Slide 1 - AccessMedicine
... The central dogma. (A) The central dogma of molecular biology as originally described by Crick. This schematic illustrates the flow of genetic information from DNA to RNA and then to protein, as well as illustrates that DNA serves as its own template in replication. (B) The new central dogma reflect ...
... The central dogma. (A) The central dogma of molecular biology as originally described by Crick. This schematic illustrates the flow of genetic information from DNA to RNA and then to protein, as well as illustrates that DNA serves as its own template in replication. (B) The new central dogma reflect ...
The Epigenetic Code regulates Chromatin Structure and
... Epigenetic regulation of Cellular Identity Epi- (greek: “over, above”) genetic: heritable changes in gene expression via mechanisms that are separate from the DNA sequence and regulated by environmental signals. Environment Organism level: Nutrition, Behavior, Pollution, Sun light, Toxins, Circadia ...
... Epigenetic regulation of Cellular Identity Epi- (greek: “over, above”) genetic: heritable changes in gene expression via mechanisms that are separate from the DNA sequence and regulated by environmental signals. Environment Organism level: Nutrition, Behavior, Pollution, Sun light, Toxins, Circadia ...
EpigEnEtiCS: A pRiMER
... goes back to Lamarck in the early 19th century, but still only correlative evidence exists in humans. In contrast, many cellular epigenetic phenomena are now well understood on the molecular level. In humans, they include the parent-of-origin specific expression of genes (imprinting) and the shuttin ...
... goes back to Lamarck in the early 19th century, but still only correlative evidence exists in humans. In contrast, many cellular epigenetic phenomena are now well understood on the molecular level. In humans, they include the parent-of-origin specific expression of genes (imprinting) and the shuttin ...
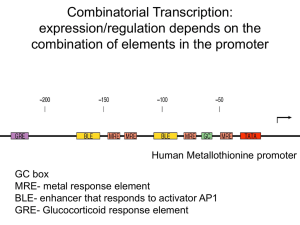
Combinatorial Transcription: expression/regulation depends on the
... • Addition of H1 further represses transcription (by binding to the linker DNA), but this can be overcome by activators such as Sp1. • There are regulatory proteins, such as the glucocorticoid-receptor complex, that can remove histones from certain promoters. ...
... • Addition of H1 further represses transcription (by binding to the linker DNA), but this can be overcome by activators such as Sp1. • There are regulatory proteins, such as the glucocorticoid-receptor complex, that can remove histones from certain promoters. ...
Molecular Genetics (Unit 6 and Unit 6.2) Study Guide Each of the
... o Endomembranes, plasmids, F plasmid, R Plasmid, Toxins, Asexual and sexual reproduction, pillus o Conjugation, transformation, transduction o Transposons, episome Reverse Transcription o The process o Components involved/job of each o The purpose Gene expression o Promote, regulatory, operator, str ...
... o Endomembranes, plasmids, F plasmid, R Plasmid, Toxins, Asexual and sexual reproduction, pillus o Conjugation, transformation, transduction o Transposons, episome Reverse Transcription o The process o Components involved/job of each o The purpose Gene expression o Promote, regulatory, operator, str ...
Ch 18.2-18.5 PPT
... methyl groups added to DNA; tightly packed; transcription Histone acetylation: acetyl groups added to histones; loosened; transcription ...
... methyl groups added to DNA; tightly packed; transcription Histone acetylation: acetyl groups added to histones; loosened; transcription ...
LOYOLA COLLEGE (AUTONOMOUS), CHENNAI – 600 034
... 4. Define DNA methylation. 5. List out the functions of UTRs. 6. What are Tumour suppressor genes? Give examples. 7. What are non -viral retro-transposons? 8. What is the importance of HMG proteins in nucleosome organization? 9. Define Transcription attenuation. 10. Comment on RNA life span. PART B ...
... 4. Define DNA methylation. 5. List out the functions of UTRs. 6. What are Tumour suppressor genes? Give examples. 7. What are non -viral retro-transposons? 8. What is the importance of HMG proteins in nucleosome organization? 9. Define Transcription attenuation. 10. Comment on RNA life span. PART B ...
Problems in Replication and Protein Synthesis
... Eukaryotic Regulation • Enhancers – sites that call for specific activators to stimulate the production of certain proteins. ...
... Eukaryotic Regulation • Enhancers – sites that call for specific activators to stimulate the production of certain proteins. ...
Regulation of Gene Expression
... o Since there are so many steps to gene expression, there are many possible levels of control o Each gene typically has its own promoter and regulatory regions o Chromatin structure affects gene expression o Activator proteins are more prevalent in eukaryotes o When DNA is tightly coiled around hist ...
... o Since there are so many steps to gene expression, there are many possible levels of control o Each gene typically has its own promoter and regulatory regions o Chromatin structure affects gene expression o Activator proteins are more prevalent in eukaryotes o When DNA is tightly coiled around hist ...
Dual function of histone H3K76 methylation in cell cycle regulation
... structure with an emphasis on posttranslational histone modifications (PTMs) and the enzymes that are responsible for them. After dissecting the repertoire of PTMs in two life cycle stages in trypanosomes, we focused on methylation of histone H3 on lysine 76 in the histone core, which is mediated by ...
... structure with an emphasis on posttranslational histone modifications (PTMs) and the enzymes that are responsible for them. After dissecting the repertoire of PTMs in two life cycle stages in trypanosomes, we focused on methylation of histone H3 on lysine 76 in the histone core, which is mediated by ...
Inquiry into Life Twelfth Edition
... chromosome also depends on the SIR proteins • SIR3 and SIR4 interact directly with histones H3 and H4 in nucleosomes – Acetylation of Lys 16 on H4 in nucleosomes prevents interaction with SIR3 ...
... chromosome also depends on the SIR proteins • SIR3 and SIR4 interact directly with histones H3 and H4 in nucleosomes – Acetylation of Lys 16 on H4 in nucleosomes prevents interaction with SIR3 ...
transcription
... 1. Histones form nucleosomes on TATA boxes, blocking transcription. Promoter-binding proteins cannot disrupt the nucleosomes. Enhancer-binding proteins bind to enhancers, displacing any histones, and then cause the histones at the TATA box to free the DNA. 2. Histone Acetylation with increased trans ...
... 1. Histones form nucleosomes on TATA boxes, blocking transcription. Promoter-binding proteins cannot disrupt the nucleosomes. Enhancer-binding proteins bind to enhancers, displacing any histones, and then cause the histones at the TATA box to free the DNA. 2. Histone Acetylation with increased trans ...
Regulation of Gene Expression
... Gene regulatory proteins, general transcription factors, coactivators, corepressors, chromatin and histone modifiers Common structural classes of gene regulatory proteins Zinc finger (762) Homeobox (199) Basic helix-loop-helix (117) Beta-scaffold (87) Basic-leucine zipper (72) Nuclear hormone recept ...
... Gene regulatory proteins, general transcription factors, coactivators, corepressors, chromatin and histone modifiers Common structural classes of gene regulatory proteins Zinc finger (762) Homeobox (199) Basic helix-loop-helix (117) Beta-scaffold (87) Basic-leucine zipper (72) Nuclear hormone recept ...
Presentación de PowerPoint
... Do not allow histone wrapped around DNA. Most of the DNA of a human cell is contained in the nucleus. Distinguish between unique and highly repetitive sequences in nuclear DNA. ...
... Do not allow histone wrapped around DNA. Most of the DNA of a human cell is contained in the nucleus. Distinguish between unique and highly repetitive sequences in nuclear DNA. ...
Chapter 19 Organization and Control of Eukaryotic Genomes
... Can cause Genetic disorders. Typically found in centromeres and telomeres so it is thought to be used for structure. Interspersed Repetitive DNA—Copies of similar sequences but not repetitive. ...
... Can cause Genetic disorders. Typically found in centromeres and telomeres so it is thought to be used for structure. Interspersed Repetitive DNA—Copies of similar sequences but not repetitive. ...
Table S11 Properties of the transcription factors for which binding
... bZIP transcription factor (ATF/CREB1 homolog) that regulates the unfolded protein response, via UPRE binding, and membrane biogenesis; ER stress-induced splicing pathway utilizing Ire1p, Trl1p and Ada5p facilitates efficient Hac1p synthesis ...
... bZIP transcription factor (ATF/CREB1 homolog) that regulates the unfolded protein response, via UPRE binding, and membrane biogenesis; ER stress-induced splicing pathway utilizing Ire1p, Trl1p and Ada5p facilitates efficient Hac1p synthesis ...
Control of Gene Expression
... Complementary strands bind to one another Gene sequence may allow formation of a ...
... Complementary strands bind to one another Gene sequence may allow formation of a ...
Histone depleted metaphase chromosomes Scaffold Attachment
... the interior • Some are also found on loops outside of the territory ...
... the interior • Some are also found on loops outside of the territory ...
Ch 15 - .Gene Regulation
... cellular activity? What word describes the attachment of groups of particular amino acids of specific proteins to nucleosomes as thought to be an important control mechanism for gene ...
... cellular activity? What word describes the attachment of groups of particular amino acids of specific proteins to nucleosomes as thought to be an important control mechanism for gene ...
cancer epigenetics - Experimental oncology
... to all heritable changes in gene expression and chromatin organization that do not involve sequence changes in DNA. It includes three distinct and self-reinforcing mechanisms: aberrations in DNA methylation, posttranslational modifications of histones and chromatin remodeling; non-protein-coding RNA ...
... to all heritable changes in gene expression and chromatin organization that do not involve sequence changes in DNA. It includes three distinct and self-reinforcing mechanisms: aberrations in DNA methylation, posttranslational modifications of histones and chromatin remodeling; non-protein-coding RNA ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.