Study questions - Pre-lab
... a. Predict whether or not you will exhibit the PTC taster phenotype. b. If you are a taster of PTC, what are your possible genotypes at the TAS2R38 locus? PAV/AVI or PAV/PAV (T/t or T/T) c. In which ways can single nucleotide polymorphisms (SNPs) affect the function of a gene? Non-sense mutations (t ...
... a. Predict whether or not you will exhibit the PTC taster phenotype. b. If you are a taster of PTC, what are your possible genotypes at the TAS2R38 locus? PAV/AVI or PAV/PAV (T/t or T/T) c. In which ways can single nucleotide polymorphisms (SNPs) affect the function of a gene? Non-sense mutations (t ...
Presentazione di PowerPoint
... – Building datasets of several strains of the same organism to investigate intra-species evolution. ...
... – Building datasets of several strains of the same organism to investigate intra-species evolution. ...
High-resolution mapping of the leaf rust disease resistance gene Lr1
... Keller 1999). Comparison of the gene composition at orthologous Lrk loci in wheat, barley and rice showed that the high density of genes is conserved at syntenic loci of large and small grass genomes (Feuillet and Keller 1999). Therefore, gene-rich regions in the wheat genome may be amenable to mole ...
... Keller 1999). Comparison of the gene composition at orthologous Lrk loci in wheat, barley and rice showed that the high density of genes is conserved at syntenic loci of large and small grass genomes (Feuillet and Keller 1999). Therefore, gene-rich regions in the wheat genome may be amenable to mole ...
Phasing Analysis Service for Whole Human Genome Sequencing
... By capturing gene information from homologous chromosomes, phasing technology eliminates the traditional reliance on haplotype inference based solely on statistical information, which can be subject to error. Other traditional phasing methods include trio studies, which compare maternal and paternal ...
... By capturing gene information from homologous chromosomes, phasing technology eliminates the traditional reliance on haplotype inference based solely on statistical information, which can be subject to error. Other traditional phasing methods include trio studies, which compare maternal and paternal ...
6 Possible Alleles
... • Set up PCR reactions • Electrophorese PCR products • Analysis and interpretation of results ...
... • Set up PCR reactions • Electrophorese PCR products • Analysis and interpretation of results ...
Single-Nucleotide Polymorphism Mapping
... SNPs have two advantages over conventional marker mutations. First, unlike conventional visible markers, SNPs in general have no phenotype, allowing a mutation of interest to be scored in a neutral phenotypic background. As a result, many markers can be assayed simultaneously, without worrying about ...
... SNPs have two advantages over conventional marker mutations. First, unlike conventional visible markers, SNPs in general have no phenotype, allowing a mutation of interest to be scored in a neutral phenotypic background. As a result, many markers can be assayed simultaneously, without worrying about ...
C2005/F2401 Key to Exam #3
... determines which strand is transcribed. If the enzyme Z gene can be transcribed (successfully) in either orientation, then the fragment itself must contain the promoter of the Z gene as well as the coding region for the enzyme. Therefore, the gene is always in the same orientation to its promoter no ...
... determines which strand is transcribed. If the enzyme Z gene can be transcribed (successfully) in either orientation, then the fragment itself must contain the promoter of the Z gene as well as the coding region for the enzyme. Therefore, the gene is always in the same orientation to its promoter no ...
synthesis Gene Cluster of Streptomyces clavuligerus
... for a argCJBDFRGH cluster have been deposited in EMBL database (Accession numbers Z85982 and L78811, respectively). Arginine biosynthesis genes in Bacillus subtilis are organized in two operons, one is argCJBDcarAB-argF for early steps in the arginine pathway, including those involved in carbamoyl p ...
... for a argCJBDFRGH cluster have been deposited in EMBL database (Accession numbers Z85982 and L78811, respectively). Arginine biosynthesis genes in Bacillus subtilis are organized in two operons, one is argCJBDcarAB-argF for early steps in the arginine pathway, including those involved in carbamoyl p ...
Lecture_8
... Euler Theorem: Extension • Theorem: A connected graph has an Eulerian path if and only if it contains at most two semi-balanced vertices (one has one more outgoing edge and the other has one more incoming edge) and all other vertices are balanced. ...
... Euler Theorem: Extension • Theorem: A connected graph has an Eulerian path if and only if it contains at most two semi-balanced vertices (one has one more outgoing edge and the other has one more incoming edge) and all other vertices are balanced. ...
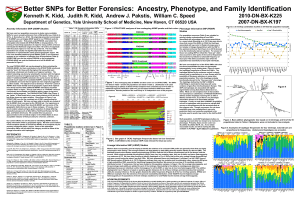
Better SNPs for Better Forensics
... Markers that are especially good at helping to identify the relatives of an unknown DNA profile are generally those that are highly polymorphic while having a low enough mutation rate that identity by state (IBS) generally implies identity by descent (IBD). The standard forensic short tandem repeat ...
... Markers that are especially good at helping to identify the relatives of an unknown DNA profile are generally those that are highly polymorphic while having a low enough mutation rate that identity by state (IBS) generally implies identity by descent (IBD). The standard forensic short tandem repeat ...
Query Results
... Download Sequence: If the tag matches an ORF (as it is in this example), the transcript sequence is given. The 5’ and 3’ UTRs, the start and stop codons, and the tag sequence are all highlighted. If the tag matches an intergenic region, the 500 flanking nucleotides upstream and downstrem the tag ar ...
... Download Sequence: If the tag matches an ORF (as it is in this example), the transcript sequence is given. The 5’ and 3’ UTRs, the start and stop codons, and the tag sequence are all highlighted. If the tag matches an intergenic region, the 500 flanking nucleotides upstream and downstrem the tag ar ...
FISH – Technical Considerations - San Antonio Society of Pathologists
... independently identifiable target on a solid stained chromosome (e.g., structural rearrangements, trisomy, etc.); other methods that localize the probe at a level of resolution appropriate to the intended chromosome target. ...
... independently identifiable target on a solid stained chromosome (e.g., structural rearrangements, trisomy, etc.); other methods that localize the probe at a level of resolution appropriate to the intended chromosome target. ...
ParSNP Hash
... • Implement NTFreq module to calculate nucleotide frequencies for each population and combined population • Implement TajimasD module to calculate Tajima’s D • Implement GO module to annotate identified SNPs ...
... • Implement NTFreq module to calculate nucleotide frequencies for each population and combined population • Implement TajimasD module to calculate Tajima’s D • Implement GO module to annotate identified SNPs ...
Cepheid Xpert Carba-R
... Each cartridge includes the following internal controls: Sample Processing Control (SPC - dry spore cake of noninfectious spores) ensures samples are adequately extracted and detects specimen-associated PCR-inhibition; SPC must be POSITIVE in NEGATIVE samples, but can be negative or positive in POSI ...
... Each cartridge includes the following internal controls: Sample Processing Control (SPC - dry spore cake of noninfectious spores) ensures samples are adequately extracted and detects specimen-associated PCR-inhibition; SPC must be POSITIVE in NEGATIVE samples, but can be negative or positive in POSI ...
Document
... DNA profiling is the use of molecular genetic methods to determine the exact genotype of a DNA sample in a way that can basically distinguish one human being from another The unique genotype of each sample is called a DNA profile. ...
... DNA profiling is the use of molecular genetic methods to determine the exact genotype of a DNA sample in a way that can basically distinguish one human being from another The unique genotype of each sample is called a DNA profile. ...
DNA Duplication Associated with Charcot-Marie-Tooth Disease Type 1A. Lupski, et al., 1991 Cell, Vol. 66, 219-232, July 26, 1991,
... Associated with CMTIA We screened CMTl A-linked 17p DNA probes for the presence of simple sequence repeats such as (GT),, which are known to be highly polymorphic and can be rapidly analyzed by the polymerase chain reaction (PCR) (Weber and May, 1989; Litt and Luty, 1989). (CT), sequences were ident ...
... Associated with CMTIA We screened CMTl A-linked 17p DNA probes for the presence of simple sequence repeats such as (GT),, which are known to be highly polymorphic and can be rapidly analyzed by the polymerase chain reaction (PCR) (Weber and May, 1989; Litt and Luty, 1989). (CT), sequences were ident ...
A general method for gene isolation in tagging approaches
... fragments larger than 500 base pairs is difficult since artefacts of uncut DNA might be displayed, or the fragments might not always be amplified thoroughly by the Taq polymerase. Premature termination of DNA synthesis leads to an accumulation of small-sized bands (less than 80 bp) which also makes ...
... fragments larger than 500 base pairs is difficult since artefacts of uncut DNA might be displayed, or the fragments might not always be amplified thoroughly by the Taq polymerase. Premature termination of DNA synthesis leads to an accumulation of small-sized bands (less than 80 bp) which also makes ...
Analysis and Characterization of Nucleic Acids and Proteins
... Under nonstandard conditions, some restriction enzymes will bind to and cut sequences other than their defined recognition sequence. This altered specificity is called star activity. The propensity for star activity varies among enzymes. Thus, the nature and degree of star activity depends on ...
... Under nonstandard conditions, some restriction enzymes will bind to and cut sequences other than their defined recognition sequence. This altered specificity is called star activity. The propensity for star activity varies among enzymes. Thus, the nature and degree of star activity depends on ...
Document
... AS occurs in 59% of human genes (Graveley, 2001) AS expands protein diversity (generates from a single gene multiple transcripts) AS is tissue-specific (Graveley, 2001) AS is related to human diseases ...
... AS occurs in 59% of human genes (Graveley, 2001) AS expands protein diversity (generates from a single gene multiple transcripts) AS is tissue-specific (Graveley, 2001) AS is related to human diseases ...
Genetics and Genomics of Core Short Tandem Repeat Loci
... (1) STR markers : important tools for human identity testing continue to be widely used for many years : their high degree of variability, ease of use in multiplex amplification formats and implementation in National DNA Databases (2) A uniform set of core STR loci : the capability for national and ...
... (1) STR markers : important tools for human identity testing continue to be widely used for many years : their high degree of variability, ease of use in multiplex amplification formats and implementation in National DNA Databases (2) A uniform set of core STR loci : the capability for national and ...
Department of Biomedical Informatics
... • Array-based technology • Massive sequencing • Genome wide association study (GWAS) • SNP array exome sequencing genome resequencing • Expression quantitative trait loci (eQTL) • Allelic specific ********ion Department of Biomedical Informatics ...
... • Array-based technology • Massive sequencing • Genome wide association study (GWAS) • SNP array exome sequencing genome resequencing • Expression quantitative trait loci (eQTL) • Allelic specific ********ion Department of Biomedical Informatics ...
Prader-Willi syndrome - type 1 deletion, a
... There was no history of repeated miscarriages in any other family members; however, intellectual disability was observed in one of the members on the paternal side. Cytogenetic analysis was carried out from the peripheral blood of the coupleto rule out inheritance of Robertsonian translocation [rob( ...
... There was no history of repeated miscarriages in any other family members; however, intellectual disability was observed in one of the members on the paternal side. Cytogenetic analysis was carried out from the peripheral blood of the coupleto rule out inheritance of Robertsonian translocation [rob( ...
Molecular Inversion Probe

Molecular Inversion Probe (MIP) belongs to the class of Capture by Circularization molecular techniques for performing genomic partitioning, a process through which one captures and enriches specific regions of the genome. Probes used in this technique are single stranded DNA molecules and, similar to other genomic partitioning techniques, contain sequences that are complementary to the target in the genome; these probes hybridize to and capture the genomic target. MIP stands unique from other genomic partitioning strategies in that MIP probes share the common design of two genomic target complementary segments separated by a linker region. With this design, when the probe hybridizes to the target, it undergoes an inversion in configuration (as suggested by the name of the technique) and circularizes. Specifically, the two target complementary regions at the 5’ and 3’ ends of the probe become adjacent to one another while the internal linker region forms a free hanging loop. The technology has been used extensively in the HapMap project for large-scale SNP genotyping as well as for studying gene copy alterationsand characteristics of specific genomic loci to identify biomarkers for different diseases such as cancer. Key strengths of the MIP technology include its high specificity to the target and its scalability for high-throughput, multiplexed analyses where tens of thousands of genomic loci are assayed simultaneously.