* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Objective
Immunoprecipitation wikipedia , lookup
Circular dichroism wikipedia , lookup
Structural alignment wikipedia , lookup
List of types of proteins wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Degradomics wikipedia , lookup
Rosetta@home wikipedia , lookup
Protein domain wikipedia , lookup
Protein design wikipedia , lookup
Protein folding wikipedia , lookup
Protein moonlighting wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Homology modeling wikipedia , lookup
Protein structure prediction wikipedia , lookup
Western blot wikipedia , lookup
Protein purification wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
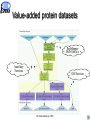
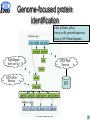
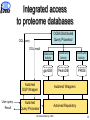
Proteome data integration characteristics and challenges K. Belhajjame1, R. Cote4, S.M. Embury1, H. Fan2, C. Goble1, H. Hermjakob, S.J. Hubbard1, D. Jones3, P. Jones4, N. Martin2, S. Oliver1, C. Orengo3, N.W. Paton1, M. Pentony3, A. Poulovassilis2, J. Siepen, R.D. Stevens1, C. Taylor4, L. Zamboulis2, and W. Zhu4 1University of Manchester 2Birkbeck College 3University College London 4European Bioinformatics Institute Outline Experimental proteomics ISPIDER architecture Example use cases Conclusion All Hands Meetings, 2005 2 Experimental proteomics An essential component for elucidation of the biological functions of proteins The study of the set of proteins produced by an organism with the aim of understanding their behaviour under varying conditions Separation 2D gel electrophoresis Protein digestion Enzymatic digestion Mass Spectrometry Maldi TOF Identification Protein DB Protein ID All Hands Meetings, 2005 3 Experimental proteomics Development of new technologies for: – protein separation (2D-SDS-PAGE, HPLC, Capillary Electrophoresis) – mass spectrometry (Multi-Dimensional protein identification) Availability of publicly accessible protein sequence databases Proteomics databases (PedroDB, gpmDB, PepSeeker, Pride, …) Building experiments involving analysis services orchestration and data processing and integration All Hands Meetings, 2005 4 Objectives of ISPIDER A Grid dedicated to the creation of bioinformatics experiments for proteomics Develop, or make, existing Proteome databases and Grid-enabled services Develop Middleware support for developing and executing new proteome analyses, based on distributed query processing and workflow technologies Undertake proteomic studies that demonstrate the effectiveness of the resulting infrastructure All Hands Meetings, 2005 5 Outline Experimental proteomics ISPIDER architecture Example use cases Conclusion and future directions All Hands Meetings, 2005 6 ISPIDER Vanilla Query Client + Phosph. Extensions 2D Gel Visualisation Client PPI Validation + Analysis Client + Aspergil. Extensions Protein ID Client Web services Proteome Request Handler Proteomic Ontologies/ Vocabularies myGrid WS WS WS PEDRo PID Phos ISPIDER Resources WS GS WS TR ISPIDER Proteomics Grid Infrastructure Data Cleaning Services Existing E-Science Infrastructure AutoMed Ontology Services DQP Workflows PRIDE Instance Ident/Mapping Services myGrid myGrid WS Source Selection Services ISPIDER Proteomics Clients WS WS WS WS PS PF FA PPI Public Proteomics Resources Existing Resources KEY: WS = Web services, GS = Genome sequence, TR = transcriptomic data, PS = protein structure, PF = protein family, FA = functional annotation, PPI = protein-protein interaction data, Work 2005 Package All WP Hands=Meetings, 7 Outline Experimental proteomics ISPIDER architecture Example use cases Conclusion and future directions All Hands Meetings, 2005 8 Value-added protein datasets Motivation Protein identification experiments are usually used as input into further analysis processes. – Gathering evidence for a biological hypothesis – Suggesting new hypotheses Objective Augment the identification results with additional information on the identified protein Implementation Taverna workflow system All Hands Meetings, 2005 9 Value-added protein datasets PepMapper Web Service Auxiliary Services GO Services All Hands Meetings, 2005 10 Genome-focused protein identification Motivation Currently, protein identification searches performed over large data sets. This means fewer false negatives, but false positives are also more likely. Objective More focused and thus more efficient protein identification Implementation Taverna workflow system DQP, a service-based query processor All Hands Meetings, 2005 11 Genome-focused protein identification select p.Name, p.Seq from p in db_proteinSequences where p.OS='HomoSapiens'; PepMapper web service DQP Web Service GOA Web Service IPI All Hands Meetings, 2005 12 Integrated access to proteome databases Motivation Ability to analyse existing proteomics results en masse is limited, because of the heterogeneities between the schemas of the different databases Objective Providing integrated access to proteome databases through a common schema Implementation AutoMed, a framework for mapping heterogeneous schemata DQP, a service-based query processor All Hands Meetings, 2005 13 Integrated access to proteome databases OGSA Distributed Query Processor OQL query OQL result User query Result OGSA-DAI Activity OGSA-DAI Activity OGSA-DAI Activity gpmDB PedroDB PRIDE Automed DQP Wrapper Automed Wrappers Automed Query Processor Automed Repository All Hands Meetings, 2005 14 Conclusions + Available e-science technologies provide rapid prototyping facilities for bioinformatics analyses + Combining such technologies is possible and opens up more possibilities Taverna + DQP Automed + DQP - Writing custom code is usually required – Processing service output to extract inputs for following services – Transforming results between data formats – Dealing with mismatches between identifiers Developing a user-guided environment for the detection and resolution of mismatches Development of Proteomics client applications (PepMapper, PepSeeker and PRIDE) All Hands Meetings, 2005 15