DNA methylation
... histone deacetylases (HDACs). • 6 HAT complexes and 18 HDACs have been identified in mammals. • Acetylation brings in a negative charge, acting to neutralize the positive charge on the histones and decreases the interaction of the N termini of histones with the negatively charged phosphate groups of ...
... histone deacetylases (HDACs). • 6 HAT complexes and 18 HDACs have been identified in mammals. • Acetylation brings in a negative charge, acting to neutralize the positive charge on the histones and decreases the interaction of the N termini of histones with the negatively charged phosphate groups of ...
Lecture 14 Student Powerpoint
... Eukaryotes Previously we discussed aspects of transcription that involved the production of a primary transcript and the processing of this into aspects of how this is processed into an mRNA in Eukaryotes. This lecture focuses on these topics and on how signals are perceived that modulate the expres ...
... Eukaryotes Previously we discussed aspects of transcription that involved the production of a primary transcript and the processing of this into aspects of how this is processed into an mRNA in Eukaryotes. This lecture focuses on these topics and on how signals are perceived that modulate the expres ...
SBI 4U Genetics 5
... cell’s DNA and causes substitution or frameshift changes. EG. Gasoline fumes, nitrites and compounds found in cigarette smoke Physical mutagens: physically change the DNA ...
... cell’s DNA and causes substitution or frameshift changes. EG. Gasoline fumes, nitrites and compounds found in cigarette smoke Physical mutagens: physically change the DNA ...
Signaling pathway
... components of the WNT signaling pathways e.g. important for maintaining the stem cell population in gastro-intestinal tract, Over-activation by APC mutation - cancer ...
... components of the WNT signaling pathways e.g. important for maintaining the stem cell population in gastro-intestinal tract, Over-activation by APC mutation - cancer ...
on-chip
... (MeDIP) or the Methylated CpG Island Recovery Assay (MIRA), followed by microarray analysis. CpG islands are genomic regions that contain dense clusters of CG dinucleotides that are often ...
... (MeDIP) or the Methylated CpG Island Recovery Assay (MIRA), followed by microarray analysis. CpG islands are genomic regions that contain dense clusters of CG dinucleotides that are often ...
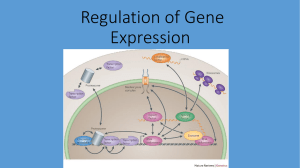
Regulation of Gene Expression
... • Pre-transcriptional control: the cell controls the extent to which DNA is exposed to transcription enzymes, regulating DNA’s availability for transcription. • Transcriptional control: the cell controls whether or not exposed DNA is transcribed into Pre-mRNA. • Post-transcriptional control: the cel ...
... • Pre-transcriptional control: the cell controls the extent to which DNA is exposed to transcription enzymes, regulating DNA’s availability for transcription. • Transcriptional control: the cell controls whether or not exposed DNA is transcribed into Pre-mRNA. • Post-transcriptional control: the cel ...
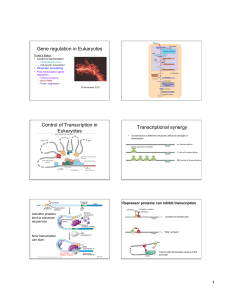
Regulation of Gene Expression in Eukaryotes
... – Function independent of position – upstream, downstream, etc. (different than promotors‐ close to gene and only one orientation) ...
... – Function independent of position – upstream, downstream, etc. (different than promotors‐ close to gene and only one orientation) ...
Microsoft Word
... could indicate a cycle of functional trafficking between the nucleus and cytoplasm in early embryogenesis. Small RNA pathways are vital mechanisms for genome regulation at the epigenetic level. Per se epigenetic regulation is a phenomenon that is responsible for generating and maintaining diversity ...
... could indicate a cycle of functional trafficking between the nucleus and cytoplasm in early embryogenesis. Small RNA pathways are vital mechanisms for genome regulation at the epigenetic level. Per se epigenetic regulation is a phenomenon that is responsible for generating and maintaining diversity ...
Read Project Details
... Brief Description. The research project is to determine how genes involved in anaerobiosis are induced by oxygen limitation and nitric oxide (NO) in Bacillus subtilis. Our previous study showed that the ResD-ResE two-component regulatory proteins and the NO-sensitive NsrR repressor play major roles ...
... Brief Description. The research project is to determine how genes involved in anaerobiosis are induced by oxygen limitation and nitric oxide (NO) in Bacillus subtilis. Our previous study showed that the ResD-ResE two-component regulatory proteins and the NO-sensitive NsrR repressor play major roles ...
Note 7.4 - Controlling Gene Expression
... Posttranslational: before many proteins become functional, they must pass through the cell membrane. A number of control mechanisms affect the rate at which a protein becomes active and the time it remains functional, including the addition of various chemical groups. ...
... Posttranslational: before many proteins become functional, they must pass through the cell membrane. A number of control mechanisms affect the rate at which a protein becomes active and the time it remains functional, including the addition of various chemical groups. ...
Histone Modifications - Life Science Saga
... http://lifesciencesaga.weebly.com http://purnasrinivas.weebly.com ...
... http://lifesciencesaga.weebly.com http://purnasrinivas.weebly.com ...
HDAC inhibitor drug protects memory in HD mice
... hippocampus gets its name from the Greek for ‘seahorse’ because it has the same shape.) Levels of CBP were much lower in the hippocampus of the HD mice. Interestingly, the CBP that was seen, appeared to be stuck to blobs of mutant huntingtin - as if the mutant protein was ‘trapping’ the CBP. Since C ...
... hippocampus gets its name from the Greek for ‘seahorse’ because it has the same shape.) Levels of CBP were much lower in the hippocampus of the HD mice. Interestingly, the CBP that was seen, appeared to be stuck to blobs of mutant huntingtin - as if the mutant protein was ‘trapping’ the CBP. Since C ...
Nuclear gene expression 1
... 1. Supercoiling of the promoter DNA 2. Sliding of the complex to the promoter 3. Looping out of DNA between enhancer and promoter ...
... 1. Supercoiling of the promoter DNA 2. Sliding of the complex to the promoter 3. Looping out of DNA between enhancer and promoter ...
Advanced Biology\Stem Cells, histones, etc
... factors, miRs, etc. can affect it as can protein activation. Our protein synthesis/use can be controlled via phosphorylation. Using a kinase, such as tyrosine kinase, an inactive protein can become activated by adding a phosphate (from ATP). A protein can be deactivated by removing a phosphate using ...
... factors, miRs, etc. can affect it as can protein activation. Our protein synthesis/use can be controlled via phosphorylation. Using a kinase, such as tyrosine kinase, an inactive protein can become activated by adding a phosphate (from ATP). A protein can be deactivated by removing a phosphate using ...
A1979HZ32700001
... "Soon after the publication of this paper, Dr. Geschwind and I had mostly, but not exclusively favorable comments on it. A late, well-known biochemist from New York expressed his disdain for this and similar quantitative methods when they are used in conjunction with a microscope, rather than a test ...
... "Soon after the publication of this paper, Dr. Geschwind and I had mostly, but not exclusively favorable comments on it. A late, well-known biochemist from New York expressed his disdain for this and similar quantitative methods when they are used in conjunction with a microscope, rather than a test ...
Eukaryotic Expression 1
... • Catabolite Activator Protein (CAP) is one such protein • CAP is a homodimer of 22.5 kD peptides • N-term binds cAMP; C-term binds DNA • Binding of CAP-(cAMP)2 to DNA assists formation of closed promoter complex [also called CRP (cAMP receptor protein)] ...
... • Catabolite Activator Protein (CAP) is one such protein • CAP is a homodimer of 22.5 kD peptides • N-term binds cAMP; C-term binds DNA • Binding of CAP-(cAMP)2 to DNA assists formation of closed promoter complex [also called CRP (cAMP receptor protein)] ...
Transcription Factors
... • “Downstream” = being 3’ to the start site – Positive numbers of bases ...
... • “Downstream” = being 3’ to the start site – Positive numbers of bases ...
Gene Section POU2AF1 (POU domain, class 2, associating factor 1)
... Spans on a 30 kb genomic fragment; five exons; large fifth exon, with many 3'-UTR repetitive elements, two pyrimidine rich regions (a duplicated CT-rich region and a [CCTT]n tetranucleotide tandem repeat) and a ...
... Spans on a 30 kb genomic fragment; five exons; large fifth exon, with many 3'-UTR repetitive elements, two pyrimidine rich regions (a duplicated CT-rich region and a [CCTT]n tetranucleotide tandem repeat) and a ...
BSN/Briefing 24 - British Society for Neuroendocrinology
... independent of DNA sequence. Epigenetic regulation therefore imparts a memory of transcriptional states through modification of histones, histone variants, chromatin remodelling by ATP-dependent complexes and DNA methylation (see figure). Cellular phenotypes transmitted in this mode include parental ...
... independent of DNA sequence. Epigenetic regulation therefore imparts a memory of transcriptional states through modification of histones, histone variants, chromatin remodelling by ATP-dependent complexes and DNA methylation (see figure). Cellular phenotypes transmitted in this mode include parental ...
Notes
... Both alleles of BRCA1 or both alleles of BRCA2 must be mutant for cancer to develop. Why would in follow a dominant inheritance pattern? ...
... Both alleles of BRCA1 or both alleles of BRCA2 must be mutant for cancer to develop. Why would in follow a dominant inheritance pattern? ...
First Annual Research EXTRAVANGA - 2010
... molecular changes similar to SZ postmortem brain: a significant increase in DNMT1 and TET1 in the FC and HP but not in cerebellum, no changes in HDACs, histone methytransferases/demethylases or MeCP2, and a significant decrease in BDNF variants. The decrease of the corresponding BDNF transcript leve ...
... molecular changes similar to SZ postmortem brain: a significant increase in DNMT1 and TET1 in the FC and HP but not in cerebellum, no changes in HDACs, histone methytransferases/demethylases or MeCP2, and a significant decrease in BDNF variants. The decrease of the corresponding BDNF transcript leve ...
Gene Expression
... All levels of transcription and translation are involved: 1. DNA sequence will encode for specific regulation – promoters, exons/introns, etc 2. RNAs – will affect which genes complete the process to become proteins 3. Proteins – function as enzymes and machinery to activate or silence specific gene ...
... All levels of transcription and translation are involved: 1. DNA sequence will encode for specific regulation – promoters, exons/introns, etc 2. RNAs – will affect which genes complete the process to become proteins 3. Proteins – function as enzymes and machinery to activate or silence specific gene ...
Gene Expression
... All levels of transcription and translation are involved: 1. DNA sequence will encode for specific regulation – promoters, exons/introns, etc 2. RNAs – will affect which genes complete the process to become proteins 3. Proteins – function as enzymes and machinery to activate or silence specific gene ...
... All levels of transcription and translation are involved: 1. DNA sequence will encode for specific regulation – promoters, exons/introns, etc 2. RNAs – will affect which genes complete the process to become proteins 3. Proteins – function as enzymes and machinery to activate or silence specific gene ...
Chapter 12
... histones, thereby loosening their interaction with the DNA surrounding nucleosomes. This loosening activates the neighboring DNA by promoting expression of genes in this area. Histone deactylases remove the acetyl groups. – Histone methylation creates a binding site for heterochromatin proteins and ...
... histones, thereby loosening their interaction with the DNA surrounding nucleosomes. This loosening activates the neighboring DNA by promoting expression of genes in this area. Histone deactylases remove the acetyl groups. – Histone methylation creates a binding site for heterochromatin proteins and ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.