaccine (vector). Three weeks later, all mice were infected with 106 PFU of LCMV clone 13 i.v. Points represents individual mice. ⴱ, p ⬍ 0.05 relative to vector-vaccinated mice (by Mann-Whitney rank-sum test). Single-epitope DNA vaccination protects against reinfections with viral variants The above results suggest that although the immune response in single-epitope-vaccinated mice is focused toward the immunizing epitope, a sufficiently broad T cell response is generated upon viral challenge to prevent viral recrudescence due to escape variants. However, there are two uncertainties regarding this interpretation. First, we do not know whether escape variants are generated in sufficient numbers to present a real challenge, e.g., Abs might suffice to control viral variants mutated in the vaccine epitope (45, 46). Secondly, it could be argued that viral challenge with a high dose of rapidly replicating virus allows the generation of a broader T cell response than would be seen under more normal circumstances; certainly, it could be argued that the immune response in clone 13-infected mice represents a special case. To address these issues, a second type of experimental set-up was used. Single-epitope (gp33 or NP396)-vaccinated mice were infected i.v. with 104 PFU of the slowly replicating LCMV Arm and allowed to rest for 3 mo to become LCMV immune. For comparison, vector-vaccinated mice were similarly infected; these mice were included to represent LCMV-immune mice with an unperturbed CD8⫹ T cell memory response. To evaluate the quality of the LCMV-specific immune response generated under these conditions, some mice were challenged i.c. with a relatively high dose (200 PFU ⬇ 1000 LD50 in naive mice) of virus, representing an escape variant in the epitope used for DNA vaccination. Mortality was registered for a period of 14 days after challenge. The outcome of infection by the i.c. route is determined by a race between the virus and the CD8⫹ T cell-mediated immune response: naive mice die from the extensive cell damage induced in the attempt to clear an infection that is already too extensive, whereas fully immune mice completely resist this challenge due to rapid CD8⫹ T cell-mediated virus clearance and minimal cell damage (47– 49). Thus, the outcome of i.c. infection with known loss variants would immediately indicate whether a biologically relevant level of CD8⫹ T cell-mediated protection against potential escape variants could be generated during viral infection of singleepitope-vaccinated mice. A summary of the results is listed in Table I. As expected, all LCMV-immune control mice were solidly immune and resisted i.c. challenge with either gp33-nil or NP396nil. Also nine of 10 DNA-vaccinated mice survived i.c. infection with a viral variant mutated in the vaccine epitope. In parallel experiments we studied the composition of the antiviral CD8⫹ T cell response in the mice before secondary challenge with escape virus. This analysis provides detailed information regarding the epitope hierarchy needed to interpret the outcome of the challenge experiment. Single-epitope- and vector-vaccinated mice were infected with 104 PFU of LCMV Arm i.v. as described above, and CD8⫹ T cell responses were analyzed 8 and 90 days p.i. (Fig. 7A). Although frequencies of LCMV-specific, CD8⫹ T cells were lower on day 90 p.i. compared with day 8 p.i., the overall picture was similar, consistent with the idea that the memory response reflects initial clonal burst size (50). Thus, as expected, higher frequencies of CD8⫹ T cells specific for the vaccine epitope were found in vaccinated mice 8 days and 3 mo p.i. (Fig. 7B). However, single-epitope DNA-vaccinated mice did not appear to have a dramatically impaired response to the other epitopes in either the early or the late phase of FIGURE 4. Representative plots of Ag-specific, CD8⫹ T cells from gp33-vaccinated, NP396-vaccinated, or vector-vaccinated C57BL/6 mice on day 10 p.i. (A), 20 p.i. (B), or 2–3 mo p.i. (C and D). Mice were immunized twice, 4 wk apart, with gp33 vaccine (GP33), NP396 vaccine (NP396), or empty vector (vector) for the controls; in some experiments, unvaccinated mice served as controls (not shown). Three weeks later, mice were infected with 106 PFU of LCMV clone 13 i.v. On the days indicated, splenocytes were stimulated in vitro with MHC class I-restricted LCMV peptides (gp33– 41, NP396 – 404, or gp276 –286) for 5 h, surface-stained for CD8 and the activation marker VLA-4, permeabilized, and stained for IFN-␥; (A–C). D, NP396-vaccinated and control vaccinated mice were surface-stained for CD8, permeabilized, and costained for IFN-␥ and TNF-␣ intracellularly. Splenocytes incubated without peptide served as negative controls; VLA-4⫹,IFN-␥⫹CD8⫹ T cells or TNF-␣⫹,IFN-␥⫹CD8⫹ T cells were in this case ⬍0.3%. Gated CD8⫹ T cells from a representative mouse are shown. Numbers in parentheses refer to mean fluorescence intensities (MFI). Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 appeared, and this cell population persisted throughout a 6-mo observation period (Fig. 6 and data not shown). In contrast, gp283– 291-specific, CD8⫹ T cells could only be detected at very low levels in control mice throughout the 6-mo detection period (Fig. 6 and data not shown). Notably, very few gp283–291-specific, CD8⫹ T cells were generated during acute transient infection of unvaccinated mice (Fig. 6, inset) (44). Thus, single-epitope DNA vaccination not only elicits a strong vaccine-specific antiviral CD8⫹ T cell response, but also favors epitope spreading during protracted LCMV infection. 6290 SINGLE EPITOPE VACCINATION AND IMMUNOSELECTION infection. Thus, the frequency of CD8⫹ T cells specific for the other immunodominant epitopes was only marginally lower in singleepitope-vaccinated mice compared with that in mice with an unperturbed LCMV-specific memory repertoire. A reduced response to the subdominant gp276 –286 epitope was observed, but this was a consistent finding only in gp33-vaccinated mice. Single-epitope DNA vaccination thus does not substantially reduce the breadth of the normal antiviral immune response, which readily explains why singleepitope-vaccinated, LCMV-immune mice were well protected against i.c. challenge with CD8⫹ T cell escape variants. Discussion The present study was undertaken to investigate the protective capacity of a monospecific, CD8⫹ T cell memory response induced by single-epitope DNA vaccination, with special focus on the potential risk associated with induction of a narrow immune response. At the level of the individual host, single-epitope DNA vaccination may select for viral CD8⫹ T cell escape variants during chronic conditions due to the narrow immune response induced. Second, prior immunization targeted to a single dominant epitope may dramatically alter the dominance hierarchy upon subsequent viral infection, which may lead to increased susceptibility to reinfections with viral CD8⫹ T cell escape variants circulating in a population. With regard to the first phenomenon, emergence of Table I. Susceptibility to i.c. infection with viral variants of LCMV Arm in DNA vaccinated, LCMV-immune micea Vaccine gp33 Vector Vector NP396 Vector Vector 1° Infection (104 PFU i.v.) 2° Infection (200 PFU i.c.) Mortalityb LCMV Arm 53b LCMV Arm 53b gp33-nil gp33-nil gp33-nil NP396-nil NP396-nil NP396-nil 1/5 0/5 4/4 0/5 0/5 4/4 LCMV Arm 53b LCMV Arm 53b a Mice were vaccinated twice 4 wk apart with gp33 vaccine (gp33), NP396 vaccine, or empty vector (vector) and infected with 104 PFU of LCMV Arm 3 wk later (1° infection); groups of vector-vaccinated mice were left uninfected to represent naive mice. All mice were infected i.c. with 200 PFU of either gp33-nil or NP396-nil 3 mo later (2° infection). b Mortality was registered twice daily for 2 wk after i.c. challenge. viral escape variants has been observed in several viral animal models after epitope vaccination (18, 21, 51). In humans, generation of viral variants is a common phenomenon in HIV patients who have a highly focused CD8⫹ T cell response (13, 14), and viral CD8⫹ T cell escape has also been observed in patients with chronic hepatitis B and C infection (52, 53). In H-2b mice we found that single-epitope vaccination with either gp33 or NP396 vaccine, followed by high dose infection with the vicerotropic LCMV clone 13 led to rapid and sustained virus control, and this correlated with a strong and broadly reactive CD8⫹ T cell response. Even though the response was dominated by CD8⫹ T cells specific for the epitope encoded by the DNA vaccine, fully functional cells specific for the other immunodominant as well as at least one subdominant LCMV class I-restricted epitope were also generated in significant numbers. In contrast, control mice suffered a chronic infection, reflecting loss of CD8⫹ T cell effector function evidenced by impaired or absent IFN-␥ production. Notably, NP396 – 404-specific, CD8⫹ T cells lost effector functions earlier and more completely than did gp33– 41 and gp276 –286-specific cells in control mice during chronic conditions. This has been observed previously (28, 54) and probably explains why it was more difficult to maintain the NP396 – 404specific T cell population in gp33-vaccinated mice than to maintain the gp33– 41-specific T cell population in NP396-vaccinated mice. The generation of a broad CD8⫹ T cell response combined with the rapid virus clearance may explain why we did not observe viral recrudescence in DNA-vaccinated, LCMV-infected mice. Thus, the virus has a limited time to mutate, and any potential viral variants that might be generated are rapidly controlled by the small, but significant, CD8⫹ T cell responses to the other LCMV epitopes. We were, in that respect, surprised to find that even NP118 DNA-vaccinated BALB/c mice completely controlled a high dose LCMV clone 13 infection with no indication of biologically relevant viral escape. Thus, in H-2d mice only one immunodominant and one extremely subdominant CD8⫹ T cell LCMV epitope is known, and selection of viral escape variants has previously been observed in NP118 –126 peptide-immunized mice infected with LCMV (21). Flow cytometric analysis revealed that the antiviral CD8⫹ T cell response in our single-epitope DNA-vaccinated Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 FIGURE 6. Kinetics of Ag-specific, CD8⫹ T cells in DNA-vaccinated and vector-vaccinated BALB/c mice after i.v. challenge with LCMV clone 13. Mice were immunized twice, 4 wk apart, with NP118 vaccine (NP118) or empty vector (vector) for the controls. Three weeks later, mice were infected with 106 PFU of LCMV clone 13 i.v. On the days indicated, splenocytes were stimulated in vitro with MHC class I-restricted LCMV peptides (NP118 –126 or gp283–291) for 5 h, surface-stained, permeabilized, and stained for IFN-␥. Splenocytes incubated without peptide served as negative controls; IFN␥⫹,CD8⫹ T cells were in this case ⬍0.3%. The mean and SD are depicted. ⴱ, p ⬍ 0.05 relative to vector-vaccinated mice (by Mann-Whitney rank-sum test). Inset represents data from mice infected 40 days earlier with optimal immunizing doses of LCMV Arm or clone 13 (104 PFU). The Journal of Immunology 6291 BALB/c mice was not particularly focused toward the vaccine epitope as we would have expected. Thus, a substantial population of CD8⫹ T cells specific for the subdominant gp283–291 epitope was likewise present in mice analyzed ⬎20 days p.i. Notably, gp283–291-specific, CD8⫹ T cells constituted ⬍1% of the CD8⫹ T cell population in control vaccinated counterparts as well as in LCMV-immune mice that had undergone a transient immunizing infection 40 days previously. Thus, single-epitope DNA vaccination not only protects against high dose infection and antiviral CD8⫹ T cell failure, but also favors epitope spreading in H-2d mice. The above findings are consistent with the hypothesis that the generation of a potent and broadly reactive CD8⫹ T cell population is decided by a delicate balance between the initial CD8⫹ T cell response and the viral load (42, 54). In control mice infected with LCMV clone 13 (H-2b and H-2d), the absence of primed LCMV-specific, CD8⫹ T cells results in a high viral load, which Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 FIGURE 7. Kinetics of the frequency of Ag-specific, CD8⫹ T cells with an activated phenotype (VLA4highIFN-␥⫹) in DNA-vaccinated and vector-vaccinated C57BL/6 mice after i.v. challenge with LCMV Arm. Mice were vaccinated twice 4 wk apart with gp33 vaccine (gp33), NP396 vaccine (NP396), or empty vector (vector) for the controls. Three weeks later, mice were infected with 104 PFU of LCMV Arm i.v. On the indicated days, splenocytes were incubated with gp33– 41, NP396 – 404, or gp276 –286 peptide for 5 h. After incubation, cells were surface-stained, permeabilized, and stained for IFN-␥. Splenocytes incubated without peptide served as negative controls; VLA-4⫹,IFN␥⫹,CD8⫹ T cells were in this case ⬍0.3%. A, Frequency of peptide-specific cells as a function of immunization status (mean ⫾ SD). ⴱ, p ⬍ 0.05 relative to vectorvaccinated mice (by Mann-Whitney rank-sum test). B, Representative plots of gated CD8⫹ cells from a gp33vaccinated, a NP396-vaccinated, and a vector-vaccinated mouse. subsequently leads to a progressive failure of the antiviral CD8⫹ T cell response. In vaccinated mice, the initial monospecific, CD8⫹ T cell response controls the high dose infection sufficiently fast to prevent exhaustion of the CD8⫹ T cell response. The virus, however, is kept in the host long enough to prime for a broad antiviral CD8⫹ T cell response. Thus, in H-2b mice, a response to both the other dominant and at least one subdominant epitope is efficiently induced. In vaccinated H-2d mice, virus is also cleared fast enough to prevent general CD8⫹ T cell failure. This is obviously associated with a strong NP118 –126-specific response. However, because the virus infection is not immediately controlled, extensive expansion of CD8⫹ T cells specific for the subdominant gp283– 291 epitope is also observed. This is in contrast to the situation in unvaccinated H-2d mice infected with a moderate dose of LCMV, where rapid virus clearance mediated by NP118 –126-specific, CD8⫹ T cells apparently prevents efficient expansion of gp283– 291-specific, CD8⫹ T cells. Thus, the conditions for induction of 6292 Previous studies have shown that prior immunization targeted to one immunodominant epitope can dramatically alter the immunodominance hierarchy upon subsequent viral infection (22, 23). The reason that we get a different result may again be that our vaccine induces relatively moderate expansion of vaccine-specific cells (29). Overall, this suggests that vaccines that elicit weak/moderate responses might actually be of benefit in some circumstances. In conclusion, our study shows that single-epitope DNA vaccination has the ability to protect against potential escape mutants regardless of whether these mutants arise inside the host or are presented during subsequent challenge. A likely limiting factor for this vaccine strategy to work, however, is the mutation rate of the virus, which may explain why monospecific, CD8⫹ T cell responses do not protect against rapidly mutating HIV infections (13, 14). Acknowledgments We thank Lone Malte and Grethe Thørner Andersen for expert technical assistance. References 1. Gammon, G., N. Shastri, J. Cogswell, S. Wilbur, S. Sadegh-Nasseri, U. Krzych, A. Miller, and E. Sercarz. 1987. The choice of T-cell epitopes utilized on a protein antigen depends on multiple factors distant from, as well as at the determinant site. Immunol. Rev. 98:53. 2. Sercarz, E. E., P. V. Lehmann, A. Ametani, G. Benichou, A. Miller, and K. Moudgil. 1993. Dominance and crypticity of T cell antigenic determinants. Annu. Rev. Immunol. 11:729. 3. Kojima, M., K. B. Cease, G. K. Buckenmeyer, and J. A. Berzofsky. 1988. Limiting dilution comparison of the repertoires of high and low responder MHCrestricted T cells. J. Exp. Med. 167:1100. 4. Bennink, J. R., and J. W. Yewdell. 1988. Murine cytotoxic T lymphocyte recognition of individual influenza virus proteins: high frequency of nonresponder MHC class I alleles. J. Exp. Med. 168:1935. 5. Frelinger, J. A., A. Orn, P. R. Brayton, and L. Hood. 1983. Use of cloned H-2 genes for study of H-2-restricted cytotoxicity: Ld is the LCMV restriction element for H-2d. Transplant. Proc. 15:2024. 6. van der Most, R. G., R. J. Concepcion, C. Oseroff, J. Alexander, S. Southwood, J. Sidney, R. W. Chesnut, R. Ahmed, and A. Sette. 1997. Uncovering subdominant cytotoxic T-lymphocyte responses in lymphocytic choriomeningitis virusinfected BALB/c mice. J. Virol. 71:5110. 7. Mylin, L. M., T. D. Schell, D. Roberts, M. Epler, A. Boesteanu, E. J. Collins, J. A. Frelinger, S. Joyce, and S. S. Tevethia. 2000. Quantitation of CD8⫹ Tlymphocyte responses to multiple epitopes from simian virus 40 (SV40) large T antigen in C57BL/6 mice immunized with SV40, SV40 T-antigen-transformed cells, or vaccinia virus recombinants expressing full-length T antigen or epitope minigenes. J. Virol. 74:6922. 8. Allan, J. E., and P. C. Doherty. 1985. Consequences of a single Ir-gene defect for the pathogenesis of lymphocytic choriomeningitis. Immunogenetics 21:581. 9. Thomsen, A. R., and O. Marker. 1989. Class I gene regulation of haplotype preference may influence antiviral immunity in vivo. Cell. Immunol. 122:365. 10. Ulmer, J. B., J. C. Sadoff, and M. A. Liu. 1996. DNA vaccines. Curr. Opin. Immunol. 8:531. 11. Borrow, P., and G. M. Shaw. 1998. Cytotoxic T-lymphocyte escape viral variants: how important are they in viral evasion of immune clearance in vivo? Immunol. Rev. 164:37. 12. Holland, J. J., J. C. de la Torre, and D. A. Steinhauer. 1992. RNA virus populations as quasispecies. Curr. Top. Microbiol. Immunol. 176:1. 13. Borrow, P., H. Lewicki, X. Wei, M. S. Horwitz, N. Peffer, H. Meyers, J. A. Nelson, J. E. Gairin, B. H. Hahn, M. B. Oldstone, et al. 1997. Antiviral pressure exerted by HIV-1-specific cytotoxic T lymphocytes (CTLs) during primary infection demonstrated by rapid selection of CTL escape virus. Nat. Med. 3:205. 14. Phillips, R. E., S. Rowland-Jones, D. F. Nixon, F. M. Gotch, J. P. Edwards, A. O. Ogunlesi, J. G. Elvin, J. A. Rothbard, C. R. Bangham, and C. R. Rizza. 1991. Human immunodeficiency virus genetic variation that can escape cytotoxic T cell recognition. Nature 354:453. 15. Webster, R. G., W. J. Bean, O. T. Gorman, T. M. Chambers, and Y. Kawaoka. 1992. Evolution and ecology of influenza A viruses. Microbiol. Rev. 56:152. 16. Gorman, O. T., W. J. Bean, and R. G. Webster. 1992. Evolutionary processes in influenza viruses: divergence, rapid evolution, and stasis. Curr. Top. Microbiol. Immunol. 176:75. 17. O’Connor, D. H., T. M. Allen, and D. I. Watkins. 2002. Cytotoxic T-lymphocyte escape monitoring in simian immunodeficiency virus vaccine challenge studies. DNA Cell Biol. 21:659. 18. Mortara, L., F. Letourneur, H. Gras-Masse, A. Venet, J. G. Guillet, and I. Bourgault-Villada. 1998. Selection of virus variants and emergence of virus escape mutants after immunization with an epitope vaccine. J. Virol. 72:1403. 19. Pircher, H., D. Moskophidis, U. Rohrer, K. Burki, H. Hengartner, and R. M. Zinkernagel. 1990. Viral escape by selection of cytotoxic T cell-resistant virus variants in vivo. Nature 346:629. Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 a strong gp283–291-specific, CD8⫹ T cell response appears to be very stringent: a sustained, high virus load exhausts this population, and rapid virus clearance prevents its expansion. Our findings are somewhat at odds with those of other studies in different viral models. Several studies have shown that singleepitope vaccination selects for viral escape mutants during chronic infection (17, 18, 21, 55, 56). These conflicting findings may in part be attributed to differences in mutation rates and escape mechanisms between viruses. Thus, for genetically stable viruses such as DNA viruses, our results clearly indicate that vaccination with a single epitope will suffice for induction of protective immunity. However, because LCMV is an RNA virus expressing considerable genetic instability (57), our results could suggest that singleepitope vaccination may also be a feasible approach for this category of viruses. In the latter case, viral escape strategies may play an important role, and clearly, virus-induced impairment of the immune system (e.g., HIV) will reduce the flexibility of the immune system and thus increase the likelihood that viral variants will escape immune-mediated control. Time is also a critical factor; the longer the virus stays in the host, the higher the chance for a relevant mutation. Because virus was cleared relatively fast in our model, fewer escape mutants may have had time to develop. Additionally, as indicated above, the strength of the vaccineprimed immune response seems to be a determining factor for the rate of virus clearance, which influences the composition of the antiviral CD8⫹ T cell response that finally determines the outcome of infection. Our DNA vaccine seems to prime for a moderate response (between 0.05 and 0.12% vaccine-specific, CD8⫹ T cells) (29), which upon viral infection results in a delicate balance, leading to viral control as well as generation of a broad antiviral CD8⫹ T cell response. This report also addresses a second issue, which needs to be considered before introducing single-epitope vaccination. The focused selection pressure toward a single epitope is likely to favor that escape variants, at some point, will become more prevalent, which will lead to vaccine failure. However, this is acceptable as long as it is not a frequently occurring event. Much more damaging is the possibility that vaccination might impair naturally induced immunity. This could happen if single-epitope vaccination leads to a skewed immunodominance hierarchy in hosts undergoing normal immunizing infection. Few studies have looked at the functional effects of this. However, findings from the LCMV as well as the influenza models suggest, that pre-existence of a monospecific, CD8⫹ T cell response may affect the CD8⫹ T cell memory response generated to subsequent viral infections by suppressing CD8⫹ T cell responses to other immunogenic viral epitopes (22, 23). DNA vaccination followed by wild-type LCMV infection protected almost all mice against i.c. reinfection with high doses of escape variants. Thus, five of five NP396-vaccinated and four of five gp33-vaccinated mice survived a relatively high i.c. challenge dose, as did all LCMV-immune control mice. Notably, all surviving mice had cleared virus from the brain and had detectable LCMV-specific, CD8⫹ T cell responses in the spleen (data not shown). This eliminates the possibility that mice survived as asymptomatic virus carriers; a phenomenon seen in T cell-deficient or anergic mice (58). Taken together it indicates that mice in both groups have had a broad antiviral CD8⫹ T cell response before i.c. challenge. This was indeed confirmed by analyzing the composition of the antiviral CD8⫹ T cell response. Thus, DNA-vaccinated mice only developed a slightly skewed CD8⫹ T cell response to wild-type LCMV infection in the acute as well as the memory phase of infection; only responses to the subdominant epitope were suppressed, and this was consistent only in gp33-vaccinated mice. SINGLE EPITOPE VACCINATION AND IMMUNOSELECTION The Journal of Immunology 40. Andreasen, S. O., J. P. Christensen, O. Marker, and A. R. Thomsen. 1999. Virusinduced non-specific signals cause cell cycle progression of primed CD8⫹ T cells but do not induce cell differentiation. Int. Immunol. 11:1463. 41. Slifka, M. K., and J. L. Whitton. 2000. Activated and memory CD8⫹ T cells can be distinguished by their cytokine profiles and phenotypic markers. J. Immunol. 164:208. 42. Wherry, E. J., and R. Ahmed. 2004. Memory CD8 T-cell differentiation during viral infection. J. Virol. 78:5535. 43. Thomsen, A. R., and O. Marker. 1989. MHC and non-MHC genes regulate elimination of lymphocytic choriomeningitis virus and antiviral cytotoxic T lymphocyte and delayed-type hypersensitivity mediating T lymphocyte activity in parallel. J. Immunol. 142:1333. 44. De Boer, R. J., M. Oprea, R. Antia, K. Murali-Krishna, R. Ahmed, and A. S. Perelson. 2001. Recruitment times, proliferation, and apoptosis rates during the CD8⫹ T-cell response to lymphocytic choriomeningitis virus. J. Virol. 75:10663. 45. Thomsen, A. R., J. Johansen, O. Marker, and J. P. Christensen. 1996. Exhaustion of CTL memory and recrudescence of viremia in lymphocytic choriomeningitis virus-infected MHC class II-deficient mice and B cell-deficient mice. J. Immunol. 157:3074. 46. Ciurea, A., P. Klenerman, L. Hunziker, E. Horvath, B. M. Senn, A. F. Ochsenbein, H. Hengartner, and R. M. Zinkernagel. 2000. Viral persistence in vivo through selection of neutralizing antibody-escape variants. Proc. Natl. Acad. Sci. USA 97:2749. 47. Doherty, P. C., J. E. Allan, F. Lynch, and R. Ceredig. 1990. Dissection of an inflammatory process induced by CD8⫹ T cells. Immunol. Today 11:55. 48. Hany, M., S. Oehen, M. Schulz, H. Hengartner, M. Mackett, D. H. Bishop, H. Overton, and R. M. Zinkernagel. 1989. Anti-viral protection and prevention of lymphocytic choriomeningitis or of the local footpad swelling reaction in mice by immunization with vaccinia-recombinant virus expressing LCMV-WE nucleoprotein or glycoprotein. Eur. J. Immunol. 19:417. 49. Thomsen, A. R., M. Volkert, and O. Marker. 1979. The timing of the immune response in relation to virus growth determines the outcome of the LCM infection. Acta Pathol. Microbiol. Scand. Suppl. 87C:47. 50. Hou, S., L. Hyland, K. W. Ryan, A. Portner, and P. C. Doherty. 1994. Virusspecific CD8⫹ T-cell memory determined by clonal burst size. Nature 369:652. 51. Yasutomi, Y., S. Koenig, R. M. Woods, J. Madsen, N. M. Wassef, C. R. Alving, H. J. Klein, T. E. Nolan, L. J. Boots, and J. A. Kessler. 1995. A vaccine-elicited, single viral epitope-specific cytotoxic T lymphocyte response does not protect against intravenous, cell-free simian immunodeficiency virus challenge. J. Virol. 69:2279. 52. Maini, M. K., S. Reignat, C. Boni, G. S. Ogg, A. S. King, F. Malacarne, G. J. Webster, and A. Bertoletti. 2000. T cell receptor usage of virus-specific CD8 cells and recognition of viral mutations during acute and persistent hepatitis B virus infection. Eur. J. Immunol. 30:3067. 53. Erickson, A. L., Y. Kimura, S. Igarashi, J. Eichelberger, M. Houghton, J. Sidney, D. McKinney, A. Sette, A. L. Hughes, and C. M. Walker. 2001. The outcome of hepatitis C virus infection is predicted by escape mutations in epitopes targeted by cytotoxic T lymphocytes. Immunity 15:883. 54. Fuller, M. J., A. Khanolkar, A. E. Tebo, and A. J. Zajac. 2004. Maintenance, loss, and resurgence of T cell responses during acute, protracted, and chronic viral infections. J. Immunol. 172:4204. 55. Price, G. E., R. Ou, H. Jiang, L. Huang, and D. Moskophidis. 2000. Viral escape by selection of cytotoxic T cell-resistant variants in influenza A virus pneumonia. J. Exp. Med. 191:1853. 56. Chua, M. M., K. C. MacNamara, L. San Mateo, H. Shen, and S. R. Weiss. 2004. Effects of an epitope-specific CD8⫹ T-cell response on murine coronavirus central nervous system disease: protection from virus replication and antigen spread and selection of epitope escape mutants. J. Virol. 78:1150. 57. Sevilla, N., E. Domingo, and J. C. de la Torre. 2002. Contribution of LCMV towards deciphering biology of quasispecies in vivo. Curr. Top. Microbiol. Immunol. 263:197. 58. Andersen, I. H., O. Marker, and A. R. Thomsen. 1991. Breakdown of blood-brain barrier function in the murine lymphocytic choriomeningitis virus infection mediated by virus-specific CD8⫹ T cells. J. Neuroimmunol. 31:155. Downloaded from http://www.jimmunol.org/ by guest on June 18, 2017 20. Aebischer, T., D. Moskophidis, U. H. Rohrer, R. M. Zinkernagel, and H. Hengartner. 1991. In vitro selection of lymphocytic choriomeningitis virus escape mutants by cytotoxic T lymphocytes. Proc. Natl. Acad. Sci. USA 88:11047. 21. Weidt, G., W. Deppert, O. Utermohlen, J. Heukeshoven, and F. Lehmann-Grube. 1995. Emergence of virus escape mutants after immunization with epitope vaccine. J. Virol. 69:7147. 22. Rodriguez, F., S. Harkins, M. K. Slifka, and J. L. Whitton. 2002. Immunodominance in virus-induced CD8⫹ T-cell responses is dramatically modified by DNA immunization and is regulated by ␥ interferon. J. Virol. 76:4251. 23. Chen, W., L. C. Anton, J. R. Bennink, and J. W. Yewdell. 2000. Dissecting the multifactorial causes of immunodominance in class I-restricted T cell responses to viruses. Immunity 12:83. 24. Moskophidis, D., and R. M. Zinkernagel. 1995. Immunobiology of cytotoxic T-cell escape mutants of lymphocytic choriomeningitis virus. J. Virol. 69:2187. 25. Marker, O., and M. Volkert. 1973. Studies on cell-mediated immunity to lymphocytic choriomeningitis virus in mice. J. Exp. Med. 137:1511. 26. Moskophidis, D., S. P. Cobbold, H. Waldmann, and F. Lehmann-Grube. 1987. Mechanism of recovery from acute virus infection: treatment of lymphocytic choriomeningitis virus-infected mice with monoclonal antibodies reveals that Lyt-2⫹ T lymphocytes mediate clearance of virus and regulate the antiviral antibody response. J. Virol. 61:1867. 27. Moskophidis, D., F. Lechner, H. Pircher, and R. M. Zinkernagel. 1993. Virus persistence in acutely infected immunocompetent mice by exhaustion of antiviral cytotoxic effector T cells. Nature 362:758. 28. Zajac, A. J., J. N. Blattman, K. Murali-Krishna, D. J. Sourdive, M. Suresh, J. D. Altman, and R. Ahmed. 1998. Viral immune evasion due to persistence of activated T cells without effector function. J. Exp. Med. 188:2205. 29. Bartholdy, C., A. Stryhn, N. J. Hansen, S. Buus, and A. R. Thomsen. 2003. Incomplete effector/memory differentiation of antigen-primed CD8⫹ T cells in gene gun DNA-vaccinated mice. Eur. J. Immunol. 33:1941. 30. Matloubian, M., R. J. Concepcion, and R. Ahmed. 1994. CD4⫹ T cells are required to sustain CD8⫹ cytotoxic T-cell responses during chronic viral infection. J. Virol. 68:8056. 31. Ahmed, R., A. Salmi, L. D. Butler, J. M. Chiller, and M. B. Oldstone. 1984. Selection of genetic variants of lymphocytic choriomeningitis virus in spleens of persistently infected mice: role in suppression of cytotoxic T lymphocyte response and viral persistence. J. Exp. Med. 160:521. 32. Lewicki, H., A. Tishon, P. Borrow, C. F. Evans, J. E. Gairin, K. M. Hahn, D. A. Jewell, I. A. Wilson, and M. B. Oldstone. 1995. CTL escape viral variants. I. Generation and molecular characterization. Virology 210:29. 33. Lewicki, H. A., M. G. von Herrath, C. F. Evans, J. L. Whitton, and M. B. Oldstone. 1995. CTL escape viral variants. II. Biologic activity in vivo. Virology 211:443. 34. Battegay, M., S. Cooper, A. Althage, J. Banziger, H. Hengartner, and R. M. Zinkernagel. 1991. Quantification of lymphocytic choriomeningitis virus with an immunological focus assay in 24- or 96-well plates. J. Virol. Methods 33:191. 35. Yanagi, Y., A. Tishon, H. Lewicki, B. A. Cubitt, and M. B. Oldstone. 1992. Diversity of T-cell receptors in virus-specific cytotoxic T lymphocytes recognizing three distinct viral epitopes restricted by a single major histocompatibility complex molecule. J. Virol. 66:2527. 36. Gairin, J. E., H. Mazarguil, D. Hudrisier, and M. B. Oldstone. 1995. Optimal lymphocytic choriomeningitis virus sequences restricted by H-2Db major histocompatibility complex class I molecules and presented to cytotoxic T lymphocytes. J. Virol. 69:2297. 37. Schulz, M., P. Aichele, M. Vollenweider, F. W. Bobe, F. Cardinaux, H. Hengartner, and R. M. Zinkernagel. 1989. Major histocompatibility complex– dependent T cell epitopes of lymphocytic choriomeningitis virus nucleoprotein and their protective capacity against viral disease. Eur. J. Immunol. 19:1657. 38. Murali-Krishna, K., J. D. Altman, M. Suresh, D. J. Sourdive, A. J. Zajac, J. D. Miller, J. Slansky, and R. Ahmed. 1998. Counting antigen-specific CD8 T cells: a reevaluation of bystander activation during viral infection. Immunity 8:177. 39. Christensen, J. P., J. P. Stenvang, O. Marker, and A. R. Thomsen. 1996. Characterization of virus-primed CD8⫹ T cells with a type 1 cytokine profile. Int. Immunol. 8:1453. 6293