6.4 Gene Regulation - Ms. Franklin`s Classroom
... By regulating the transcription of genes into mRNA this can control the amount of proteins/enzymes produced for the metabolic pathway of tryptophan. ...
... By regulating the transcription of genes into mRNA this can control the amount of proteins/enzymes produced for the metabolic pathway of tryptophan. ...
Gene Regulation and Expression Notes
... Lactose Turns On the Operon When lactose is added to the medium, it diffuses into the cell and attaches to the lac repressor. ...
... Lactose Turns On the Operon When lactose is added to the medium, it diffuses into the cell and attaches to the lac repressor. ...
A Tour of the Cell - Ursuline High School
... Free in the cytosol….. they make proteins for use in the cytosol. When Attached to the Endoplasmic Reticulum….. they make proteins that are exported from the cell. When ...
... Free in the cytosol….. they make proteins for use in the cytosol. When Attached to the Endoplasmic Reticulum….. they make proteins that are exported from the cell. When ...
micrebiology - Microbiology
... DNA-binding proteins: correlations between structural differences, properties and functions Structural studies of histones and histone-like proteins have revealed a distinction into two classes depending on whether a typical fold, characteristic for eukaryotic histones (1,2), is present or not. In t ...
... DNA-binding proteins: correlations between structural differences, properties and functions Structural studies of histones and histone-like proteins have revealed a distinction into two classes depending on whether a typical fold, characteristic for eukaryotic histones (1,2), is present or not. In t ...
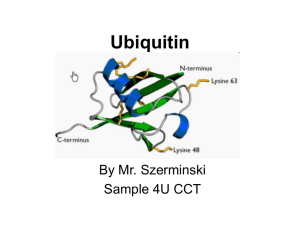
Ubiquitin
... Topics to be discussed • General info: - it is a regulatory protein that has been found in almost all tissues of eukaryotes - one of its functions: it directs protein recycling - can attach to proteins and label them for destruction. - discovery won the Nobel Prize for chemistry in 2004 ...
... Topics to be discussed • General info: - it is a regulatory protein that has been found in almost all tissues of eukaryotes - one of its functions: it directs protein recycling - can attach to proteins and label them for destruction. - discovery won the Nobel Prize for chemistry in 2004 ...
LKCMedicine Lecture Series by Asst Prof Chng Toh Hean
... transcription of new genes and synthesis of new proteins. The transport of synaptically-localised transcriptional regulators during neuronal activity provides a method of coupling synaptic activation with changes in the transcription. Asst Prof Chng’s lab is interested in studying the molecular basi ...
... transcription of new genes and synthesis of new proteins. The transport of synaptically-localised transcriptional regulators during neuronal activity provides a method of coupling synaptic activation with changes in the transcription. Asst Prof Chng’s lab is interested in studying the molecular basi ...
MCB Lecture 4 – Genes and Chromosomes
... histone proteins (wrapped 1.8 times), and the other 54 are considered linker DNA. o Nuclease – can digest the linker proteins but cannot digest the nucleotides wrapped around the histone Histone Proteins – H2A, H2B, H3, H4. Extremely well conserved. They have large amount of Arginine and Lysine (+Am ...
... histone proteins (wrapped 1.8 times), and the other 54 are considered linker DNA. o Nuclease – can digest the linker proteins but cannot digest the nucleotides wrapped around the histone Histone Proteins – H2A, H2B, H3, H4. Extremely well conserved. They have large amount of Arginine and Lysine (+Am ...
Advanced Techniques in Molecular Biology
... kidney) plays an essential role in generating energy for metabolism, synthesizing the brain neurotransmitter glutamate and maintaining acid-base balance in the kidney. Thus it makes sense that in case of decreasing the SNAT6( siRNA condition), the production of the Gls protein decreases, as there is ...
... kidney) plays an essential role in generating energy for metabolism, synthesizing the brain neurotransmitter glutamate and maintaining acid-base balance in the kidney. Thus it makes sense that in case of decreasing the SNAT6( siRNA condition), the production of the Gls protein decreases, as there is ...
p-5-wwu_wp3_talk-wagenknecht-kolkenbrock
... Department of Plant Biology and Biotechnology, Westphalian Wilhelm’s-University Münster, Schlossplatz 8, 48143 Münster, Germany ...
... Department of Plant Biology and Biotechnology, Westphalian Wilhelm’s-University Münster, Schlossplatz 8, 48143 Münster, Germany ...
PTM
... Mononucleotide addition is used to regulate the activity of some enzymes. Two different examples are found among the system that regulates Nitrogen utilization in E. coli: • Glutamine synthetase is adenylylated (i.e. AMP is added) at a specific tyrosine residue. The enzyme is inactive when it is ade ...
... Mononucleotide addition is used to regulate the activity of some enzymes. Two different examples are found among the system that regulates Nitrogen utilization in E. coli: • Glutamine synthetase is adenylylated (i.e. AMP is added) at a specific tyrosine residue. The enzyme is inactive when it is ade ...
Control of Gene Expression
... Individual genes are usually more methylated in cells in which the genes are not expressed. Once methylated, genes usually stay that way through successive cell divisions in an individual Removal of the extra methyl groups can turn on some of these genes Inheritance of traits transmitted by me ...
... Individual genes are usually more methylated in cells in which the genes are not expressed. Once methylated, genes usually stay that way through successive cell divisions in an individual Removal of the extra methyl groups can turn on some of these genes Inheritance of traits transmitted by me ...
Ch 18 Notes - FacStaff Home Page for CBU
... Eukaryotic Gene Expression: Regulated at many stages Differential Gene Expression Almost all the cells in an organism are genetically identical. Differences between cell types result from differential gene expression, the expression of different genes by cells with the same genome. Abnormalities in ...
... Eukaryotic Gene Expression: Regulated at many stages Differential Gene Expression Almost all the cells in an organism are genetically identical. Differences between cell types result from differential gene expression, the expression of different genes by cells with the same genome. Abnormalities in ...
ppt
... • Not all genes are active in all cells • Not all genes in a given cell are activated all the time • There MUST be some way to control when a gene is turned "on" or "off" • The activation of a gene results in transcription (mRNA made) which in turn results in the formation of a protein • Chromosomes ...
... • Not all genes are active in all cells • Not all genes in a given cell are activated all the time • There MUST be some way to control when a gene is turned "on" or "off" • The activation of a gene results in transcription (mRNA made) which in turn results in the formation of a protein • Chromosomes ...
The Organization and Control of Eukaryotic Genomes
... development. In all organisms, the expression of specific genes is most commonly regulated at the level of transcription by DNA-binding proteins. ...
... development. In all organisms, the expression of specific genes is most commonly regulated at the level of transcription by DNA-binding proteins. ...
Control of Gene Expression
... • Tryptophan fits in repressor blocks RNA polymerase. • Once out of tryp, repressor changes shape to allow promoter available to make more tryptophan turns transcription on. ...
... • Tryptophan fits in repressor blocks RNA polymerase. • Once out of tryp, repressor changes shape to allow promoter available to make more tryptophan turns transcription on. ...
File - EUREKA! Science
... Regulation of Transcription Genes Controlling Genes Some genes code for proteins that control the expression of other genes ...
... Regulation of Transcription Genes Controlling Genes Some genes code for proteins that control the expression of other genes ...
Control of Gene Expression
... • Methylation (the addition of –CH3) of DNA or histone proteins is associated with the control of gene expression. • Clusters of methylated cytosine nucleotides bind to a protein that prevents activators from binding to DNA. • Methylated histone proteins are associated with inactive regions of chrom ...
... • Methylation (the addition of –CH3) of DNA or histone proteins is associated with the control of gene expression. • Clusters of methylated cytosine nucleotides bind to a protein that prevents activators from binding to DNA. • Methylated histone proteins are associated with inactive regions of chrom ...
DNA-binding motifs
... • Methylation (the addition of –CH3) of DNA or histone proteins is associated with the control of gene expression. • Clusters of methylated cytosine nucleotides bind to a protein that prevents activators from binding to DNA. • Methylated histone proteins are associated with inactive regions of chrom ...
... • Methylation (the addition of –CH3) of DNA or histone proteins is associated with the control of gene expression. • Clusters of methylated cytosine nucleotides bind to a protein that prevents activators from binding to DNA. • Methylated histone proteins are associated with inactive regions of chrom ...
Regulation of Gene Expression
... pathway are often found in the same operon – All under the control of the same promoter region – Thus these genes are transcribed all together into one continuous mRNA strand: polycistronic mRNA • Proteins are then synthesized from that mRNA ...
... pathway are often found in the same operon – All under the control of the same promoter region – Thus these genes are transcribed all together into one continuous mRNA strand: polycistronic mRNA • Proteins are then synthesized from that mRNA ...
Control of Gene Expression Control of Gene Expression Regulatory
... • Methylation (the addition of –CH3 to DNA or histone proteins) is associated with the control of gene expression. • Clusters of methylated cytosine nucleotides bind to a protein that prevents activators from binding to DNA. • Methylated histone proteins are associated with inactive regions of chrom ...
... • Methylation (the addition of –CH3 to DNA or histone proteins) is associated with the control of gene expression. • Clusters of methylated cytosine nucleotides bind to a protein that prevents activators from binding to DNA. • Methylated histone proteins are associated with inactive regions of chrom ...
Control of Gene Expression 3 - Dr. Kordula
... glutaminerich domain of Sp1 and the prolinerich domain of CTF1. See Figure 3 below. C. Enhancers These DNA elements, located 200 bp to 50 kb from the +1, affect gene expression despite their distance from the promoter region. Enhancers can be located upstream, downstream, or perhaps in an int ...
... glutaminerich domain of Sp1 and the prolinerich domain of CTF1. See Figure 3 below. C. Enhancers These DNA elements, located 200 bp to 50 kb from the +1, affect gene expression despite their distance from the promoter region. Enhancers can be located upstream, downstream, or perhaps in an int ...
Bio200 Au13 Lec19 10-29 Slides
... • Eukaryotic mRNA is heavily processed before being used • A 5’ protein cap and a 3’ poly-A tail are added to give stability • Non-coding introns are spliced out of the mRNA by the spliceosome ...
... • Eukaryotic mRNA is heavily processed before being used • A 5’ protein cap and a 3’ poly-A tail are added to give stability • Non-coding introns are spliced out of the mRNA by the spliceosome ...
Slide 1 - Ommbid.com
... The left half of the figure represents the state of several proteins and mRNAs under normal conditions, the right half shows the activation of the UPR in response to an overload of the ER with unfolded or malfolded proteins. Under normal conditions the three effector proteins of the UPR (PERK, IRE1 ...
... The left half of the figure represents the state of several proteins and mRNAs under normal conditions, the right half shows the activation of the UPR in response to an overload of the ER with unfolded or malfolded proteins. Under normal conditions the three effector proteins of the UPR (PERK, IRE1 ...
Chapter 19 - mrswehri.com
... chromosome. However, unlike the genes in the operons of prokaryotes, each of the eukaryotic genes have their own promoter and is individually transcribed. It is thought that the coordinate regulation of genes clustered in eukaryotic cells involves changes in chromatin structure that makes the entire ...
... chromosome. However, unlike the genes in the operons of prokaryotes, each of the eukaryotic genes have their own promoter and is individually transcribed. It is thought that the coordinate regulation of genes clustered in eukaryotic cells involves changes in chromatin structure that makes the entire ...
Gene Expression
... gene to turn on (promote) or off (repress). Eukaryotes have multiple switches. – Induction- If proteins from neighboring cells are present, gene may turn on (ex: retina) – Hormones and other molecules may attach to enhancer sequence to turn on genes. ...
... gene to turn on (promote) or off (repress). Eukaryotes have multiple switches. – Induction- If proteins from neighboring cells are present, gene may turn on (ex: retina) – Hormones and other molecules may attach to enhancer sequence to turn on genes. ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.