* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Section 1 Metabolic Processes Cell Structure and Process
Nucleic acid analogue wikipedia , lookup
Peptide synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Citric acid cycle wikipedia , lookup
Photosynthesis wikipedia , lookup
Radical (chemistry) wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Metalloprotein wikipedia , lookup
Proteolysis wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
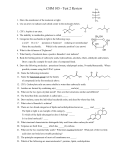
Section 1 Metabolic Processes Cell Structure and Process electronegativity: a substance’s ability to attract electrons usually, electronegativity increases from left to right and from bottom to top in a periodic table if the electronegativity difference of a bond is greater than or equal to 1.7, that bond is ionic if it is less than 1.7, it is polar covalent polarity of a molecule is dependent on the electronegativity of the constituent elements and the molecule’s shape bonding: ionic: usually between metals and nonmetals; metals lose 1 or more electrons to nonmetals; when dissolved in water, they lose their crystalline structure, ex. NaCl covalent: compounds form by sharing electrons between neighbouring atoms. A single † dash in a structural diagram indicates 1 pair electrons being shared, 2 dashes mean a double bond (4 electrons shared), etc. intermolecular forces: bonds between molecules determine the physical state of a substance at a given temperature and pressure weaker than the bonds that hold the molecule together hydrogen bonds are the strongest intermolecular force. They form between an electropositive hydrogen of one polar molecule and the electronegative N, O or F of a neighbouring molecule. These bonds are easily broken and very important biologically, as they determine the shape and thus the function of molecules acids, bases and buffers: acids: substances that increase the concentration of H 3O+ (hydronium) ions when dissolved in water † of OH - (hydroxide) ions when dissolved bases: substances that increase the concentration in water † maintain this, cells use buffers. most cells work best at a pH of approx. 7. To buffers are usually conjugate acid-base pairs in equilibrium and sometimes buffers are proteins the most common buffer (biologically) is carbonic acid and bicarbonate pH<7.35 is acidosis and pH>7.35 is alkalosis. 7.35 is normal. organic compounds: compounds containing carbon organized according to properties and molecular structure functional groups: specific groups of atoms attached to an organic molecule react with other functional groups influence physical and chemical properties organic (carboxylic) acids contain: -COOH polar weak acid † amines (amino-) contain: -NH 2 weak base found in†proteins alcohols contain: -OH polar, forms hydrogen bonds † water soluble aldehydes and ketones (carboxyls) contain: = O if the “ = O ” is on a terminal carbon, the molecule is an aldehyde † after drinking alcohol aldehydes are exhaled † if the “ = O ” is on a mid chain carbon, the molecule is a ketone ketones are expelled in urine if eating a high-fat diet † aldehydes and ketones are water-soluble, common in sugars phosphates contain: found in ADP and ATP sulfhydryls contain: -SH macromolecules † large molecules of repeating subunits four major classes: carbohydrates (CHOs) lipids (made of fatty acids and glycerol) proteins (made of amino acids) nucleic acids (made of nucleotides) condensation reactions aka anabolic reactions or dehydration synthesis reactions form larger molecules a molecule of water is removed, which comes from a H - from one functional group and a -OH from another one are removed these reactions absorb energy † † hydrolysis reactions: aka catabolic reactions water is added to break a large molecule into smaller molecules produces energy enzymes are usually required carbohydrates general formula: (CH 2O) n , where “n” is some number three general groups: monosaccharides (single sugars) ex. glucose † monosaccharides are a single chain or ring of carbon atoms to which hydroxyl groups are attached. CH 2OH H O O C H H H C OH † H † †HO C H † OH H C † OH † H OH OH H C OH † H C OH † † H † H ex: † OH (solid and dissolved glucose, C6 H12O6 molecules) † † † † † † oligosaccharides (double sugars) ex. sucrose † 2 or more simple sugars attached by glycosidic linkages + Æ ex: glucose + glucose Æ maltose + H 2O † † † † + H 2O polysaccharides (many sugars) ex. cellulose, starch complex carbohydrates with 100-1000 simple sugars there are two kinds: energy sources and polysaccharides isomers: molecules with the same chemical formula, but a different arrangement of atoms, ex. glucose and galactose glycosidic linkages: 1-4 glycosidic linkage 1-6 glycosidic linkage, see page 31 lipids: usually hydrophobic composed of C, H and O, but has more C-H than O-H bonds used for storing energy, building membranes and chemical signals four families: fats, phospholipids, steroids, waxes carry more energy than proteins or CHOs, can act as insulation triglycerides (most common fat) are composed of 3 fatty acids attached to 1 molecule of glycerol ( ) fatty acid: long hydrocarbon chains with -COOH at the end saturated fatty acids have no double bonds (solid @ room temp.) † multiple bonds (liquid @ room temp.) unsaturated fatty acids have ester linkages are formed by condensation reactions between fatty acids and glycerol ( ), called esterification steroids (sterols) are hydrophobic molecules containing four fused hydrocarbon rings and other functional groups phospholipids are glycerol +2 fatty acids +1 polar phosphate group waxes contain long-chain fatty acids linked to alcohols or carbon rings waxes are hydrophobic, firm, pliable amino acids: general structure: proteins: can be biological catalysts transport molecules, in haemoglobin, keratin, etc can be shown as ionized: amino acid monomers are called peptides peptides are synthesized in cytoplasm through ‘protein synthesis’ bonds between amino acid monomers are called peptide bonds, formed by condensation reactions. These bonds are called amide linkages + Æ DNA specifies † the order of the amino acids when they are assembled for review questions see p. 50# 19, 20, 23, 24, 25, 26, pp. 86-87# 1-15