* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Loss of Mismatched HLA in Leukemia after Stem
Molecular mimicry wikipedia , lookup
Adaptive immune system wikipedia , lookup
Lymphopoiesis wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Innate immune system wikipedia , lookup
Sjögren syndrome wikipedia , lookup
A30-Cw5-B18-DR3-DQ2 (HLA Haplotype) wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
X-linked severe combined immunodeficiency wikipedia , lookup
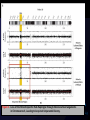
Loss of Mismatched HLA in Leukemia after Stem-Cell Transplantation N Engl J Med 2009;361:478-88 R2 Jung Kook Wi / prof. Kyung Sam Cho Background Transplantation of hematopoietic stem cells – from a haploidentical family donor – Curative option for patients with high-risk hematologic cancers The major limitation of this strategy – the risk of severe GVHD – alloreactions mediated by donor T cells – against the recipient’s unshared HLA haplotype The feasibility & efficacy of infusions of haploidentical donor T cells Background The infusion of donor T cells Rapid reconstitution of the immune system The graft-versus-leukemia Relapse still ??? Mechanism ??? Genomic loss Of the recipient’s mismatched HLA haplotype Mechanism of tumor escape from the selective pressure of a patient-specific graft-versus-leukemia reaction Methods Patients Retrospectively – Hematologic cancers – One or more haploidentical hematopoietic stem-cell transplantations – At the San Raffaele Hospital in Milan ( 2002 - 2007 ) All (43 patient) : high-risk hematologic myeloid cancers – 36 with AML & 7 with high-risk MDS – 26 with transplantation & 17 more than one Methods Chimerism Analyses Monthly in samples of bone marrow aspirate – short-tandem-repeat amplification (STR) – genomic HLA typing Loss of Heterozygosity – The use of polymerase-chainreaction amplification of 12 highly polymorphic short-tandem-repeat markers spanning the entire length of chromosome 6 – The Illumina Human CNV370-Quad Bead Array – The Affymetrix Human SNP Array 6.0 single-nucleotidepolymorphism (SNP) array Methods In Vitro Evaluation of Graft-versus-Leukemia Effect Separated peripheral-blood mononuclear cells – from the stem-cell donor for Patient 16 – from Patient 16 • 85 days after the first hematopoietic stem-cell transplantation • 96 days after the second transplantation – from a healthy HLA-mismatched subject Cells plated with irradiated mononuclear cells The function of responder cells from the mixed lymphocyte culture – tested after each stimulation – the use of 51Cr-release, enzyme-linked immunospot (ELISpot) – [3H] thymidine-incorporation assays Results Clinical Observations Studies of donor–host hematopoietic chimerism – carried out monthly after transplantation in all 43 patients – short-tandem-repeat amplification and HLA typing – looking for a reappearance of the host hematopoiesis in the BM – often predicts relapse In all 17 patients, relapse was confirmed – host origin on the basis of short-tandem-repeat chimerism Surprisingly, in five of these patients, genomic HLA typing of BM cells did not detect host-specific HLA alleles. Figure 1 Identification of Mutant Variants of Leukemic Cells Not Detected by Chimerism Analysis with HLA Typing. Figure 2. Genomic Loss of the Patient-Specific HLA Haplotype by Leukemic Cells after Transplantation in Patient 16. Figure 3 . Loss of the Patient-Specific HLA Haplotype through Extensive Rearrangements in Chromosome 6, Leading to Acquired Uniparental Disomy. Figure 3 . Loss of the Patient-Specific HLA Haplotype through Extensive Rearrangements in Chromosome 6, Leading to Acquired Uniparental Disomy. Functional Study of the Graft-versusLeukemia Response Conclusions After transplantation of haploidentical hematopoietic stem cells and infusion of donor T cells, leukemic cells can escape from the donor’s anti - leukemic T cells through the loss of the mismatched HLA haplotype. This event leads to relapse.