* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Genome-wide characteristics of sequence coverage by next
Transposable element wikipedia , lookup
Public health genomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
X-inactivation wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Genomic imprinting wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Human genome wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Non-coding DNA wikipedia , lookup
RNA interference wikipedia , lookup
Pathogenomics wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Genomic library wikipedia , lookup
Epigenetics of depression wikipedia , lookup
History of RNA biology wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epitranscriptome wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Primary transcript wikipedia , lookup
Genome evolution wikipedia , lookup
Nutriepigenomics wikipedia , lookup
RNA silencing wikipedia , lookup
Non-coding RNA wikipedia , lookup
Metagenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
Long non-coding RNA wikipedia , lookup
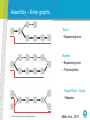
NESCENT : NGS : Measuring expression Jen Taylor Bioinformatics Team CSIRO Plant Industry Measuring Expression • What & Why • What is expression and why do we care? • How • Platforms / Technology • Closed approaches – Microarray • Open approaches - Sequencing • Experimental Design • Analysis • Biases • Bioinformatics • Statistical Issues and Analysis • In action • Workshop – Detection of Differential Expression • Case Studies in Plant functional genomics CSIRO. Nescent August 2011 - Measuring Expression What is expression / transcriptome ? mRNA tRNA rRNA DNA siRNA microRNA piRNA tasiRNA CSIRO. Nescent August 2011 - Measuring Expression lncRNA Beyond the Genome: 1995 Human Genome sequencing begins in earnest “Mapping the Book of Life” 2000 - First Draft = approx 140, 000 genes 2003 - Essential Completion = 30, 000 – 40,000 genes ?? = 24, 195 genes !!!??? Commemorative stained glass window for F.C. Crick, designed by Maria McClafferty.(Photograph: Paul Forster) CSIRO. Nescent August 2011 - Measuring Expression Gonville & Caius College, Cambridge, UK. “The failure of the human genome” “despite more than 700 genome-scanning publications and nearly $100bn spent, geneticists still had not found more than a fractional genetic basis for human disease “ Manolio et al., Nature, 2009 “The most likely explanation for why genes for common diseases have not been found is that, with few exceptions, they do not exist. …., if inherited genes are not to blame for our commonest illnesses, can we find out what is? “ Guardian, 2011 CSIRO. Nescent August 2011 - Measuring Expression Beyond the Genome: Gene Number ≠ Complexity Complexity Gene Regulation Transcriptome CSIRO. Nescent August 2011 - Measuring Expression Commemorative stained glass window for F.C. Crick, designed by Maria McClafferty.(Photograph: Paul Forster) Gonville & Caius College, Cambridge, UK. Why the expression ? High-throughput friendly Genome Predicts Biology ** Regulatory network Transcriptome Context dependent Proteome CSIRO. Nescent August 2011 - Measuring Expression **Li et al., 2004 Measuring Expression ? Parts Description • Function? • Interconnectedness? CSIRO. Nescent August 2011 - Measuring Expression Comparisons • Population - level • Between genomes Measuring Expression ? What are important members of a transcriptome? mRNA • polyadenylated, coding • alternatively spliced Noncoding RNA (small RNA) • varying lengths, functions (18 – 32 bases) • microRNA, siRNA, piRNA, tasiRNA, long non-coding RNA “Dark” RNA • transcription outside of annotated genes • Non-polyadenylated Anti-sense transcription CSIRO. Nescent August 2011 - Measuring Expression Measuring Expression ? How does the transcriptome vary to give rise to phenotype ? Changes in Abundance • Abundance = Rate of Transcription – Rate of Decay Changes in Function • Availability for function – polyadenylation, silencing, localisation • Suitability for function – alternate splicing CSIRO. Nescent August 2011 - Measuring Expression How to measure Expression PLATFORMS / TECHNOLOGY CSIRO. Nescent August 2011 - Measuring Expression Measuring Expression : platforms • Closed systems – microarray • Probes immobilised on a substrate profile target species in the transcriptome CSIRO. Nescent August 2011 - Measuring Expression CSIRO. Nescent August 2011 - Measuring Expression Single and two colour arrays Two colour Single colour Experimental Control Labelling Labelling Sample A Hybridisation Probe Library Array Manufacture CSIRO. Nescent August 2011 - Measuring Expression Array Scanning Array profiling Affymetrix Array Targets • Arabidopsis Genome 24,000 • C. elegans Genome 22,500 • Drosophila Genome 18, 500 • E. coli Genome 20, 366 • Human Genome U133 Plus 47,000 • Mouse Genome 39, 000 • Yeast Genome • S.cerevisiae 5, 841 • S. pombe 5, 031 • Rat Genome 30, 000 • Zebrafish 14, 900 • Plasmodium / Anopheles • P. faciparum 4,300 • A. gambiae 14,900 • Barley (25,500), Soybean (37,500 + 23,300 pathogen), Grape (15,700) • Canine (21,700), Bovine (23,000) • B.subtilis (5,000), S. aureus (3,300 ORFS), Xenopus (14, 400) CSIRO. Nescent August 2011 - Measuring Expression CSIRO. Nescent August 2011 - Measuring Expression CSIRO. Nescent August 2011 - Measuring Expression Closed System – Microarray • Pros • • • • High-throughput Targeted profiling Inexpensive – “population friendly” Analytical methods are standardised • Negative • “Closed system” , novel = invisible • Difficult to see allelle-specific expression • Biases due to hybridisation • SNPs • Competitive and non-specific hybridisation CSIRO. Nescent August 2011 - Measuring Expression Open systems – RNA Sequencing Technology: • Illumina • SOLiD, IonTorrent • 454 Pros: • Transcript discovery • Allelic expression • High resolution abundance measures Cons: • Analysis can be complex • Expensive • Sensitivity is sequencing depth dependent CSIRO. Nescent August 2011 - Measuring Expression RNA Sequencing CSIRO. Nescent August 2011 - Measuring Expression Mortazavi et al., 2008 RNASeq - Correspondence • Range > 5 orders of magnitude • Better detection of low abundance transcripts CSIRO. Nescent August 2011 - Measuring Expression Marioni et al., 2009 Platform Choice / Sample Preparation Choice What do you want to profile ? • Polyadenylated • PolyA RNA extraction • Small RNA (< 100 bases) • Size filtering by gel • Strand-specific • RNA – Protein Interactions • RNA Immunoprecipitation (IP) CSIRO. Nescent August 2011 - Measuring Expression RNASeq - Workflow Sample Total RNA PolyA RNA Mapping to Genome Small RNA Differential Expression SNP detection Transcript structure Library Construction Secondary structure Assembly to Contigs Sequencing Base calling & QC CSIRO. Nescent August 2011 - Measuring Expression Targets or Products Illumina RNASeq : TruSeq CSIRO. Nescent August 2011 - Measuring Expression Small RNA sequencing Small RNA smallRNA separation: PAGE 134 110 75 25 small RNA < 35bp CSIRO. Nescent August 2011 - Measuring Expression Strand - specificity Using adaptors Ligation : 3’ and 5’ adaptors added sequentially Using chemical modification dUTP : Addition and removal after selection SMART : addition of C’s on 5’ end CSIRO. Nescent August 2011 - Measuring Expression Levin et al., 2010 CSIRO. Nescent August 2011 - Measuring Expression Levin et al., 2010 Non-polyA methods • Total RNA extraction • Ribosomal RNA and tRNA > 95-97% of total RNA • Ribosomal reduction methods • Subtractive hybridisation with rRNA probes • Exonuclease cleave of rRNA • NuGen – “proprietary combination of reverse transcriptase and primers in the Ovation RNA-Seq System” • cDNA normalisation methods • Partial digestion of any highly abundant species (Evrogen) CSIRO. Nescent August 2011 - Measuring Expression Platform Choice / Sample Preparation Choice What do you want to profile ? • Polyadenylated • PolyA RNA extraction • Small RNA (< 100 bases) • Size filtering by gel • Strand-specific • RNA – Protein Interactions • RNA Immunoprecipitation (IP) CSIRO. Nescent August 2011 - Measuring Expression • Non - PolyA • rRNA reduction EXPERIMENTAL DESIGN and ANALYSIS CSIRO. Nescent August 2011 - Measuring Expression RNASeq Experimental Design • Issues: • sequencing depth - how much ? • number of replicates – how many ? • Aims of the data : • Transcriptome assembly / transcript characterisation • Maximise depth • Detection of differential expression (denovo or reference) • Balance depth and replication CSIRO. Sequencing Depth V.S. Number of Replicates Library 1 Defining Replicates Library 2 • Technical Replicates Individual Library 1 Lane 1 Individual 1 Library 2 Lane 2 Lane 3 • Biological Replicates Lane 4 Library 3 Individual 2 Library 4 Library 1 Library 2 Multiplex Lane 1 Lane 2 Lane 1 , L1 L2 L3 L4 Depth = 2 x 100% lane / sample CSIRO. Sequencing Depth V.S. Number of Replicates 100% lane / sample 25% lane / sample CSIRO. Sequencing Depth V.S. Number of Replicates Coverage Depth CSIRO. Sequencing Depth V.S. Number of Replicates Number of Replicates # Reps 2 4 6 8 False P 0.03 0.03 0.03 0.03 0.03 0.03 False N 0.84 0.72 0.64 0.59 0.54 0.50 True P 0.16 0.28 0.36 0.41 0.46 0.50 True N 0.97 0.97 0.97 0.97 0.97 0.97 10 12 edgeR <= 0.01 , DESeq <= 0.01 More information in biological replicates than depth For differential expression CSIRO. Sequencing Depth V.S. Number of Replicates RNASeq Analysis • Overall Aim : • To get an accurate measurement of transcript abundance, structure and identity • Biases and Compositions • Alignment • TopHat / Cufflinks • Assembly • ABySS CSIRO. Nescent August 2011 - Measuring Expression Assumptions Every transcript / k-mer has equal chance of being sequenced No. sequences observed ≈ transcript abundance Gene A = z Reads / million Gene B = y Reads / million z=2xy Gene A > Gene B CSIRO. Nescent August 2011 - Measuring Expression Length Bias Oshlack and Wakefield, 2009 CSIRO. Nescent August 2011 - Measuring Expression Alignment Bias CSIRO. Nescent August 2011 - Measuring Expression Alignment Bias CSIRO. Nescent August 2011 - Measuring Expression Sequencing Bias CSIRO. Nescent August 2011 - Measuring Expression Hansen et al., 2010 Bias Every transcript / k-mer has equal chance of being sequenced No. sequences observed ≈ transcript abundance Gene A = z Reads / million / kb Weighting schemas (e.g. Cufflinks) : • Mapability • kmer / fragment frequencies CSIRO. Nescent August 2011 - Measuring Expression Gene B = y Reads / million / kb Bias Every transcript / k-mer has equal chance of being sequenced No. sequences observed ≈ transcript abundance Sample A vs Sample B Gene A1 = z Reads per million z=2xy CSIRO. Nescent August 2011 - Measuring Expression Gene A2 = y Reads per million Read density variability CSIRO. Nescent August 2011 - Measuring Expression RNASeq – Compositional properties Depth of Sequence • Sequence count ≈ Transcript Abundance • Majority of the data can be dominated by a small number of highly abundant transcripts • Ability to observe transcripts of smaller abundance is dependent upon sequence depth • Fixed budget of reads CSIRO. Nescent August 2011 - Measuring Expression A simple example – compositional bias Sequencing budget / depth: 4000 reads sample I A B C D Expected counts 1000 1000 1000 1000 CSIRO. Nescent August 2011 - Measuring Expression sample II A B Expected counts 2000 2000 Soil diversity by phylogenetic analysis - Phylum level 454-sequence analysis of bacterial 16S rRNA gene ~410,000 sequences Recognized bacterial phyla A B C 0% 20% 40% 60% 80% 100% % distribution CSIRO. Nescent August 2011 - Measuring Expression A. Richardson, CSIRO RNASeq Bioinformatics Analysis • Aims: • To get an accurate measurement of transcript abundance, structure and identity • Biases and Compositions • Relative abundances NOT absolute • Alignment • TopHat • Assembly • ABySS CSIRO. Nescent August 2011 - Measuring Expression RNA Sequencing analysis Sequence Data Genome? Alignment Assembly Read Density Contigs Differential Expression SNPs Transcript Characterisation CSIRO. Nescent August 2011 - Measuring Expression RNASeq – Alignment Considerations Reads with multiple locations • Discard / Random Allocation • Clustering - local coverage • Weighting Reads Spanning Exons • Make and align to exon junction libraries • Denovo junction detection Summarisation of counts • Exons • Transcript boundaries • Inferred read boundaries CSIRO. Nescent August 2011 - Measuring Expression TopHat Multimapping : ≤10 sites Assembly : consensus ‘island’ exon CSIRO. Nescent August 2011 - Measuring Expression Trapnell et al., 2009; Roberts et al., 2011 TopHat / Cufflinks Heuristics : • “Correct” errors in low coverage areas • Grabs 45 bp either side of islands to capture splice sites • Collapse small islands • Looks for junctions within larger islands, highly covered Cufflinks : • calculates the probability of observing a certain fragment within a given transcript given surrounding fragments. CSIRO. Nescent August 2011 - Measuring Expression Trapnell et al., 2009; Roberts et al., 2011 Alignment • Great if you have a fully annotated, reference • Okay.. If you have a partially annotated reference • “Different” if you have a big bunch of ESTs Options: • Align to a neighbouring genome or EST library • Denovo transcriptome assembly Tools: • ABySS, Mira, Trinity, HT-Seq, SAMtools CSIRO. Nescent August 2011 - Measuring Expression RNA Sequencing analysis Sequence Data Genome? Alignment Assembly Read Density Contigs Differential Expression SNPs Transcript Characterisation CSIRO. Nescent August 2011 - Measuring Expression Denovo transcriptome assembly • • • • • • • • • • • • • ABySS MIRA Trinity Velvet AllPaths Soap-denovo Euler CABOG Edena SHARCGS VCAKE SSAKE CAP3 CSIRO. Nescent August 2011 - Measuring Expression • Will run on reasonable computer resources for large genomes • (e.g. < 1 TB of RAM) • Paired end data handling • Platform flexible • Handles haplotype complexity and polyploid genomes Denovo transcriptome assembly • • • • • • • • • • • • • ABySS MIRA Trinity Velvet AllPaths Soap-denovo Euler CABOG Edena SHARCGS VCAKE SSAKE CAP3 CSIRO. Nescent August 2011 - Measuring Expression • Will run on reasonable computer resources for large genomes • (e.g. < 1 TB of RAM) • Handles paired end data • Handles data from all platforms • Handles haplotype complexity and polyploid genomes Assembly – Kmer graphs K=4 CSIRO. Nescent August 2011 - Measuring Expression Miller et al., 2010 Assembly – Kmer graphs Spurs • Sequencing error Bubbles • Sequencing error • Polymorphism Frayed Rope / Cycles • Repeats CSIRO. Nescent August 2011 - Measuring Expression Miller et al., 2010 Assembly – Kmer graphs Spurs • Sequencing error Bubbles • Sequencing error • Polymorphism Frayed Rope / Cycles • Repeats CSIRO. Nescent August 2011 - Measuring Expression Miller et al., 2010 ABySS & TransABySS • User specifies k • Optimal k depends on sequencing depth CSIRO. Nescent August 2011 - Measuring Expression ABySS & TransABySS • Sequencing depth is relative to transcript abundance • Iterate over multiple k and merge • Contigs contained within a large contig are “buried” CSIRO. Nescent August 2011 - Measuring Expression Assessing assembly quality ? • Comparisons between assembly algorithms • Contig summary statistics • Comparisons to known resources (e.g. ESTs) Trial on Rice Transcriptome: • 120 Million 75 bp single end Illumina reads – embryo • ABySS : • Number of contigs = 6, 804 • Contig length range = 38 – 2,818 [mean = 203] • Database comparisons : • Rice public cDNA sequences : 67, 393 • Contigs with high quality matches to cDNA : 6,555 (96%) CSIRO. Nescent August 2011 - Measuring Expression RNASeq Bioinformatics Analysis • Aims: • To get an accurate measurement of transcript abundance, structure and identity • Biases and Compositions • Relative abundances NOT absolute • Alignment • Assembly CSIRO. Nescent August 2011 - Measuring Expression STATISTICAL ISSUES CSIRO. Nescent August 2011 - Measuring Expression Measuring Expression – Statistical Issues • Data elements • Normalisation • Detection of Differential Expression CSIRO. Nescent August 2011 - Measuring Expression Count Data : of what ? CSIRO. Nescent August 2011 - Measuring Expression Count Data : of what ? CSIRO. Nescent August 2011 - Measuring Expression Garber et al., 2011 Statistical analysis of RNASeq • Count data • Distribution is positively skewed, not normal • Between sample variability in counts - normalisation CSIRO. Nescent August 2011 - Measuring Expression Normalization is required Two scenarios : 1. Different sizes of total reads (library size) 2. Fixed library size, subset of highly expressed reads in 1 sample. Both reduce sequencing budget available for the majority of transcripts CSIRO. Nescent August 2011 - Measuring Expression Normalisation • Assume the majority of log ratios = 0 [No change] TMM : Trimmed Mean of M values (log ratios) Adjust TMM to be equal between samples CSIRO. Nescent August 2011 - Measuring Expression Robinson and Oshlack, 2010 DE genes with and without TMM normalization CSIRO. Nescent August 2011 - Measuring Expression RNASeq data – Poisson Distributions • Poisson distributions are used when things are counted • The probability of seeing n events in a fixed time or space • The number of lions on a 1 day safari • The number of raindrops on a tennis court • The number of flying elephants in a year • Requires λ : rate of events • Variance = mean = λ CSIRO. Nescent August 2011 - Measuring Expression RNASeq data – Negative Binomial • RNASeq data is more variable than Poisson • Variance > mean = λ • Less prominent for large mean • Over-dispersed Poisson Noise types • Shot noise • Unavoidable, prominent for low mean • Technical noise • Small, hopefully, can be managed • Biological noise • Sample differences CSIRO. Nescent August 2011 - Measuring Expression RNA Seq • Variance also depends on the mean CSIRO. Nescent August 2011 - Measuring Expression Anders, 2010 RNASeq Model The total counts for a transcript in sample j from condition c : s j c s j vc 2 Library normalisation Mean Value Fitted Variance (overdispersion) For a given gene , test for a difference in counts between conditions. Is mean c1 + mean c2 statistically different to mean c1 + mean c1? CSIRO. Nescent August 2011 - Measuring Expression RNASeq DE Testing • • • • • DESeq – Anders and Huber, 2010 EdgeR – Robinson et al., 2009 – R BaySeq – Hardcastle and Kelley, 2010 – R DEGSeq – Wang et al., 2010 – R NBP - Di et al., 2011 • LOX – Zhang et al., 2010 • Infers expression measures allowing for incorporation of noise from different methodologies in the one experimental design CSIRO. Nescent August 2011 - Measuring Expression Measuring Expression • What & Why • What is expression and why do we care? • How • Platforms / Technology • Closed approaches – Microarray • Open approaches - Sequencing • Experimental Design • Analysis • Biases • Bioinformatics • Statistical Issues and Analysis • In action • Workshop – Detection of Differential Expression • Case Studies in Plant functional genomics CSIRO. Nescent August 2011 - Measuring Expression Acknowledgements Plant Industry Jennifer M Taylor Bionformatics Leader Phone: +61 2 62464929 Email: [email protected] Jose Robles Stuart Stephen Hua Ying Andrew Spriggs Alexie Pa NESCENT Funding Thank you Contact Us Phone: 1300 363 400 or +61 3 9545 2176 Email: [email protected] Web: www.csiro.au