Screening of RYR1 genotypes in swine population by a rapid and
... make sequence discrimination easier. However, the primers used for HRM must generate short amplicons. According to the manufacturer's recommendation the best results can be obtained with amplicons up to 300 bp (www. roche-applied-science.com) because reducing amplicon size increases the difference i ...
... make sequence discrimination easier. However, the primers used for HRM must generate short amplicons. According to the manufacturer's recommendation the best results can be obtained with amplicons up to 300 bp (www. roche-applied-science.com) because reducing amplicon size increases the difference i ...
Bchem 4200 Part13 - U of L Class Index
... Sliding is the most important process in target site location. → Leaving the target side might also involve sliding etc. Sliding accelerates target site location: → under optimum conditions it allows for scanning of ~106 bases per binding event. → but it’s a random walk →the effective sliding distan ...
... Sliding is the most important process in target site location. → Leaving the target side might also involve sliding etc. Sliding accelerates target site location: → under optimum conditions it allows for scanning of ~106 bases per binding event. → but it’s a random walk →the effective sliding distan ...
View Poster - Technology Networks
... unicellular organisms. This has often led to the speculation that miRNAs have co-evolved with multicellularity in plants and animals. In contrast, we found that miRNA precursors are present in the single-celled green alga Chlamydomonas reinhardtii and that they give rise to mature miRNAs that regula ...
... unicellular organisms. This has often led to the speculation that miRNAs have co-evolved with multicellularity in plants and animals. In contrast, we found that miRNA precursors are present in the single-celled green alga Chlamydomonas reinhardtii and that they give rise to mature miRNAs that regula ...
Title: FISH analysis comparing the gene composition of the Onager
... The onager [E. hemionus onager, EHO] and the domestic horse [E. caballus, ECA] have evolved over the course of 3.7 million years. The closely related EHO and ECA have diploid chromosome numbers of 2n=56 and 2n=64, respectively. Comparative gene mapping was done by FISH [fluorescent in-situ hybridiza ...
... The onager [E. hemionus onager, EHO] and the domestic horse [E. caballus, ECA] have evolved over the course of 3.7 million years. The closely related EHO and ECA have diploid chromosome numbers of 2n=56 and 2n=64, respectively. Comparative gene mapping was done by FISH [fluorescent in-situ hybridiza ...
Introduction to Preprocessing: RMA (Robust Multi
... (“the approximately right scale”) By most reasonable metrics, RMA performs well (at least well enough to justify using it without losing ...
... (“the approximately right scale”) By most reasonable metrics, RMA performs well (at least well enough to justify using it without losing ...
Poster - Pacific Biosciences
... As a cost-effective alternative to whole genome human sequencing, targeted sequencing of specific regions, such as exomes or panels of relevant genes, has become increasingly common. These methods typically include direct PCR amplification of the genomic DNA of interest, or the capture of these targ ...
... As a cost-effective alternative to whole genome human sequencing, targeted sequencing of specific regions, such as exomes or panels of relevant genes, has become increasingly common. These methods typically include direct PCR amplification of the genomic DNA of interest, or the capture of these targ ...
Sequencing Requirements Requirements for DNA sequencing: Only
... an agarose gel or a NanoDrop spectrophotometer. a. Purified DNA should run as a single band on an agarose gel b. For NanoDrop, the A260/A280 ratio should be 1.8. The NanoDrop only requires 1-2 uL of sample. 7. Do you supply primers? -The GCF DOES NOT supply primers. Please submit ONE primer (fwd or ...
... an agarose gel or a NanoDrop spectrophotometer. a. Purified DNA should run as a single band on an agarose gel b. For NanoDrop, the A260/A280 ratio should be 1.8. The NanoDrop only requires 1-2 uL of sample. 7. Do you supply primers? -The GCF DOES NOT supply primers. Please submit ONE primer (fwd or ...
Illumina Infinium HumanMethylation450 BeadChip Data
... workflow through Partek® Genomics Suite™ software where you can perform analysis in a single, user-friendly application. Start by importing native .idat files directly into Partek Genomics Suite and applying innovative techniques for signal processing, background correction, and normalization. Follo ...
... workflow through Partek® Genomics Suite™ software where you can perform analysis in a single, user-friendly application. Start by importing native .idat files directly into Partek Genomics Suite and applying innovative techniques for signal processing, background correction, and normalization. Follo ...
Question 1 _____/ 30 points Question 2 _____/ 20 points Question 3
... protein is only expressed when there are very low nutrients to reduce protein synthesis and conserve resources. The location of the RP28 gene and its two neighbors is illustrated below (the arrows indicate the direction of transcription). ...
... protein is only expressed when there are very low nutrients to reduce protein synthesis and conserve resources. The location of the RP28 gene and its two neighbors is illustrated below (the arrows indicate the direction of transcription). ...
In situ hybridization
... 2. Incubate with the blocking reagent (10x Roche Blocking solution, in Buffer 1) for 30 minutes. 3. Incubate anti-DIG antibody-AP conjugate in Buffer 1 + blocking serum. (1 hour at room temperature to overnight at 4oC 1/5000). 4. Remove unbound antibody by washing in Buffer 1 for 10 minutes, 3x. 5. ...
... 2. Incubate with the blocking reagent (10x Roche Blocking solution, in Buffer 1) for 30 minutes. 3. Incubate anti-DIG antibody-AP conjugate in Buffer 1 + blocking serum. (1 hour at room temperature to overnight at 4oC 1/5000). 4. Remove unbound antibody by washing in Buffer 1 for 10 minutes, 3x. 5. ...
biotechnology
... • Because the expensive disposal of radioactive waste, nonradioactive probes have been developed. The vitamin biotin, can be chemically coupled to the nucleotides used to synthesize the probe. Biotin binds very tenaciously to avidin-a readily available protein contained in chicken egg whites • Avidi ...
... • Because the expensive disposal of radioactive waste, nonradioactive probes have been developed. The vitamin biotin, can be chemically coupled to the nucleotides used to synthesize the probe. Biotin binds very tenaciously to avidin-a readily available protein contained in chicken egg whites • Avidi ...
Genotyping errors - Proceedings of the Royal Society B
... higher (ef = 0.59) than error rate per phenotype (i.e., per locus x sample) but only slightly so. Overall, error rates are low and likely to have a negligible effect on results. For example, for a fixed (monomorphic) fragment to become erroneously polymorphic under the 5%-criterion, it would necess ...
... higher (ef = 0.59) than error rate per phenotype (i.e., per locus x sample) but only slightly so. Overall, error rates are low and likely to have a negligible effect on results. For example, for a fixed (monomorphic) fragment to become erroneously polymorphic under the 5%-criterion, it would necess ...
A computational platform for whole genome association analysis
... Test for correlation between unlinked loci Test for difference in correlation between loci, in cases and controls Increases multiple testing: e.g. 500,000 SNPs leads to 124,999,750,000 possible pairs of SNPs ...
... Test for correlation between unlinked loci Test for difference in correlation between loci, in cases and controls Increases multiple testing: e.g. 500,000 SNPs leads to 124,999,750,000 possible pairs of SNPs ...
APOC3 rs2854116 single nucleotide polymorphism
... polymorphism have been widely developed because of its significant association with elevated plasma lipid concentrations, risk of coronary heart disease and metabolic syndromein multiple populations. The study aimed to apply the restriction fragment length polymorphism (RFLP) method for identifying ...
... polymorphism have been widely developed because of its significant association with elevated plasma lipid concentrations, risk of coronary heart disease and metabolic syndromein multiple populations. The study aimed to apply the restriction fragment length polymorphism (RFLP) method for identifying ...
Bioinformatics Variant Analysis
... adapter sequence set (Illumina TruSeq adapters) with two or less mismatches and an alignment quality of at least 12. To remove noise introduced by sequencing errors, reads are filtered and clipped by quality. By default, the reads are filtered using a phred score of Q22 as a minimum threshold. Bases ...
... adapter sequence set (Illumina TruSeq adapters) with two or less mismatches and an alignment quality of at least 12. To remove noise introduced by sequencing errors, reads are filtered and clipped by quality. By default, the reads are filtered using a phred score of Q22 as a minimum threshold. Bases ...
Title Actual distribution of bacteriocytes in the trophosome of a beard
... a special tissue called the trophosome in the latter half of their extremely slender bodies. In the trophosome, specific cells, named bacteriocytes by Schwemmler (1980), are present. ...
... a special tissue called the trophosome in the latter half of their extremely slender bodies. In the trophosome, specific cells, named bacteriocytes by Schwemmler (1980), are present. ...
Case Study Learning via Simulations of Molecular Biology Techniques
... malignancy. Several BRCA1 mutations, including point mutations, deletions, and insertions, have been identified that may contribute to loss of tumor suppressor function. These mutations can be identified by amplifying portions of the BRCA1 gene by PCR and then using RFLP analysis, direct sequencing, ...
... malignancy. Several BRCA1 mutations, including point mutations, deletions, and insertions, have been identified that may contribute to loss of tumor suppressor function. These mutations can be identified by amplifying portions of the BRCA1 gene by PCR and then using RFLP analysis, direct sequencing, ...
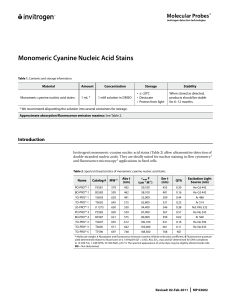
Monomeric Cyanine Nucleic Acid Stains
... Email: [email protected] Technical Services: [email protected] ...
... Email: [email protected] Technical Services: [email protected] ...
Biostat Jhsph Edu Hji Courses Genomics Sequencing Ppt
... (d) Changes in the fluorophore cross-talk cause misinterpretation of the received signal (teal arrows; d). For simplicity, the noise factors are presented separately from each other. ...
... (d) Changes in the fluorophore cross-talk cause misinterpretation of the received signal (teal arrows; d). For simplicity, the noise factors are presented separately from each other. ...
PowerPoint-presentatie - the biopsychology research group
... In conclusion: The Perlegen SNP-microarrays used in the GAIN projects contain copy number information. The CNV analysis can potentially identify new candidate genes and new risk alleles for ADHD. ...
... In conclusion: The Perlegen SNP-microarrays used in the GAIN projects contain copy number information. The CNV analysis can potentially identify new candidate genes and new risk alleles for ADHD. ...
Lab #1: Alu Lab, Part 1
... Part B: Amplifying the Target Sequence by PCR With our DNA successfully isolated from our cheek cells, we’re ready to begin the PCR. We’ll set-up this reaction today and allow it to run overnight. We’ll analyze our samples using gel electrophoresis during our next lab session. A). Set up group contr ...
... Part B: Amplifying the Target Sequence by PCR With our DNA successfully isolated from our cheek cells, we’re ready to begin the PCR. We’ll set-up this reaction today and allow it to run overnight. We’ll analyze our samples using gel electrophoresis during our next lab session. A). Set up group contr ...
PLEIOTROPIC MULTI-TRAIT GENOME
... dataset, management group, flock, date of observation, drop year, sex, birth type, and rear type as fixed effects. The FA traits were corrected for intramuscular fat content. The individual trait results were combined using the meta-analysis described by Bolormaa et al. (2014). To avoid identifying ...
... dataset, management group, flock, date of observation, drop year, sex, birth type, and rear type as fixed effects. The FA traits were corrected for intramuscular fat content. The individual trait results were combined using the meta-analysis described by Bolormaa et al. (2014). To avoid identifying ...
Molecular Inversion Probe

Molecular Inversion Probe (MIP) belongs to the class of Capture by Circularization molecular techniques for performing genomic partitioning, a process through which one captures and enriches specific regions of the genome. Probes used in this technique are single stranded DNA molecules and, similar to other genomic partitioning techniques, contain sequences that are complementary to the target in the genome; these probes hybridize to and capture the genomic target. MIP stands unique from other genomic partitioning strategies in that MIP probes share the common design of two genomic target complementary segments separated by a linker region. With this design, when the probe hybridizes to the target, it undergoes an inversion in configuration (as suggested by the name of the technique) and circularizes. Specifically, the two target complementary regions at the 5’ and 3’ ends of the probe become adjacent to one another while the internal linker region forms a free hanging loop. The technology has been used extensively in the HapMap project for large-scale SNP genotyping as well as for studying gene copy alterationsand characteristics of specific genomic loci to identify biomarkers for different diseases such as cancer. Key strengths of the MIP technology include its high specificity to the target and its scalability for high-throughput, multiplexed analyses where tens of thousands of genomic loci are assayed simultaneously.