Inositol 1,3,4,5,6-Pentakisphosphate 2-Kinase
... seeds. A conserved W-box cis-element, TTGACC, is located at position 20 to 25 bp of this sequence. It is well documented that a family of plant-specific zinc finger transcription factors, the WRKY transcription factors, is able to bind to this cis-element and regulate gene expression (Ülker and Som ...
... seeds. A conserved W-box cis-element, TTGACC, is located at position 20 to 25 bp of this sequence. It is well documented that a family of plant-specific zinc finger transcription factors, the WRKY transcription factors, is able to bind to this cis-element and regulate gene expression (Ülker and Som ...
A mixed group ll/group III twintron in the Euglena
... of introns and the role of introns during gene evolution. The debate over this question has focused on two views of intron evolution: 'introns early' versus 'introns late'. In the introns early model, genes are viewed as being assembled from exons that code for structural or functional domains. Intr ...
... of introns and the role of introns during gene evolution. The debate over this question has focused on two views of intron evolution: 'introns early' versus 'introns late'. In the introns early model, genes are viewed as being assembled from exons that code for structural or functional domains. Intr ...
Analysis of Two Genes Encoding Prothrombin Activators in
... a step forward for the study. It has given us another set of direction to which aspects of the gene structure is worth investigating in order to elucidate the different mechanisms involved in the regulating of transcription activities of the two genes. However no direct conclusion can be made for th ...
... a step forward for the study. It has given us another set of direction to which aspects of the gene structure is worth investigating in order to elucidate the different mechanisms involved in the regulating of transcription activities of the two genes. However no direct conclusion can be made for th ...
New lysosomal acid lipase gene mutants explain the phenotype of
... tion at the 21 position of exon 8 donor site (the 59 splice site of the corresponding intron) that results in exon skipping and the subsequent loss of 24 amino acids from the protein is the most frequent defect observed in patients with CESD (4, 7–9). Skipping of the same exon has also been observed ...
... tion at the 21 position of exon 8 donor site (the 59 splice site of the corresponding intron) that results in exon skipping and the subsequent loss of 24 amino acids from the protein is the most frequent defect observed in patients with CESD (4, 7–9). Skipping of the same exon has also been observed ...
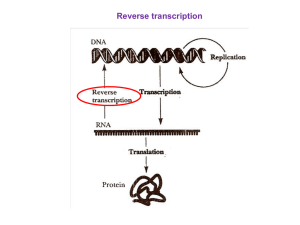
Insert Overview of Translation here 2 pages.
... In prokaryotes, this is fairly well understood. Prokaryotic mRNAs contain a ribosome binding site that is located 5' to (in front of) the start codon. This sequence is 5' AGGAGG 3'. It is called a Shine-Dalgarno sequence and it is found about 10 bases 5' to the start codon. The 16S rRNA, in turn, co ...
... In prokaryotes, this is fairly well understood. Prokaryotic mRNAs contain a ribosome binding site that is located 5' to (in front of) the start codon. This sequence is 5' AGGAGG 3'. It is called a Shine-Dalgarno sequence and it is found about 10 bases 5' to the start codon. The 16S rRNA, in turn, co ...
Chapter 15
... Eukaryotic pre-mRNA splicing • Introns – non-coding sequences • Exons – sequences that will be translated • Small ribonucleoprotein particles (snRNPs) recognize the intron–exon boundaries • snRNPs cluster with other proteins to form spliceosome – Responsible for removing introns ...
... Eukaryotic pre-mRNA splicing • Introns – non-coding sequences • Exons – sequences that will be translated • Small ribonucleoprotein particles (snRNPs) recognize the intron–exon boundaries • snRNPs cluster with other proteins to form spliceosome – Responsible for removing introns ...
RNA-Quant™ cDNA Synthesis Kit
... RNA-Quant™ cDNA Synthesis Kit Efficiently make cDNA to measure any RNA by qPCR Total RNA was harvested from human HT1080 cells using standard Trizol extraction protocols. The RNA-Quant kit was used to tail and tag all RNAs into quantifiable cDNA for qPCR analysis. Sample amplification plots and spe ...
... RNA-Quant™ cDNA Synthesis Kit Efficiently make cDNA to measure any RNA by qPCR Total RNA was harvested from human HT1080 cells using standard Trizol extraction protocols. The RNA-Quant kit was used to tail and tag all RNAs into quantifiable cDNA for qPCR analysis. Sample amplification plots and spe ...
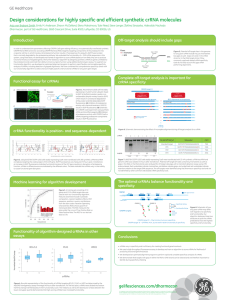
Design considerations for highly specific and efficient
... Off-target analysis should include gaps ...
... Off-target analysis should include gaps ...
Biomart/ GENOME ALIGNMENT III
... and the mouse sequence. Even in remote organisms such as fugu and zebrafish some of the human exons are conserved. Between rat and mouse and human in the region 1000 bp upstream of the first exons parts of the sequence are conserved as well. These might correspond to the regulatory motifs, responsib ...
... and the mouse sequence. Even in remote organisms such as fugu and zebrafish some of the human exons are conserved. Between rat and mouse and human in the region 1000 bp upstream of the first exons parts of the sequence are conserved as well. These might correspond to the regulatory motifs, responsib ...
Glimpses of a few literatures on snRNA
... Number of sequenced examples is a snapshot as of 2002 and is influenced by DNA-sequencing strategies and database upkeep; it may provide a rough indication of relative abundance. RNAs in any group vary in size; the size provided here indicates the lower end of the length distribution for the natura ...
... Number of sequenced examples is a snapshot as of 2002 and is influenced by DNA-sequencing strategies and database upkeep; it may provide a rough indication of relative abundance. RNAs in any group vary in size; the size provided here indicates the lower end of the length distribution for the natura ...
Information Encoding in Biological Molecules: DNA and
... • exons can be added to an existing transcript by shift + left-click + drag exon to the transcript • to create an exon without support right-click and select “Create exon” • to delete exon/transcript choose “Delete feature” Lab 5.2 ...
... • exons can be added to an existing transcript by shift + left-click + drag exon to the transcript • to create an exon without support right-click and select “Create exon” • to delete exon/transcript choose “Delete feature” Lab 5.2 ...
RNA Polymerase - California Lutheran University
... Alternative splicing • Single primary transcript can be spliced into different mRNAs by the inclusion of different sets of exons • 15% of known human genetic disorders are due to altered splicing • 35 to 59% of human genes exhibit some form of alternative splicing • Explains how 25,000 genes of the ...
... Alternative splicing • Single primary transcript can be spliced into different mRNAs by the inclusion of different sets of exons • 15% of known human genetic disorders are due to altered splicing • 35 to 59% of human genes exhibit some form of alternative splicing • Explains how 25,000 genes of the ...
Complete Characterization of the 3 Mouse Hereditary Hemochromatosis HFE Gene and
... right, a photograph of the RT-PCR experiment is shown indicating the correspondence between the splicing forms detected after sequencing (left) and the PCR amplification products (lane 1 from the photograph). M, 1-kb size marker. The primers used for the RT (oligo dT) and PCR (Ex1U and Ex6L) are ind ...
... right, a photograph of the RT-PCR experiment is shown indicating the correspondence between the splicing forms detected after sequencing (left) and the PCR amplification products (lane 1 from the photograph). M, 1-kb size marker. The primers used for the RT (oligo dT) and PCR (Ex1U and Ex6L) are ind ...
Brooker Genetics 5e Sample Chapter 16
... altering phenotype. Even so, many recent studies have suggested that environmentally induced changes in an organism’s characteristics are rooted in epigenetic changes that alter gene expression. For example, several studies have indicated that temperature changes have epigenetic effects. In certain ...
... altering phenotype. Even so, many recent studies have suggested that environmentally induced changes in an organism’s characteristics are rooted in epigenetic changes that alter gene expression. For example, several studies have indicated that temperature changes have epigenetic effects. In certain ...
6.
... nine sequences (median 3), the average EST and mRNA support for non-CAS exons was 2.2 sequences (median 1). However, sequence support by itself is not sufficient for detecting functional alternative splicing: in our study, 30% of the CAS exons were supported by a single human expressed sequence. We ...
... nine sequences (median 3), the average EST and mRNA support for non-CAS exons was 2.2 sequences (median 1). However, sequence support by itself is not sufficient for detecting functional alternative splicing: in our study, 30% of the CAS exons were supported by a single human expressed sequence. We ...
Inquiry into Life Twelfth Edition
... • Extra nucleotides are removed from the 5’ends of pre-tRNA in one step by an endonucleolytic cleavage catalyzed by RNase P • RNase P from bacteria and eukaryotic nuclei have a catalytic RNA subunit called M1 RNA • Spinach chloroplast RNase P appears to lack an RNA subunit ...
... • Extra nucleotides are removed from the 5’ends of pre-tRNA in one step by an endonucleolytic cleavage catalyzed by RNase P • RNase P from bacteria and eukaryotic nuclei have a catalytic RNA subunit called M1 RNA • Spinach chloroplast RNase P appears to lack an RNA subunit ...
An accessible database for mouse and human whole transcriptome
... (2) Browse to your gene of interest. (3) To view the track ‘Whole transcriptome qPCR primers’ scroll down the page to the panel of suggested tracks named ‘Expression’. Turn the ‘qPCR primers’ to ‘pack’ (see Fig. 2A) and press refresh. (4) Use the UCSC browser tools to zoom in to specific regions of ...
... (2) Browse to your gene of interest. (3) To view the track ‘Whole transcriptome qPCR primers’ scroll down the page to the panel of suggested tracks named ‘Expression’. Turn the ‘qPCR primers’ to ‘pack’ (see Fig. 2A) and press refresh. (4) Use the UCSC browser tools to zoom in to specific regions of ...
The Wiskott-Aldrich Syndrome and X-Linked
... Exon 2: Intron 2: Exon 3: Intron 3: Exon 4: Intron 2: Exon 3: Intron 3: Exon 4: Intron 4 Intron 7: Exon 8: Intron 8: Exon 9: Intron 9: Exon 1 0 Exon 10: Intron 10: Exon 11: Intron 10: ...
... Exon 2: Intron 2: Exon 3: Intron 3: Exon 4: Intron 2: Exon 3: Intron 3: Exon 4: Intron 4 Intron 7: Exon 8: Intron 8: Exon 9: Intron 9: Exon 1 0 Exon 10: Intron 10: Exon 11: Intron 10: ...
PowerPoint 演示文稿
... Spliceosome contains snRNPs which are composed of snRNAs and proteins. snRNAs in snRNPs are U-rich, and called URNAs. ...
... Spliceosome contains snRNPs which are composed of snRNAs and proteins. snRNAs in snRNPs are U-rich, and called URNAs. ...
Glimpses of a few literatures on snRNA
... Number of sequenced examples is a snapshot as of 2002 and is influenced by DNA-sequencing strategies and database upkeep; it may provide a rough indication of relative abundance. RNAs in any group vary in size; the size provided here indicates the lower end of the length distribution for the natura ...
... Number of sequenced examples is a snapshot as of 2002 and is influenced by DNA-sequencing strategies and database upkeep; it may provide a rough indication of relative abundance. RNAs in any group vary in size; the size provided here indicates the lower end of the length distribution for the natura ...
NAME: AKALABU, MAUREEN CHIDINMA COURSE: BCH 301 MAT
... nucleus ribonucleoprotein particles, abbreviated as snRNPs. This class of splicing is a very common feature of messenger RNA (mRNA) processing in "higher" eukaryotes such as humans. It is not yet known if snRNP-mediated splicing is catalyzed by the RNA components. Note also that some RNA splicing re ...
... nucleus ribonucleoprotein particles, abbreviated as snRNPs. This class of splicing is a very common feature of messenger RNA (mRNA) processing in "higher" eukaryotes such as humans. It is not yet known if snRNP-mediated splicing is catalyzed by the RNA components. Note also that some RNA splicing re ...
PBI 6 Features on Teacher`s Map 2-08.qxp
... Nucleotides 62,187 to 62,278: Exon I (92 nucleotides, 30 2/3 codons) The Translation Start Site (AUG) is located at nucleotides 62,187- 62,189. All proteins begin with the amino acid methionine, Met, encoded by nucleotides AUG. This rule is a consequence of the mechanism that cells use to begin prot ...
... Nucleotides 62,187 to 62,278: Exon I (92 nucleotides, 30 2/3 codons) The Translation Start Site (AUG) is located at nucleotides 62,187- 62,189. All proteins begin with the amino acid methionine, Met, encoded by nucleotides AUG. This rule is a consequence of the mechanism that cells use to begin prot ...
Alternative splicing

Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. Consequently the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes. Alternative splicing is sometimes termed differential splicing.Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome; in humans, ~95% of multi-exonic genes are alternatively spliced. There are numerous modes of alternative splicing observed, of which the most common is exon skipping. In this mode, a particular exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.The production of alternatively spliced mRNAs is regulated by a system of trans-acting proteins that bind to cis-acting sites on the primary transcript itself. Such proteins include splicing activators that promote the usage of a particular splice site, and splicing repressors that reduce the usage of a particular site. Mechanisms of alternative splicing are highly variable, and new examples are constantly being found, particularly through the use of high-throughput techniques. Researchers hope to fully elucidate the regulatory systems involved in splicing, so that alternative splicing products from a given gene under particular conditions could be predicted by a ""splicing code"".Abnormal variations in splicing are also implicated in disease; a large proportion of human genetic disorders result from splicing variants. Abnormal splicing variants are also thought to contribute to the development of cancer.