* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Slide 1
Expanded genetic code wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Genetic code wikipedia , lookup
Gene expression wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
List of types of proteins wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Biochemistry wikipedia , lookup
Protein design wikipedia , lookup
Protein moonlighting wikipedia , lookup
Rosetta@home wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Circular dichroism wikipedia , lookup
Folding@home wikipedia , lookup
Metalloprotein wikipedia , lookup
Western blot wikipedia , lookup
Implicit solvation wikipedia , lookup
Homology modeling wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Interactome wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein adsorption wikipedia , lookup
Protein–protein interaction wikipedia , lookup
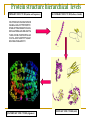
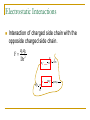
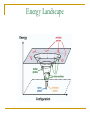
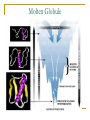
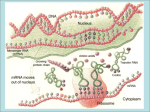
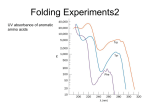
CMSE SEMINAR Protein Folding mechanisms By Sefer Baday OUTLINE Proteins Protein Folding Forces Driving Folding Energy landscape The folding mechanism models Conclusion Some Facts about Proteins Composed of amino acids. Each sequence fold in unique structure-native structure Proteins are functional only in their native states Folding is reversibe unfolding or re-folding is possible Modest changes in the environment can cause structural changes in the protein,thus affecting its function Protein structure hierarchical levels PRIMARY STRUCTURE (amino acid sequence) SECONDARY STRUCTURE (helices, strands) VHLTPEEKSAVTALWGKVNVDE VGGEALGRLLVVYPWTQRFFE SFGDLSTPDAVMGNPKVKAHG KKVLGAFSDGLAHLDNLKGTFA TLSELHCDKLHVDPENFRLLGN VLVCVLAHHFGKEFTPPVQAAY QKVVAGVANALAHKYH QUATERNARY STRUCTURE (oligomers) TERTIARY STRUCTURE (fold) What is Protein Folding ? • Protein folding is the process by which a protein assumes its functional shape or conformation. Random Coil Native conformation Why is the “Protein Folding” so important Most of the proteins should fold in order to function Misfolding cause some diseases. Cystic Fibrosis ,affects lungs and digestive system and cause early death Alzheimers’s and Parkinson's disease It may help us to understand the structure of proteins which has not been known LEVINTHAL PARADOX Let have Protein composed of 100 amino acids. Assume that each amino acid has only 3 possible conformations. Total number of conformations = 3100 ~= 5x1047 . If 100 psec (10-10 sec) were required to convert from a conformation to another one, a random search of all conformations would require 5x1047 x 10-10 sec = 1.6 x 1030 years. However, folding of proteins takes place in msec to sec order. Forces that stabilize protein structure Interactions between atoms within the protein chain Interactions between the protein and the solvent Electrostatic Interactions Interaction of charged side chain with the opposide charged side chain. q1q 2 F O Dr 2 C CH + NH3 - O H2C C O 2 O + NH3 (CH2)4 Hydrogen Bonds Noncovalent bond Energy:10-40 kJ/ mol Strength varies with angle of hydrogen bond interaciton. van der Waals forces Between all atoms Approximately 1kj/mol Van der Waals potential Van der Waals Force Van der waals radii of common atoms (nm): H 0.1 nm C 0.17 nm N 0.15 nm O 0.14 nm P 0.19 nm S 0.185 nm r0 r0 r Average Strength of Interactions Bond Type kJ/mol Covalent Bond 250 Electrostatic 5 van der Waals 5 Hydrogen bond 20 The Hydrophobic Interaction Hydrophobic means afraid of water Hydrophobic residues are buried in while hyrophilic residues stay outside. Hydrogen Bonds The kinetic Theory of Protein Folding Folding proceeds through a definite series of steps or a Pathway. A protein does not try out all possible rotations of conformational angles, but only enough to find the pathway. Energy Landscape Energy Landscape Molten Globule Contact Order The average separation in the sequence between residues that are in contact with each other in native structure Phi Value Analysis Experimental method to study of the structure of the transition state Using mutations as a structural report Phi=1, transition state has native like structure Phi=0, transition state has denatured like structure The Framework Model Local interactions are main determinants of protein structures native state unfolded state Transition state Hydrophobic Collapse Hydrophobic core forms first. collapse unfolded state native state Hydrophobic Collapse Formation of hydrophobic globule may hinder the reorganization of both side chains and whole protein Nucleation Model Unites hydrophobic collapse and frame work model unfolded state formation of a nucleus native state Nucleation Model Substantial expulsion of water from the burial of non polar surfaces Good correlation between decrease in hyrodynamic volume and increase in secondary structure Unfolding simulation of Ci2 The folding Pathways of Barnase Conclusions Non local interactions( Hydrophobic effect and van der waals ) are needed to bring protein into a globular conformation. Chemically specific interactions( hydrogen bonds, electorstatic interactions) determine the fine detail of the protein structure Conclusions The folding process is hierarchical Native topology affects the folding mechanism. Nucleation method explains folding mechanism better than framework and hydrophobic collapse methods. THANK YOU QUESTIONS ???