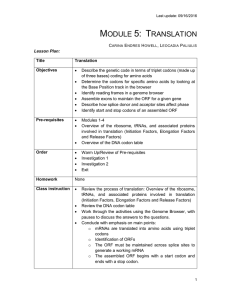
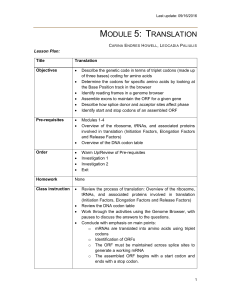
Lecture 8: RNA-sequence analysis: Expression, isoforms
... isoform i with random noise added • Hypothesis H1 – Condition A and B differentially express isoform • Degrees of freedom (dof) is the number of free parameters in H1 minus the number of free parameters in H0; in this case degrees of freedom is 4 – 2 = 2 (H1 has an extra mean and ...
... isoform i with random noise added • Hypothesis H1 – Condition A and B differentially express isoform • Degrees of freedom (dof) is the number of free parameters in H1 minus the number of free parameters in H0; in this case degrees of freedom is 4 – 2 = 2 (H1 has an extra mean and ...
Ehlers-Danlos syndrome type VIIA and VIIB result from splice
... gocele in the right occipital region and had a right intracranial hemorrhage in the perinatal period. Her hips were treated with a Pavlik harness but without success. An open reduction was performed at the age of 2 months on both hips. Because of difficulty walking, a second bilateral open reduction ...
... gocele in the right occipital region and had a right intracranial hemorrhage in the perinatal period. Her hips were treated with a Pavlik harness but without success. An open reduction was performed at the age of 2 months on both hips. Because of difficulty walking, a second bilateral open reduction ...
Figure 5 - GEP Community Server
... looking at the genome. You saw an example of this previously in Module 1. Sometimes we can infer the correct reading frame given the pattern of start and stop codons within the region of the exon, identified by RNA-Seq data. But that sort of information does not always give a definitive answer – the ...
... looking at the genome. You saw an example of this previously in Module 1. Sometimes we can infer the correct reading frame given the pattern of start and stop codons within the region of the exon, identified by RNA-Seq data. But that sort of information does not always give a definitive answer – the ...
module 5: translation - GEP Community Server
... looking at the genome. You saw an example of this previously in Module 1. Sometimes we can infer the correct reading frame given the pattern of start and stop codons within the region of the exon, identified by RNA-Seq data. But that sort of information does not always give a definitive answer – the ...
... looking at the genome. You saw an example of this previously in Module 1. Sometimes we can infer the correct reading frame given the pattern of start and stop codons within the region of the exon, identified by RNA-Seq data. But that sort of information does not always give a definitive answer – the ...
this help page as PDF
... The Exhaust Align Size is the maximum sequence length for exhaustive search: because the Needleman-Wunsch algorithm is slow, the optimal alignment will not be computed in DNA regions longer than the given value (default: 500 nucleotides). Instead, Scipio will try here to place additional amino acids ...
... The Exhaust Align Size is the maximum sequence length for exhaustive search: because the Needleman-Wunsch algorithm is slow, the optimal alignment will not be computed in DNA regions longer than the given value (default: 500 nucleotides). Instead, Scipio will try here to place additional amino acids ...
Analysis continued Each TopHat run will result in four files: a list of
... Use Sequence Data. Use sequence data for some optional classification functions, including the addition of the p_id attribute required by CuffDiff, which is the identifier for the coding ...
... Use Sequence Data. Use sequence data for some optional classification functions, including the addition of the p_id attribute required by CuffDiff, which is the identifier for the coding ...
DNA transcription 3.lecture ENG OK
... Alternative splicing allows production of a large variety of proteins from a limited amount of DNA. It is now thought that between 30 and 60% of human genes undergo alternative splicing. Many human diseases (15%) may be connected to problems with splicing. Explains how 25000 genes of the hum ...
... Alternative splicing allows production of a large variety of proteins from a limited amount of DNA. It is now thought that between 30 and 60% of human genes undergo alternative splicing. Many human diseases (15%) may be connected to problems with splicing. Explains how 25000 genes of the hum ...
Document
... • genes are also associated with additional sequences of DNA 1. core promoter sequence – for the binding of the RNA polymerase -RNA polymerase recognizes specific sequences of nt’s -binding is helped out by transcription factors 2. enhancer regions – help enhance transcription can be several thousan ...
... • genes are also associated with additional sequences of DNA 1. core promoter sequence – for the binding of the RNA polymerase -RNA polymerase recognizes specific sequences of nt’s -binding is helped out by transcription factors 2. enhancer regions – help enhance transcription can be several thousan ...
Document
... • genes are also associated with additional sequences of DNA 1. core promoter sequence – for the binding of the RNA polymerase -RNA polymerase recognizes specific sequences of nt’s -binding is helped out by transcription factors 2. enhancer regions – help enhance transcription can be several thousan ...
... • genes are also associated with additional sequences of DNA 1. core promoter sequence – for the binding of the RNA polymerase -RNA polymerase recognizes specific sequences of nt’s -binding is helped out by transcription factors 2. enhancer regions – help enhance transcription can be several thousan ...
Snímek 1
... Small nuclear ribonucleoproteins (snRNPs) are active in recognizing and removing introns from pre-mRNA in the nucleus. Each snRNP particle is composed of small nuclear RNA (snRNA) of approximately 150 nucleotides, several Sm proteins and a number of specific proteins that are unique for each snRNP. ...
... Small nuclear ribonucleoproteins (snRNPs) are active in recognizing and removing introns from pre-mRNA in the nucleus. Each snRNP particle is composed of small nuclear RNA (snRNA) of approximately 150 nucleotides, several Sm proteins and a number of specific proteins that are unique for each snRNP. ...
Sequence Alignment - Bilkent University
... duplications through non-allelic recombination Provide new regulatory elements into the neighboring genes where they insert (new enhancers, promoters, and polyA tails) ...
... duplications through non-allelic recombination Provide new regulatory elements into the neighboring genes where they insert (new enhancers, promoters, and polyA tails) ...
synthase is regulated by mRNA splicing
... ies have been done on the relationship of prostaglandin synthesis and cell division, and it is now well established that many mitogens induce PGHS activity (4-11). Significantly, it has also been shown that some nonsteroidal antiinflammatory drugs exert antiproliferative and antitumor activities in ...
... ies have been done on the relationship of prostaglandin synthesis and cell division, and it is now well established that many mitogens induce PGHS activity (4-11). Significantly, it has also been shown that some nonsteroidal antiinflammatory drugs exert antiproliferative and antitumor activities in ...
How were introns inserted into nuclear genes?
... A group II intron could mutate into a classical intron. (a) Proposed sequence of about intron insertion19, and group I events. (b) Example of a group II intron (from Ref. 22) which would have introns are now known to insert then> classical splice signals given a single-base mutation ('). selves (see ...
... A group II intron could mutate into a classical intron. (a) Proposed sequence of about intron insertion19, and group I events. (b) Example of a group II intron (from Ref. 22) which would have introns are now known to insert then> classical splice signals given a single-base mutation ('). selves (see ...
FOUR la INVARIANT CHAIN FORMS DERIVE
... vector, pcD, described by Okayama and Berg (14), using mRNA isolated from the human lay lymphoblastoid cell line, Raji. The libraries were screened by colony hybridization with a yl chain partial cDNA (15) insert. ^-150 positive colonies were isolated from a single library, with a frequency of ^-1 i ...
... vector, pcD, described by Okayama and Berg (14), using mRNA isolated from the human lay lymphoblastoid cell line, Raji. The libraries were screened by colony hybridization with a yl chain partial cDNA (15) insert. ^-150 positive colonies were isolated from a single library, with a frequency of ^-1 i ...
HL7_-_CAP_Cancer_Biomarker_Reporting_Committee
... tyrosine kinase inhibitors. ## This form of EGFR activating mutation is generally associated with resistance to EGFR tyrosine kinase inhibitors although insertions at or before position 768 can be associated with sensitivity. ### This mutation is typically secondary to other EGFR activating mutation ...
... tyrosine kinase inhibitors. ## This form of EGFR activating mutation is generally associated with resistance to EGFR tyrosine kinase inhibitors although insertions at or before position 768 can be associated with sensitivity. ### This mutation is typically secondary to other EGFR activating mutation ...
Temperature-Sensitive Mutations Made Easy: Generating
... and thus were discarded. Forty-two alleles were eliminated at this step because of leaky splicing at 30°. Step 5: To eliminate duplicate TS alleles. The remaining 49 alleles were sequenced, and 40 unique alleles were identified (Figure S2). No obvious pattern of mutations emerged from the analysis o ...
... and thus were discarded. Forty-two alleles were eliminated at this step because of leaky splicing at 30°. Step 5: To eliminate duplicate TS alleles. The remaining 49 alleles were sequenced, and 40 unique alleles were identified (Figure S2). No obvious pattern of mutations emerged from the analysis o ...
Transcript Isoform Differences Across Human Tissues Are
... isoforms can behave like completely distinct proteins when considering their proteinprotein interaction capabilities 15 . Recently, it has been reported that a large majority of alternatively spliced RNAs bind to ribosomes 16 , which suggests that most of them could be translated into proteins and t ...
... isoforms can behave like completely distinct proteins when considering their proteinprotein interaction capabilities 15 . Recently, it has been reported that a large majority of alternatively spliced RNAs bind to ribosomes 16 , which suggests that most of them could be translated into proteins and t ...
Career of Tom Muir
... ■ First example of protein splicing by small molecule ■ MBP and His are model protein ■ No structural or sequence restrictions to exteins ...
... ■ First example of protein splicing by small molecule ■ MBP and His are model protein ■ No structural or sequence restrictions to exteins ...
SMN1
... 3' splice site (3' SS) and 5' splice site (5' SS) are shown below with numbers indicating the percent prevalence of the most frequent nucleotides at each position. Shown in red are the branch point adenosine as well as the virtually invariant last two and first two nucleotides of the introns. (B) Ro ...
... 3' splice site (3' SS) and 5' splice site (5' SS) are shown below with numbers indicating the percent prevalence of the most frequent nucleotides at each position. Shown in red are the branch point adenosine as well as the virtually invariant last two and first two nucleotides of the introns. (B) Ro ...
Temperature-sensitive control of protein activity by conditionally
... responsible for their phenotype. No mutations were found in the DNA encoding the flanking regions of the Gal4 host gene, while 10 conservative and 27 missense mutations were identified within intein sequences (see Supplementary Fig. 1 online for details). To be of general use as a universal c temper ...
... responsible for their phenotype. No mutations were found in the DNA encoding the flanking regions of the Gal4 host gene, while 10 conservative and 27 missense mutations were identified within intein sequences (see Supplementary Fig. 1 online for details). To be of general use as a universal c temper ...
Document
... been revised from 5154 to 2222… – FANTOM/RIKEN Consortium Science, March 2006 Brendan Frey ...
... been revised from 5154 to 2222… – FANTOM/RIKEN Consortium Science, March 2006 Brendan Frey ...
Alternatively Spliced Genes
... and 3). This chapter will focus on the splicing of introns in the major class. This is because this class accounts for more than 99% of known introns. Most of our knowledge about pre-mRNA splicing has also come from studies of the major class of introns. Comparison of genes coding for components of ...
... and 3). This chapter will focus on the splicing of introns in the major class. This is because this class accounts for more than 99% of known introns. Most of our knowledge about pre-mRNA splicing has also come from studies of the major class of introns. Comparison of genes coding for components of ...
Transcription
... within nucleus • snRNA: a class of small RNA molecules within the nucleus snRNA ...
... within nucleus • snRNA: a class of small RNA molecules within the nucleus snRNA ...
gene transcription and rna modification
... – This in turn corresponds to the sequence of amino acid in the polypeptide ...
... – This in turn corresponds to the sequence of amino acid in the polypeptide ...
Splicing regulation: a structural biology perspective
... interactions. Mass spectrometric analyses of affinity-purified spliceosomal complexes indicate that the total number of spliceosome-associated factors is approximately 170 [1]. Among all the proteins involved in splicing, one can first distinguish the proteins which are part of the spliceosome (the ...
... interactions. Mass spectrometric analyses of affinity-purified spliceosomal complexes indicate that the total number of spliceosome-associated factors is approximately 170 [1]. Among all the proteins involved in splicing, one can first distinguish the proteins which are part of the spliceosome (the ...
Alternative splicing

Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. Consequently the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes. Alternative splicing is sometimes termed differential splicing.Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome; in humans, ~95% of multi-exonic genes are alternatively spliced. There are numerous modes of alternative splicing observed, of which the most common is exon skipping. In this mode, a particular exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.The production of alternatively spliced mRNAs is regulated by a system of trans-acting proteins that bind to cis-acting sites on the primary transcript itself. Such proteins include splicing activators that promote the usage of a particular splice site, and splicing repressors that reduce the usage of a particular site. Mechanisms of alternative splicing are highly variable, and new examples are constantly being found, particularly through the use of high-throughput techniques. Researchers hope to fully elucidate the regulatory systems involved in splicing, so that alternative splicing products from a given gene under particular conditions could be predicted by a ""splicing code"".Abnormal variations in splicing are also implicated in disease; a large proportion of human genetic disorders result from splicing variants. Abnormal splicing variants are also thought to contribute to the development of cancer.