17. Gene regulation
... and determines which allele of a gene is expressed for certain genes (immunoglobulin genes, genes on X-chromosome, etc.) 2. Histone modification histones are basic proteins that help compact DNA act as generalized repressors of transcription dense packed chromatin: transcriptionally inactive ...
... and determines which allele of a gene is expressed for certain genes (immunoglobulin genes, genes on X-chromosome, etc.) 2. Histone modification histones are basic proteins that help compact DNA act as generalized repressors of transcription dense packed chromatin: transcriptionally inactive ...
Eukaryotic Gene Control
... • Multicellular organisms have developmental pathways from zygote to adult • Developmental sequences are predominately determined and programmed by differential gene expression. ...
... • Multicellular organisms have developmental pathways from zygote to adult • Developmental sequences are predominately determined and programmed by differential gene expression. ...
Document
... Groups of structural genes with related functions + DNA responsible for controlling ...
... Groups of structural genes with related functions + DNA responsible for controlling ...
Eukaryotic Gene Expression
... specialization of cells during development • Since all cells have the same DNA, how can differentiation occur? • Gene regulation. ...
... specialization of cells during development • Since all cells have the same DNA, how can differentiation occur? • Gene regulation. ...
Eukaryotic transcriptional control
... Eukaryotic transcriptional control: major considerations 1. Interplay among multiple general transcription factors; activators/repressors; mediator/coactivators ...
... Eukaryotic transcriptional control: major considerations 1. Interplay among multiple general transcription factors; activators/repressors; mediator/coactivators ...
Epigenet-web
... CpGs are vastly underrepresented genome-wide compared to what would be expected by chance (0.23 in the human genome and 0.19 in the mouse genome, respectively) This is because deamination of cytosine gives rise to uracil, which is easily recognized as foreign within the DNA strand and replaced, wher ...
... CpGs are vastly underrepresented genome-wide compared to what would be expected by chance (0.23 in the human genome and 0.19 in the mouse genome, respectively) This is because deamination of cytosine gives rise to uracil, which is easily recognized as foreign within the DNA strand and replaced, wher ...
401Lecture8Sp2013post
... • Activators can direct histone acetylation and methylation to specific genes • Repressors can direct histone deacetylation, histone methylation and DNA methylation to specific genes • Repressors and microRNAs can recruit polycomb group complexes to promote heterochromatin formation • Epigenetic mar ...
... • Activators can direct histone acetylation and methylation to specific genes • Repressors can direct histone deacetylation, histone methylation and DNA methylation to specific genes • Repressors and microRNAs can recruit polycomb group complexes to promote heterochromatin formation • Epigenetic mar ...
Chapter 13 Chromatin Structure and its Effects on
... Is this caused by the promoter or something else near the promoter? Can this occur with the promoter at a different location? Try a modified SV40 that has a second promoter inserted. Does transcription cause this or is it caused by something else that binds the promoter? Do we need to have transcrip ...
... Is this caused by the promoter or something else near the promoter? Can this occur with the promoter at a different location? Try a modified SV40 that has a second promoter inserted. Does transcription cause this or is it caused by something else that binds the promoter? Do we need to have transcrip ...
Chromatin Remodeling and Gene Expression
... Potentiation and Activation. Histone modifications associated with various phas promoter states are shown as symbols at right. Experimentally verified and putative pathways leading to phas activation are shown as blue (solid) and red (dotted) lines, respectively. (A) In the repressed state during ve ...
... Potentiation and Activation. Histone modifications associated with various phas promoter states are shown as symbols at right. Experimentally verified and putative pathways leading to phas activation are shown as blue (solid) and red (dotted) lines, respectively. (A) In the repressed state during ve ...
Analytical and Chromatography - Sigma
... Linkage Between DNA Repair and Chromatin Modification • Biochemical experiments have permitted the identification of acidic factors that can form complexes with histones and enhance the process of histone deposition. They act as histone chaperones by facilitating the formation of nucleosome cores w ...
... Linkage Between DNA Repair and Chromatin Modification • Biochemical experiments have permitted the identification of acidic factors that can form complexes with histones and enhance the process of histone deposition. They act as histone chaperones by facilitating the formation of nucleosome cores w ...
Eukaryotic Gene Regulation Exercise - KEY
... 4. Spliced mRNA differently… Some genes "loosened", others not… Some proteins targeted for degradation 5. A. Before transcription: Histone acetylation Molecule affected: DNA ...
... 4. Spliced mRNA differently… Some genes "loosened", others not… Some proteins targeted for degradation 5. A. Before transcription: Histone acetylation Molecule affected: DNA ...
DNA Methylation Histone Acetylation
... Epigenetic therapy with DNA methylation inhibitors (DNMTi) and HDAC inhibitors (HDACi). Combination therapies are likely to gain traction in the future because of the inherent self – reinforcing nature of silencing mechanisms. ...
... Epigenetic therapy with DNA methylation inhibitors (DNMTi) and HDAC inhibitors (HDACi). Combination therapies are likely to gain traction in the future because of the inherent self – reinforcing nature of silencing mechanisms. ...
Histone Methylation
... Condensation of DNA involves coiling around proteins called histones. Acetylation is when acetyl groups (COCH3) are attached to lysines in the histone tails. This reduces condensation and promotes transcription because the transcription machinery has better access to the DNA. ...
... Condensation of DNA involves coiling around proteins called histones. Acetylation is when acetyl groups (COCH3) are attached to lysines in the histone tails. This reduces condensation and promotes transcription because the transcription machinery has better access to the DNA. ...
20141203103493
... Acetylation of histone tails promotes loose chromatin structure that permits transcription ...
... Acetylation of histone tails promotes loose chromatin structure that permits transcription ...
Go-ChIP-Grade™ Purified anti-Histone H3 (C-terminus
... approximately 146bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4) limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones play a central role in transcription regulation, DNA repair, DNA rep ...
... approximately 146bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4) limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones play a central role in transcription regulation, DNA repair, DNA rep ...
New Ligands of CRABP2 Suggest a Role for this Protein in
... Retinoic acid (RA) regulates transcription of a series of genes involved in cell proliferation, differentiation and apoptosis by binding to the RA receptor (RAR) and retinoid X receptor (RXR) heterodimers. The cellular retinoic acid-binding protein 2 (CRABP2) is involved in the transport of RA from ...
... Retinoic acid (RA) regulates transcription of a series of genes involved in cell proliferation, differentiation and apoptosis by binding to the RA receptor (RAR) and retinoid X receptor (RXR) heterodimers. The cellular retinoic acid-binding protein 2 (CRABP2) is involved in the transport of RA from ...
Go-ChIP-Grade™ Purified anti-Histone H3 (C
... approximately 146bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4) limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones play a central role in transcription regulation, DNA repair, DNA rep ...
... approximately 146bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4) limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones play a central role in transcription regulation, DNA repair, DNA rep ...
PowerPoint Presentation - Creighton Chemistry Webserver
... • Promoted by histone acetyltransferase (HAT) • Acetylation causes opening of histone protein arms, making DNA in chromatin more accessible • Acetylated histones are a mark of gene activity ...
... • Promoted by histone acetyltransferase (HAT) • Acetylation causes opening of histone protein arms, making DNA in chromatin more accessible • Acetylated histones are a mark of gene activity ...
Genetic Controls in Eukaryotes
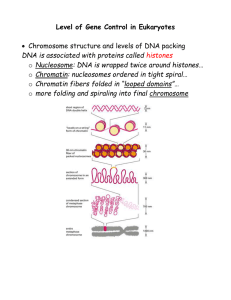
... Level of Gene Control in Eukaryotes Chromosome structure and levels of DNA packing DNA is associated with proteins called histones o Nucleosome: DNA is wrapped twice around histones… o Chromatin: nucleosomes ordered in tight spiral… o Chromatin fibers folded in “looped domains”… o more folding and ...
... Level of Gene Control in Eukaryotes Chromosome structure and levels of DNA packing DNA is associated with proteins called histones o Nucleosome: DNA is wrapped twice around histones… o Chromatin: nucleosomes ordered in tight spiral… o Chromatin fibers folded in “looped domains”… o more folding and ...
product data sheet
... peptide can bind two copies of BRD2-2 (BRD2, bromodomain 2), each interacting with one of the two acetylated lysines . In an in vitro RNA polymerase II transcription system, binding of either BRD2 or BRD3 to a chromatin template assembled with hyperacetylated ...
... peptide can bind two copies of BRD2-2 (BRD2, bromodomain 2), each interacting with one of the two acetylated lysines . In an in vitro RNA polymerase II transcription system, binding of either BRD2 or BRD3 to a chromatin template assembled with hyperacetylated ...
Molecular mechanisms of the epigenetic regulation Tatiana G
... Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO 80045 USA Plant homeodomain (PHD) fingers, YEATS, Tudor and bromodomains are found in proteins involved in a wide array of fundamental biological processes, including transcription, replication, DNA damage repair, cell ...
... Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO 80045 USA Plant homeodomain (PHD) fingers, YEATS, Tudor and bromodomains are found in proteins involved in a wide array of fundamental biological processes, including transcription, replication, DNA damage repair, cell ...
Epigenetic regulators as novel treatments
... Some definitions: Epigenetics-the study of heritable changes in gene expression without changing the DNA sequence; this occurs at 3 levels of organization: 1) methylation of cytosine nucleotides within coding sequences and at promoter sites that alter transcription rates 2) changes in chromatin pro ...
... Some definitions: Epigenetics-the study of heritable changes in gene expression without changing the DNA sequence; this occurs at 3 levels of organization: 1) methylation of cytosine nucleotides within coding sequences and at promoter sites that alter transcription rates 2) changes in chromatin pro ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.