How do I identify exon number with the UCSC Genome Browser
... couple of links that take us to a location where we can choose our genome of interest. We will click here to reset to set the browser to the default genome assembly which is the human hg19 assembly at the present time. The [submit] button takes us into the main browser viewer with a large number of ...
... couple of links that take us to a location where we can choose our genome of interest. We will click here to reset to set the browser to the default genome assembly which is the human hg19 assembly at the present time. The [submit] button takes us into the main browser viewer with a large number of ...
Two-Exon Skipping Due to a Point Mutation in p67
... proline-rich sequence and the mutant protein may have lost its stability due to a loss of these sequences. The sequences, therefore, contributetoprotein stabilitythrough intra-or intermolecular association of p67-phox with itself or with other proteins such as p47-phox or p40-phox that contain simil ...
... proline-rich sequence and the mutant protein may have lost its stability due to a loss of these sequences. The sequences, therefore, contributetoprotein stabilitythrough intra-or intermolecular association of p67-phox with itself or with other proteins such as p47-phox or p40-phox that contain simil ...
The alternative splicing of tau exon10 and its
... Jiang et al. 2003; Wang et al. 2004). Like other alternative exons, exon 10 is regulated by a finely tuned balance of sequences and trans-acting factors. Exon 10 contains two enhancers, a GAR (guanidine/adenosine-rich) and an ACE (adenosine/cytosine-enhancer) motif and two weak silencers that are dis ...
... Jiang et al. 2003; Wang et al. 2004). Like other alternative exons, exon 10 is regulated by a finely tuned balance of sequences and trans-acting factors. Exon 10 contains two enhancers, a GAR (guanidine/adenosine-rich) and an ACE (adenosine/cytosine-enhancer) motif and two weak silencers that are dis ...
Zeeberg - Gene Ontology Consortium
... • GoMiner traditionally dereplicates input files so that only one instance of a gene name is processed • When multiple alternatively spliced forms are to be analyzed, however, dereplication would result in a loss of relevant information • Consequently, we have added a new feature to GoMiner to retai ...
... • GoMiner traditionally dereplicates input files so that only one instance of a gene name is processed • When multiple alternatively spliced forms are to be analyzed, however, dereplication would result in a loss of relevant information • Consequently, we have added a new feature to GoMiner to retai ...
Chapter 14 Guided Reading
... 21. Use the diagram below to demonstrate initiation of transcription at a eukaryotic promoter. Label all parts of the diagram and discuss what is occurring at each step.. ...
... 21. Use the diagram below to demonstrate initiation of transcription at a eukaryotic promoter. Label all parts of the diagram and discuss what is occurring at each step.. ...
Chapter 17 notes
... Figure 17.2 Overview: the roles of transcription and translation in the flow of genetic ...
... Figure 17.2 Overview: the roles of transcription and translation in the flow of genetic ...
Fundamentals of Nucleic Acid Biochemistry: RNA
... response elements (HREs). HREs often occur in multiple copies in enhancer sequence regions. ...
... response elements (HREs). HREs often occur in multiple copies in enhancer sequence regions. ...
Interconnections Between RNA-Processing Pathways Revealed by
... these were factors encoding known or predicted components of the spliceosome. Included among ...
... these were factors encoding known or predicted components of the spliceosome. Included among ...
alternatively-spliced protein sequences derived
... Many proteins exist in more than one isoform, one cause of which is differential splicing: up to 30% of human genes are believed to exist in alternatively spliced isoforms. Isoforms may differ quite considerably from one another, with potentially less than 50% sequence similarity. In the SWISS-PROT ...
... Many proteins exist in more than one isoform, one cause of which is differential splicing: up to 30% of human genes are believed to exist in alternatively spliced isoforms. Isoforms may differ quite considerably from one another, with potentially less than 50% sequence similarity. In the SWISS-PROT ...
Serine/Arginine-rich proteins Physcomitrella patens Andreas Ring
... functionally different protein products from a single gene (Graveley, 2001; Manley and Tacke, 1996). This makes AS a major contributor to protein diversity, but AS is also a form of post-transcriptional gene regulation. AS of pre-mRNA transcripts in which introns or “poison exons” are spliced into a ...
... functionally different protein products from a single gene (Graveley, 2001; Manley and Tacke, 1996). This makes AS a major contributor to protein diversity, but AS is also a form of post-transcriptional gene regulation. AS of pre-mRNA transcripts in which introns or “poison exons” are spliced into a ...
Computational Definition of
... internal non-protein-coding exons of these genes as examples of exons that should have a relatively high content of ESEs and a relatively low content of ESSs, but with no protein-coding information. We compared the sequence composition of these non-coding exons with that of 2 dissimilar types of seq ...
... internal non-protein-coding exons of these genes as examples of exons that should have a relatively high content of ESEs and a relatively low content of ESSs, but with no protein-coding information. We compared the sequence composition of these non-coding exons with that of 2 dissimilar types of seq ...
authors` original image
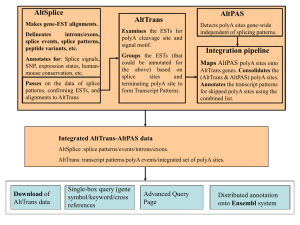
... Makes gene-EST alignments. Delineates introns/exons, splice events, splice patterns, peptide variants, etc. Annotates for: Splice signals, SNP, expression states, humanmouse conservation, etc. ...
... Makes gene-EST alignments. Delineates introns/exons, splice events, splice patterns, peptide variants, etc. Annotates for: Splice signals, SNP, expression states, humanmouse conservation, etc. ...
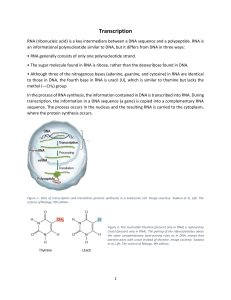
Transcription
... the first 5’-triphosphate is NOT cleaved proceeds in 5’ to 3’ direction new residues are added to the 3’ OH the template is copied in the 3’ – 5’ direction forms a temporary DNA:RNA hybrid transcription rate ( 50 to 90 nts/sec) RNA polymerase has complete processivity ...
... the first 5’-triphosphate is NOT cleaved proceeds in 5’ to 3’ direction new residues are added to the 3’ OH the template is copied in the 3’ – 5’ direction forms a temporary DNA:RNA hybrid transcription rate ( 50 to 90 nts/sec) RNA polymerase has complete processivity ...
Transcription
... O Can control Z only when on same chromosome !!! -> cis acting control I is trans acting factor Proteins are synthesized in two stages: 1. DNA is transcribed in mRNA 2. mRNA is translated into protein This model explains behavior of lac system ...
... O Can control Z only when on same chromosome !!! -> cis acting control I is trans acting factor Proteins are synthesized in two stages: 1. DNA is transcribed in mRNA 2. mRNA is translated into protein This model explains behavior of lac system ...
Exons and Introns Characterization in Nucleic Acid Sequences by
... (DNA) sequence analysis is to determine the exact locations of the genes and also in eukaryotes, the protein-coding regions in the mRNA primary transcript (pre-mRNA).The conversion into discrete numerical values of the symbols associated to the nucleotides of these sequences allows for a signal to a ...
... (DNA) sequence analysis is to determine the exact locations of the genes and also in eukaryotes, the protein-coding regions in the mRNA primary transcript (pre-mRNA).The conversion into discrete numerical values of the symbols associated to the nucleotides of these sequences allows for a signal to a ...
The Interplay of Temperature and Genotype on Patterns
... alternative splicing is not yet clear. We studied variation in alternative splicing among four different temperatures, 13, 18, 23, and 29°, in two Drosophila melanogaster genotypes. We show plasticity of alternative splicing with up to 10% of the expressed genes being differentially spliced between ...
... alternative splicing is not yet clear. We studied variation in alternative splicing among four different temperatures, 13, 18, 23, and 29°, in two Drosophila melanogaster genotypes. We show plasticity of alternative splicing with up to 10% of the expressed genes being differentially spliced between ...
Transcription and RNA processing
... prokaryotic and eukaryotic genes, transcription begins at a DNA sequence that is upstream (to the “left” on the DNA) of the first codon (i.e., at the promoter), and ends downstream (to the “right” on the DNA) of the termination codon. In eukaryotes, there is usually a “polyadenylation” sequence (AAU ...
... prokaryotic and eukaryotic genes, transcription begins at a DNA sequence that is upstream (to the “left” on the DNA) of the first codon (i.e., at the promoter), and ends downstream (to the “right” on the DNA) of the termination codon. In eukaryotes, there is usually a “polyadenylation” sequence (AAU ...
development and mature motor function The splicing
... Received November 1, 2011; revised version accepted January 31, 2012. ...
... Received November 1, 2011; revised version accepted January 31, 2012. ...
Geuvadis Analysis Meeting
... - Quantified 615 datasets based on the Gencode v7 annotation - Sensitivity is a function of sequencing depth ...
... - Quantified 615 datasets based on the Gencode v7 annotation - Sensitivity is a function of sequencing depth ...
Alternative Splicing: How to Get More than One Protein from a Gene
... Alternative Splicing: How to Get More than One Protein from a Gene Description: Use the word key from the “Protein Synthesis and Words” activity to demonstrate how eukaryotic cells may use one DNA sequence to code for multiple proteins. Eukaryotic cells might use the same gene or DNA sequence differ ...
... Alternative Splicing: How to Get More than One Protein from a Gene Description: Use the word key from the “Protein Synthesis and Words” activity to demonstrate how eukaryotic cells may use one DNA sequence to code for multiple proteins. Eukaryotic cells might use the same gene or DNA sequence differ ...
Natalia Gromak, Alexis Rideau,
... Introduction Alternative splicing is one of the major mechanisms that allows for an expressed proteome that is far more complex than predicted by the number of genes in a genome (Black, 2000; Graveley, 2001; Maniatis and Tasic, 2002; Modrek and Lee, 2002; Roberts and Smith, 2002). It allows the prod ...
... Introduction Alternative splicing is one of the major mechanisms that allows for an expressed proteome that is far more complex than predicted by the number of genes in a genome (Black, 2000; Graveley, 2001; Maniatis and Tasic, 2002; Modrek and Lee, 2002; Roberts and Smith, 2002). It allows the prod ...
The SR Protein SRp38 Represses Splicing in M Phase Cells
... RBD showed the highest homology with that of the classical SR protein SC35 (46% identity). Based on the apparent size of the proteins, and in keeping with the naming system used for most SR proteins, we call the full-length protein SRp38 and the smaller form SRp38-2 (see Figure 1A). During the cours ...
... RBD showed the highest homology with that of the classical SR protein SC35 (46% identity). Based on the apparent size of the proteins, and in keeping with the naming system used for most SR proteins, we call the full-length protein SRp38 and the smaller form SRp38-2 (see Figure 1A). During the cours ...
Regulation of Gene Expression by Coupling of Alternative Splicing
... removing protein domains, affecting protein activity, or altering the stability of the transcript or the resulting protein.4-6 In the last few years, it has become clear that many alternative splice forms previously thought to encode truncated proteins are actually targets of NMD (Fig. 1). In mammal ...
... removing protein domains, affecting protein activity, or altering the stability of the transcript or the resulting protein.4-6 In the last few years, it has become clear that many alternative splice forms previously thought to encode truncated proteins are actually targets of NMD (Fig. 1). In mammal ...
Alternative splicing

Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. Consequently the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes. Alternative splicing is sometimes termed differential splicing.Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome; in humans, ~95% of multi-exonic genes are alternatively spliced. There are numerous modes of alternative splicing observed, of which the most common is exon skipping. In this mode, a particular exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.The production of alternatively spliced mRNAs is regulated by a system of trans-acting proteins that bind to cis-acting sites on the primary transcript itself. Such proteins include splicing activators that promote the usage of a particular splice site, and splicing repressors that reduce the usage of a particular site. Mechanisms of alternative splicing are highly variable, and new examples are constantly being found, particularly through the use of high-throughput techniques. Researchers hope to fully elucidate the regulatory systems involved in splicing, so that alternative splicing products from a given gene under particular conditions could be predicted by a ""splicing code"".Abnormal variations in splicing are also implicated in disease; a large proportion of human genetic disorders result from splicing variants. Abnormal splicing variants are also thought to contribute to the development of cancer.