Introduction to cDNA Microarray Technology complementary DNA
... microarray slide. • Dyed sequences hybridize to the complementary probes that have been spotted on the array. All spots of the same color are made at the same time. ...
... microarray slide. • Dyed sequences hybridize to the complementary probes that have been spotted on the array. All spots of the same color are made at the same time. ...
SPLIT RNA Extraction Kit: Pure Fractions for Demanding Applications
... (especially on-column DNase digestion) or the enzyme inactivation (e.g., by heat denaturation) can severely compromise RNA integrity. Similarly, size-filtration based methods such as gDNA removal columns result in either ineffective gDNA removal or exclusion of longer RNA molecules (Fig. 3). 5 min ...
... (especially on-column DNase digestion) or the enzyme inactivation (e.g., by heat denaturation) can severely compromise RNA integrity. Similarly, size-filtration based methods such as gDNA removal columns result in either ineffective gDNA removal or exclusion of longer RNA molecules (Fig. 3). 5 min ...
Differential roles of TGIF family genes in mammalian reproduction Open Access
... conserved downstream and upstream genes in most of species. Human genomic sequence encompassing MYOM1 (partial sequence and not shown)-TGIF1-DLGAP1 (chr18: 3,395,327-3,507,935) compared to the mouse (chr17: 71,147,775-71,255,269), opossum (chr3: 269,037,689269,144,991), platypus (Contig3116: 1-55,20 ...
... conserved downstream and upstream genes in most of species. Human genomic sequence encompassing MYOM1 (partial sequence and not shown)-TGIF1-DLGAP1 (chr18: 3,395,327-3,507,935) compared to the mouse (chr17: 71,147,775-71,255,269), opossum (chr3: 269,037,689269,144,991), platypus (Contig3116: 1-55,20 ...
Introduction to molecular and cell biology
... a ~300 nucleotide transcript (fasX) • not encoding for a peptide/protein ...
... a ~300 nucleotide transcript (fasX) • not encoding for a peptide/protein ...
Supplementary Material PDF
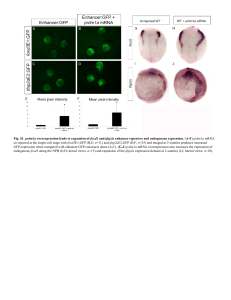
... Fig. S3. Double fluorescent ISH shows mosaic colocalization of enhancer:GFP with endogenous foxd3 and tfap2a mRNA expression. (A-F) Double fluorescent ISH for gfp (A) and foxd3 (B) in foxd3E1:GFP-injected embryos (merge in C, lateral view) and for gfp (D) and tfap2a (E) in tfap2aE2:GFP-injected emb ...
... Fig. S3. Double fluorescent ISH shows mosaic colocalization of enhancer:GFP with endogenous foxd3 and tfap2a mRNA expression. (A-F) Double fluorescent ISH for gfp (A) and foxd3 (B) in foxd3E1:GFP-injected embryos (merge in C, lateral view) and for gfp (D) and tfap2a (E) in tfap2aE2:GFP-injected emb ...
Connections between mRNA 3( end processing and transcription
... step is CTD-independent whereas the second is not [29]. An interesting possibility is that the pausing step reflects a processivity change in the elongation complex consistent with the anti-terminator model, while the CTD-dependent release step could be related to recruitment of the polyadenylation ...
... step is CTD-independent whereas the second is not [29]. An interesting possibility is that the pausing step reflects a processivity change in the elongation complex consistent with the anti-terminator model, while the CTD-dependent release step could be related to recruitment of the polyadenylation ...
ARGONAUTE1 Acts in Arabidopsis Root Radial
... miRNAs and their target genes (Rajagopalan et al. 2006, Fahlgren et al. 2007). Processed from imperfectly complementary stem–loop precursor RNAs (Bartel 2004), miRNAs regulate gene expression by targeting their complementary mRNA for cleavage or translational inhibition (Tang et al. 2003, Chen 2004) ...
... miRNAs and their target genes (Rajagopalan et al. 2006, Fahlgren et al. 2007). Processed from imperfectly complementary stem–loop precursor RNAs (Bartel 2004), miRNAs regulate gene expression by targeting their complementary mRNA for cleavage or translational inhibition (Tang et al. 2003, Chen 2004) ...
Interfering RNA
... Gene Walk Conclusions • Probability of finding an individual functional RNAi is high – A broad claim to “An isolated RNAi that inhibits the expression of human gene X.” may be enabled by providing the sequence for gene X and gene walk data (no magic number) ...
... Gene Walk Conclusions • Probability of finding an individual functional RNAi is high – A broad claim to “An isolated RNAi that inhibits the expression of human gene X.” may be enabled by providing the sequence for gene X and gene walk data (no magic number) ...
Recent retrotransposition events have not affected
... TEs may accumulate upstream because they have a role in chromatin regulation. (Huda et al, Gene, 2009) TEs may preferentially insert upstream (as known for Drosophila P elements) ...
... TEs may accumulate upstream because they have a role in chromatin regulation. (Huda et al, Gene, 2009) TEs may preferentially insert upstream (as known for Drosophila P elements) ...
Module 1 - Bioinformatics.ca
... Why sequence RNA (versus DNA)? • Interpreting mutations that do not have an obvious effect on protein sequence – ‘Regulatory’ mutations that affect what mRNA isoform is expressed and how much • e.g. splice sites, promoters, exonic/intronic splicing motifs, etc. ...
... Why sequence RNA (versus DNA)? • Interpreting mutations that do not have an obvious effect on protein sequence – ‘Regulatory’ mutations that affect what mRNA isoform is expressed and how much • e.g. splice sites, promoters, exonic/intronic splicing motifs, etc. ...
RNA/DNA catalysts
... Understand the basics of RNA/DNA catalysts - what functional groups used for catalysis? structures formed? Know about transesterification & cleavage reactions Know four types of natural catalytic RNAs (group I introns, group II introns, RNase P, small self-cleaving), what reactions they perform, kno ...
... Understand the basics of RNA/DNA catalysts - what functional groups used for catalysis? structures formed? Know about transesterification & cleavage reactions Know four types of natural catalytic RNAs (group I introns, group II introns, RNase P, small self-cleaving), what reactions they perform, kno ...
Qβ replicase discriminates between legitimate and illegitimate
... close to one another, which favors their annealing. • These stands immediately collapse into the double helix under action of proteases and detergents that cannot affect the stability of the RNA secondary structure, but destroy or unfold the protein structure. ...
... close to one another, which favors their annealing. • These stands immediately collapse into the double helix under action of proteases and detergents that cannot affect the stability of the RNA secondary structure, but destroy or unfold the protein structure. ...
View PDF
... which it is associated in sheep (‘Callipyge’ means ‘beautiful bottom’) [27]. Many similarities have been identified between the PWS–AS and CLPG loci. For example, the CLPG locus also contains multiple paternally expressed genes, including DLK1 (overexpression of which causes the CPLG phenotype [28,2 ...
... which it is associated in sheep (‘Callipyge’ means ‘beautiful bottom’) [27]. Many similarities have been identified between the PWS–AS and CLPG loci. For example, the CLPG locus also contains multiple paternally expressed genes, including DLK1 (overexpression of which causes the CPLG phenotype [28,2 ...
Relationship between expression and methylation of obesity
... has been utilised for transcriptomics in studies investigating a variety of topics including cancer (18,19), infectious disease (20) and immunology (21), but has yet to be employed in the assessment of expression changes associated with obesity in humans. The nCounter Analysis System has a variety o ...
... has been utilised for transcriptomics in studies investigating a variety of topics including cancer (18,19), infectious disease (20) and immunology (21), but has yet to be employed in the assessment of expression changes associated with obesity in humans. The nCounter Analysis System has a variety o ...
-Chain Gene in Epididymis α Expression of the C4b
... (C4BP␣) and have demonstrated significant C4BP␣ mRNA expression in epididymis as well as liver. The level of C4BP␣ transcripts increased in the epididymis after birth, while it remained constant in the liver. C4BP␣ mRNA was also detected in the normal murine epididymis at a significant level, but it ...
... (C4BP␣) and have demonstrated significant C4BP␣ mRNA expression in epididymis as well as liver. The level of C4BP␣ transcripts increased in the epididymis after birth, while it remained constant in the liver. C4BP␣ mRNA was also detected in the normal murine epididymis at a significant level, but it ...
network - bioinf leipzig
... Regulatory interactions can also be inferred directly from data = reverse engineering of biological pathways/networks from data. In the example above time-series expression data61is used to infer a directed and signed graph based on delayed correlations. ...
... Regulatory interactions can also be inferred directly from data = reverse engineering of biological pathways/networks from data. In the example above time-series expression data61is used to infer a directed and signed graph based on delayed correlations. ...
PowerPoint 簡報
... •Initiation: A promoter is the DNA sequence that initially binds the RNA polymerase. Only one of the DNA strands acts as a template. The choice of promoter determines which stretch of DNA is transcribed and is the main step at which regulation is imposed. •Elongation: Once the RNA polymerase has sy ...
... •Initiation: A promoter is the DNA sequence that initially binds the RNA polymerase. Only one of the DNA strands acts as a template. The choice of promoter determines which stretch of DNA is transcribed and is the main step at which regulation is imposed. •Elongation: Once the RNA polymerase has sy ...
Codon Dictionary Worksheet
... So, if the mRNA codon is UUU, what tRNA anticodon is complementary? (Answer: AAA) If the mRNA codon is UAC, what tRNA anticodon is complementary? (Answer #4 below) If the mRNA codon is GGU, what tRNA anticodon is complementary? (Answer #5 below) If the tRNA anticodon is UAC, with which mRNA codon do ...
... So, if the mRNA codon is UUU, what tRNA anticodon is complementary? (Answer: AAA) If the mRNA codon is UAC, what tRNA anticodon is complementary? (Answer #4 below) If the mRNA codon is GGU, what tRNA anticodon is complementary? (Answer #5 below) If the tRNA anticodon is UAC, with which mRNA codon do ...
Chapter 11 Transcription and RNA Processing
... • RNA synthesis, catalyzed by RNA polymerases, is similar to DNA synthesis in many respects. • RNA synthesis occurs within a localized region of strand separation (Transcription Bubble), and only one strand of DNA functions as a template for RNA synthesis. © John Wiley & Sons, Inc. ...
... • RNA synthesis, catalyzed by RNA polymerases, is similar to DNA synthesis in many respects. • RNA synthesis occurs within a localized region of strand separation (Transcription Bubble), and only one strand of DNA functions as a template for RNA synthesis. © John Wiley & Sons, Inc. ...
Document
... DNA contains the information needed to make proteins. However, DNA is too large to leave the nucleus. RNA acts as a set of working instructions for ribosomes to make proteins. This process is also known as gene expression. Gene expression is a regulated process. ...
... DNA contains the information needed to make proteins. However, DNA is too large to leave the nucleus. RNA acts as a set of working instructions for ribosomes to make proteins. This process is also known as gene expression. Gene expression is a regulated process. ...
Introduction to RNA sequencing
... Why sequence RNA (versus DNA)? • Interpreting mutations that do not have an obvious effect on protein sequence – ‘Regulatory’ mutations that affect what mRNA isoform is expressed and how much • e.g. splice sites, promoters, exonic/intronic splicing motifs, etc. ...
... Why sequence RNA (versus DNA)? • Interpreting mutations that do not have an obvious effect on protein sequence – ‘Regulatory’ mutations that affect what mRNA isoform is expressed and how much • e.g. splice sites, promoters, exonic/intronic splicing motifs, etc. ...
Central Dogma of Molecular Biology Chapter 28 DNA Replication
... RNA splicing is not merely a curiosity. At least 15% of all genetic diseases have been associated with mutations that affect RNA splicing. Moreover, the same pre-mRNA can be spliced differently in various cell types, at different stages of development, or in response to other biological signals. (A ...
... RNA splicing is not merely a curiosity. At least 15% of all genetic diseases have been associated with mutations that affect RNA splicing. Moreover, the same pre-mRNA can be spliced differently in various cell types, at different stages of development, or in response to other biological signals. (A ...
Glycosylphosphatidyl inositol-anchored protein (GPI
... means of quantitative RT-PCR of RNA from 48 thymoma tumors. We found that GPI-80 was significantly higher in invasive thymoma (stage IV thymoma) than in stage I thymoma. It has previously been shown that the GPI-80 is a possible regulatory molecule of cell adhesion and migration.18–20) GPI80 protein ...
... means of quantitative RT-PCR of RNA from 48 thymoma tumors. We found that GPI-80 was significantly higher in invasive thymoma (stage IV thymoma) than in stage I thymoma. It has previously been shown that the GPI-80 is a possible regulatory molecule of cell adhesion and migration.18–20) GPI80 protein ...
MicroRNA
A micro RNA (abbreviated miRNA) is a small non-coding RNA molecule (containing about 22 nucleotides) found in plants, animals, and some viruses, which functions in RNA silencing and post-transcriptional regulation of gene expression.Encoded by eukaryotic nuclear DNA in plants and animals and by viral DNA in certain viruses whose genome is based on DNA, miRNAs function via base-pairing with complementary sequences within mRNA molecules. As a result, these mRNA molecules are silenced by one or more of the following processes: 1) cleavage of the mRNA strand into two pieces, 2) destabilization of the mRNA through shortening of its poly(A) tail, and 3) less efficient translation of the mRNA into proteins by ribosomes. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1000 miRNAs, which are abundant in many mammalian cell types and appear to target about 60% of the genes of humans and other mammals.miRNAs are well conserved in both plants and animals, and are thought to be a vital and evolutionarily ancient component of genetic regulation. While core components of the microRNA pathway are conserved between plants and animals, miRNA repertoires in the two kingdoms appear to have emerged independently with different primary modes of action. Plant miRNAs usually have near-perfect pairing with their mRNA targets, which induces gene repression through cleavage of the target transcripts. In contrast, animal miRNAs are able to recognize their target mRNAs by using as little as 6–8 nucleotides (the seed region) at the 5' end of the miRNA, which is not enough pairing to induce cleavage of the target mRNAs. Combinatorial regulation is a feature of miRNA regulation in animals. A given miRNA may have hundreds of different mRNA targets, and a given target might be regulated by multiple miRNAs.The first miRNA was discovered in the early 1990s. However, miRNAs were not recognized as a distinct class of biological regulators until the early 2000s. Since then, miRNA research has revealed different sets of miRNAs expressed in different cell types and tissuesand has revealed multiple roles for miRNAs in plant and animal development and in many other biological processes. Aberrant expression of miRNAs has been implicated in numerous disease states, and miRNA-based therapies are under investigation.Estimates of the average number of unique messenger RNAs that are targets for repression by a typical microRNA vary, depending on the method used to make the estimate, but several approaches show that mammalian miRNAs can have many unique targets. For example, an analysis of the miRNAs highly conserved in vertebrate animals shows that each of these miRNAs has, on average, roughly 400 conserved targets. Likewise, experiments show that a single miRNA can reduce the stability of hundreds of unique messenger RNAs, and other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold).