Characterization of Deletions in the LDL Receptor Gene in Patients
... basis of clinical criteria alone; many hypercholesterolemic patients can only be described as "probable FH" or even "possible FH" unless they have tendon xanthomas or a well-defined family history of hypercholesterolemia and premature coronary heart disease. For this reason a diagnostic test based o ...
... basis of clinical criteria alone; many hypercholesterolemic patients can only be described as "probable FH" or even "possible FH" unless they have tendon xanthomas or a well-defined family history of hypercholesterolemia and premature coronary heart disease. For this reason a diagnostic test based o ...
Supplementary Information (doc 42K)
... dbGap dataset and quality control Genome-wide SNP data for 7 018 individuals comprising 2 339 trios in which each child was affected with any type of NSOFC (CL/P or CPO), were downloaded from dbGaP (Accession number: phs000094.v1.p1)1. Available genotype data included a total of 1 387 466 SNPs, comp ...
... dbGap dataset and quality control Genome-wide SNP data for 7 018 individuals comprising 2 339 trios in which each child was affected with any type of NSOFC (CL/P or CPO), were downloaded from dbGaP (Accession number: phs000094.v1.p1)1. Available genotype data included a total of 1 387 466 SNPs, comp ...
Cloning of genes from genomic DNA: Part 3
... Continuing from our isolation of genomic DNA and PCR amplification of either the evenskipped gene or the twist gene, we will now move on to the third step in the cloning procedure. We will use restriction enzymes to cleave off the ends of the PCR products. The oligonucleotide primers used in the PCR ...
... Continuing from our isolation of genomic DNA and PCR amplification of either the evenskipped gene or the twist gene, we will now move on to the third step in the cloning procedure. We will use restriction enzymes to cleave off the ends of the PCR products. The oligonucleotide primers used in the PCR ...
Curriculum and Training Specialist Bio
... DNA profiling is the use of molecular genetic methods to determine the exact genotype of a DNA sample in a way that can basically distinguish one human being from another The unique genotype of each sample is called a DNA profile. ...
... DNA profiling is the use of molecular genetic methods to determine the exact genotype of a DNA sample in a way that can basically distinguish one human being from another The unique genotype of each sample is called a DNA profile. ...
Document
... DNA profiling is the use of molecular genetic methods to determine the exact genotype of a DNA sample in a way that can basically distinguish one human being from another The unique genotype of each sample is called a DNA profile. ...
... DNA profiling is the use of molecular genetic methods to determine the exact genotype of a DNA sample in a way that can basically distinguish one human being from another The unique genotype of each sample is called a DNA profile. ...
Title: Gene Interactions in Corn. Introduction. The phenotype of an
... grammar and/or poor spelling. Your report should be stapled once in the top left corner. Binders or paper clips are not acceptable. Your report should be in the following format: Name and date. Title of experiment. Introduction to experiment. Description of methods and materials used. The above info ...
... grammar and/or poor spelling. Your report should be stapled once in the top left corner. Binders or paper clips are not acceptable. Your report should be in the following format: Name and date. Title of experiment. Introduction to experiment. Description of methods and materials used. The above info ...
Infectious Pancreatic Necrosis Virus
... amplification, forward and reverse primers hybridize to the IPNV cDNA. A fluorogenic probe is included in the same reaction mixture which consists of a DNA probe labeled with a 5`-dye and a 3`-quencher. During PCR amplification, the probe is cleaved and the reporter dye and quencher are separated. T ...
... amplification, forward and reverse primers hybridize to the IPNV cDNA. A fluorogenic probe is included in the same reaction mixture which consists of a DNA probe labeled with a 5`-dye and a 3`-quencher. During PCR amplification, the probe is cleaved and the reporter dye and quencher are separated. T ...
Gene quantification using real-time quantitative PCR
... DNA copy number measurements are important in determining the extent of genomic imbalance that underlies most malignancies. There are numerous techniques available for measuring DNA copy number in tumors; each method has specific advantages and disadvantages. Chromosomal CGH can detect imbalances ac ...
... DNA copy number measurements are important in determining the extent of genomic imbalance that underlies most malignancies. There are numerous techniques available for measuring DNA copy number in tumors; each method has specific advantages and disadvantages. Chromosomal CGH can detect imbalances ac ...
A familial inverted duplication/deletion of 2p25.1–25.3
... paradigmatic example of this type of rearrangement.6,7 Here, we report a family where two children and their affected father carry the same dup(2)(p25.3p25.1) detected by karyotype analysis. Molecular characterization by FISH and array-CGH (aCGH) analyses demonstrated that the rearranged chromosome ...
... paradigmatic example of this type of rearrangement.6,7 Here, we report a family where two children and their affected father carry the same dup(2)(p25.3p25.1) detected by karyotype analysis. Molecular characterization by FISH and array-CGH (aCGH) analyses demonstrated that the rearranged chromosome ...
Gabriele Marras
... Genome wide association studies (GWAS) are used to identify regions of the genome associated with the phenotypes. However, standard GWAS only identifies individual SNPs associated with traits and not directly regions of the genome or genes. Additionally, standard ...
... Genome wide association studies (GWAS) are used to identify regions of the genome associated with the phenotypes. However, standard GWAS only identifies individual SNPs associated with traits and not directly regions of the genome or genes. Additionally, standard ...
Novel cryptic chromosomal rearrangements in childhood acute
... progress in identifying and treating resistant subtypes of the disease.1,2 The classification of ALL into therapeutically relevant risk categories relies on both clinical parameters, including age, leukocyte count, immunophenotype, central nervous system (CNS) involvement, as well as on the blast ce ...
... progress in identifying and treating resistant subtypes of the disease.1,2 The classification of ALL into therapeutically relevant risk categories relies on both clinical parameters, including age, leukocyte count, immunophenotype, central nervous system (CNS) involvement, as well as on the blast ce ...
Characterization of two rice DNA methyltransferases
... The full-length sequence of OsMET1-2 was obtained by comparison of OsMET1-1 with the rice genome database available at http://www.tigr.org. Nucleotide sequence comparison using AlignX indicated that OsMET1-1 had 99% identity to a region in clone AC093713, located on chromosome 3 and showed 68.5% id ...
... The full-length sequence of OsMET1-2 was obtained by comparison of OsMET1-1 with the rice genome database available at http://www.tigr.org. Nucleotide sequence comparison using AlignX indicated that OsMET1-1 had 99% identity to a region in clone AC093713, located on chromosome 3 and showed 68.5% id ...
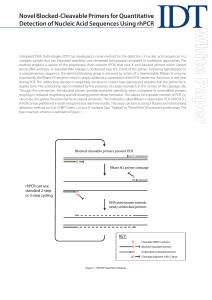
Novel Blocked-Cleavable Primers for Quantitative Detection of
... Integrated DNA Technologies (IDT) has developed a novel method for the detection of nucleic acid sequences in a complex sample that has improved specificity and decreased background compared to traditional approaches. The method employs a variant of the polymerase chain reaction (PCR) that uses 3’-e ...
... Integrated DNA Technologies (IDT) has developed a novel method for the detection of nucleic acid sequences in a complex sample that has improved specificity and decreased background compared to traditional approaches. The method employs a variant of the polymerase chain reaction (PCR) that uses 3’-e ...
The HELCOM Rec. 15/5 HELCOM Network of Marine Protected Areas
... • The effect of the legal protection of HELCOM MPAs is unclear • The 10% MPA target is reached for the Baltic Sea but not for sub-regions • The HELCOM MPA Guidelines should be fully compatible with the IUCN classification of MPAs and the selection criteria for EBSAs • MPAs should be transboundary wh ...
... • The effect of the legal protection of HELCOM MPAs is unclear • The 10% MPA target is reached for the Baltic Sea but not for sub-regions • The HELCOM MPA Guidelines should be fully compatible with the IUCN classification of MPAs and the selection criteria for EBSAs • MPAs should be transboundary wh ...
マニュアル Megaprime DNA Labelling System
... subsequently removed as monophosphates. Very small amount of input DNA can be labelled, enabling very high specific activity DNA probes to be produced with relatively small quantities of added nucleotides. These radioactive labelled fragments can then be used as sensitive hybridization probes for a ...
... subsequently removed as monophosphates. Very small amount of input DNA can be labelled, enabling very high specific activity DNA probes to be produced with relatively small quantities of added nucleotides. These radioactive labelled fragments can then be used as sensitive hybridization probes for a ...
Genome Biology and
... – Gene identification is almost trivial in bacteria and yeasts • Genes are readily recognized by ab initio analysis as ORFs coding for >100 amino acids (no introns) – Smaller ORFs and overlapping genes are missed – Gene identification is relatively straightforward in small genomes, such as worm, pla ...
... – Gene identification is almost trivial in bacteria and yeasts • Genes are readily recognized by ab initio analysis as ORFs coding for >100 amino acids (no introns) – Smaller ORFs and overlapping genes are missed – Gene identification is relatively straightforward in small genomes, such as worm, pla ...
RAPID DNA HYBRIDIZATION REACTIONS USING
... preconcentrating reactant species in simple, straight microchannels. We use peak mode ITP to focus species by up to 50,000 fold into order 10 micron zone lengths; yielding order 10 picoliter volumes which incubate reactions. We have developed and experimentally validated the detailed reaction kineti ...
... preconcentrating reactant species in simple, straight microchannels. We use peak mode ITP to focus species by up to 50,000 fold into order 10 micron zone lengths; yielding order 10 picoliter volumes which incubate reactions. We have developed and experimentally validated the detailed reaction kineti ...
Use case flow for use case: 2
... 2. The System uses semantic matching to find a database which can be used to find the chromosome location for a gene (the gene database). 3. The System takes the gene expressed and queries the gene database to find what chromosome the expressed gene is on. 4. The gene database returns the chromosom ...
... 2. The System uses semantic matching to find a database which can be used to find the chromosome location for a gene (the gene database). 3. The System takes the gene expressed and queries the gene database to find what chromosome the expressed gene is on. 4. The gene database returns the chromosom ...
PAG 2012 - Illumina
... Creation of a genomic resource and quantitative trait loci detection in Pacific Whiteleg Shrimp Litopenaeus vannamei for selective breeding applications ...
... Creation of a genomic resource and quantitative trait loci detection in Pacific Whiteleg Shrimp Litopenaeus vannamei for selective breeding applications ...
Molecular Inversion Probe

Molecular Inversion Probe (MIP) belongs to the class of Capture by Circularization molecular techniques for performing genomic partitioning, a process through which one captures and enriches specific regions of the genome. Probes used in this technique are single stranded DNA molecules and, similar to other genomic partitioning techniques, contain sequences that are complementary to the target in the genome; these probes hybridize to and capture the genomic target. MIP stands unique from other genomic partitioning strategies in that MIP probes share the common design of two genomic target complementary segments separated by a linker region. With this design, when the probe hybridizes to the target, it undergoes an inversion in configuration (as suggested by the name of the technique) and circularizes. Specifically, the two target complementary regions at the 5’ and 3’ ends of the probe become adjacent to one another while the internal linker region forms a free hanging loop. The technology has been used extensively in the HapMap project for large-scale SNP genotyping as well as for studying gene copy alterationsand characteristics of specific genomic loci to identify biomarkers for different diseases such as cancer. Key strengths of the MIP technology include its high specificity to the target and its scalability for high-throughput, multiplexed analyses where tens of thousands of genomic loci are assayed simultaneously.