Aligning reads with Galaxy
... • The basics – What is RNA-seq, alternative splicing. • Assembly techniques – Reference-based alignment – De-novo assembly ...
... • The basics – What is RNA-seq, alternative splicing. • Assembly techniques – Reference-based alignment – De-novo assembly ...
file
... Figure S2: Generation of Fzd3 mutant mice (related to Figure 7). (A) Schematic representation of the knockout first allele (modified from EUCOMM). A cassette containing FTR-Engrailed–2 exon–IRES–LacZ–loxP–neo–FRT–loxP was inserted in intron 2. In Fzd3ko/ko mice, three mRNAs isoforms are produced. I ...
... Figure S2: Generation of Fzd3 mutant mice (related to Figure 7). (A) Schematic representation of the knockout first allele (modified from EUCOMM). A cassette containing FTR-Engrailed–2 exon–IRES–LacZ–loxP–neo–FRT–loxP was inserted in intron 2. In Fzd3ko/ko mice, three mRNAs isoforms are produced. I ...
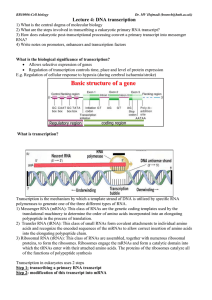
Lecture 4: DNA transcription
... Performed by spliceosomes (large RNA-protein complex made of small nuclear ribonucleoproteins) Recognise exon-intron boundaries and splice exons together by transesterification reactions Cell type-specific splicing ...
... Performed by spliceosomes (large RNA-protein complex made of small nuclear ribonucleoproteins) Recognise exon-intron boundaries and splice exons together by transesterification reactions Cell type-specific splicing ...
Ch. 10: Presentation Slides
... RNA Transcription: Splicing • Spliceosomes contain protein and specialized small RNAs complementary to the splice junctions to provide specificity to splicing reaction • Small nuclear RNAs U1, U2 and U5 recognize splice donor and acceptor sites by complementary base pairing so that intron excision ...
... RNA Transcription: Splicing • Spliceosomes contain protein and specialized small RNAs complementary to the splice junctions to provide specificity to splicing reaction • Small nuclear RNAs U1, U2 and U5 recognize splice donor and acceptor sites by complementary base pairing so that intron excision ...
Gene Section SRSF1 (serine/arginine rich splicing factor 1) -
... SF2/ASF modulates both constitutive and alternative pre-mRNA splicing. The function of SF2/ASF in splicing depends on the pre-mRNA sequence and the cellular context. Cis-acting sequences and trans-acting factors modulate SF2/ASF activity. For example, SF2/ASF antagonizes the activity of hnRNP A/B pr ...
... SF2/ASF modulates both constitutive and alternative pre-mRNA splicing. The function of SF2/ASF in splicing depends on the pre-mRNA sequence and the cellular context. Cis-acting sequences and trans-acting factors modulate SF2/ASF activity. For example, SF2/ASF antagonizes the activity of hnRNP A/B pr ...
Solid Tumour Section Lung: non-small cell carcinoma with inv(2)(p21p23)
... EML4-ALK fusion gene. Incidence of such tumors is 4-5% in non-small cell lung cancer among the Asian ethnic group, but may be lower among the others. ...
... EML4-ALK fusion gene. Incidence of such tumors is 4-5% in non-small cell lung cancer among the Asian ethnic group, but may be lower among the others. ...
pdf
... (3) These two phosphoester transfers result in a joining of the two exons and excision of the intron (with the initiating G nucleotide attached to the 5' end.) (4) The excised intron is then circularized by attack of the 3'-OH of the last nucleotide of the intron on the phosphate between the 15th an ...
... (3) These two phosphoester transfers result in a joining of the two exons and excision of the intron (with the initiating G nucleotide attached to the 5' end.) (4) The excised intron is then circularized by attack of the 3'-OH of the last nucleotide of the intron on the phosphate between the 15th an ...
Functional Analysis of A Novel Splicing Mutation in The Mutase
... The Novel mutation reported here leads to retention of the intron 12, which adds 10 amino acids to the encoding protein after exon 12 but also causes a premature stop codon leading to the complete deletion of the following exon 13. This mutation interrupts the vitamin B12 binding site. Thus, it rend ...
... The Novel mutation reported here leads to retention of the intron 12, which adds 10 amino acids to the encoding protein after exon 12 but also causes a premature stop codon leading to the complete deletion of the following exon 13. This mutation interrupts the vitamin B12 binding site. Thus, it rend ...
exon f exon g
... estimated from its sequence. The mean score of a random protein conformation is estimated by a weighted sum of protein composition over the 20 standard amino acid residue types, where each weight corresponds to the expected change in the score by inserting a specific type of amino acid residue. The ...
... estimated from its sequence. The mean score of a random protein conformation is estimated by a weighted sum of protein composition over the 20 standard amino acid residue types, where each weight corresponds to the expected change in the score by inserting a specific type of amino acid residue. The ...
The 1997 GSA Honors and Awards
... yeast snRNAs to events in mammalian cells remained, future events would reveal Christine’s pioneering work in this area to be foundational to the understanding of both premessenger RNA splicing and eukaryotic preribosomal RNA processing. In addition to leading the chase for cellular factors that mig ...
... yeast snRNAs to events in mammalian cells remained, future events would reveal Christine’s pioneering work in this area to be foundational to the understanding of both premessenger RNA splicing and eukaryotic preribosomal RNA processing. In addition to leading the chase for cellular factors that mig ...
AP Details for Protein Synthesis
... • Code for ALL life! – strongest support for a common origin for all life ...
... • Code for ALL life! – strongest support for a common origin for all life ...
Evolution of the fibrinogen γ′ chain: implications for the binding of
... is an intron at the expected location, but a mere 2-residue extension would occur in the absence of splicing. All other mammals studied (opossum, mouse, rat, dog, cat, bovids, horse and three primates) have extended c¢ sequences, the lengths of which range from eight to 20 residues, the longest bein ...
... is an intron at the expected location, but a mere 2-residue extension would occur in the absence of splicing. All other mammals studied (opossum, mouse, rat, dog, cat, bovids, horse and three primates) have extended c¢ sequences, the lengths of which range from eight to 20 residues, the longest bein ...
DNA level results in a phenotype of the patient
... are modulated exclusively by CUGBP1 or MBNL1, but all events regulated by both proteins exhibit antagonistic responses [42]. In addition, while MBNL1 nuclear levels increase during development, CUGBP1 nuclear levels decrease [22,42], supporting the hypothesis that the level and localization of these ...
... are modulated exclusively by CUGBP1 or MBNL1, but all events regulated by both proteins exhibit antagonistic responses [42]. In addition, while MBNL1 nuclear levels increase during development, CUGBP1 nuclear levels decrease [22,42], supporting the hypothesis that the level and localization of these ...
Alternative splicing induced by nonsense mutations in the
... decreases the score of an ESEfinder-predicted binding site for SC35 significantly from 2.51 (just above threshold) to 0.25, and RESCUE-ESE also predicts the elimination of an ESE by this mutation. In summary, ESEfinder and RESCUE-ESE differ in their predictions about the effect of the nonsense mutat ...
... decreases the score of an ESEfinder-predicted binding site for SC35 significantly from 2.51 (just above threshold) to 0.25, and RESCUE-ESE also predicts the elimination of an ESE by this mutation. In summary, ESEfinder and RESCUE-ESE differ in their predictions about the effect of the nonsense mutat ...
Modular proteins I
... The Kunitz type proteinase inhibitor is a single module protein: In bovine pancreatic trypsin inhibitor gene, phase 1 intons are found at both boundaries of the single inhibitor domain (protomodule stage) Lipoprotein-associated coagulation inhibitor consists of three tandem copies of this module (ta ...
... The Kunitz type proteinase inhibitor is a single module protein: In bovine pancreatic trypsin inhibitor gene, phase 1 intons are found at both boundaries of the single inhibitor domain (protomodule stage) Lipoprotein-associated coagulation inhibitor consists of three tandem copies of this module (ta ...
MS Word File
... anticodon on loop will bind to complementary codon on mRNA amino acid binding site at 3’ end appropriate amino acid is added by aminoacyl tRNA synthetase separate enzyme for each amino acid (20 total) Initiation-Ribosome binds to mRNA near AUG codon (ribosome binding site) Small ribosomal subunit, f ...
... anticodon on loop will bind to complementary codon on mRNA amino acid binding site at 3’ end appropriate amino acid is added by aminoacyl tRNA synthetase separate enzyme for each amino acid (20 total) Initiation-Ribosome binds to mRNA near AUG codon (ribosome binding site) Small ribosomal subunit, f ...
1. ELONGATION
... Consensus sequences of 5’ and 3’ splice junctions in eukaryotic mRNAs. Almost all introns begin with GU and end with AG. From the analysis of many exon intron boundaries, extended consensus sequences of preferred nucleotides at the 5’ and 3’ ends have been established. In addition to AG, other nucle ...
... Consensus sequences of 5’ and 3’ splice junctions in eukaryotic mRNAs. Almost all introns begin with GU and end with AG. From the analysis of many exon intron boundaries, extended consensus sequences of preferred nucleotides at the 5’ and 3’ ends have been established. In addition to AG, other nucle ...
Kein Folientitel
... Chimeric ESTs can be detected by genome comparison There is one particularly bad class of chimeric sequences that will be subject of the exercises. ...
... Chimeric ESTs can be detected by genome comparison There is one particularly bad class of chimeric sequences that will be subject of the exercises. ...
Figure 2 - GEP Community Server
... using a chemical method to tag the special structure that occurs at 5’ ends of transcript, fishing out the RNA molecules using these tags, and mapping the sequence back to the genome, a method called “CAGE” (cap analysis of gene expression). In addition, we will also display the "D. mel. cDNA" track ...
... using a chemical method to tag the special structure that occurs at 5’ ends of transcript, fishing out the RNA molecules using these tags, and mapping the sequence back to the genome, a method called “CAGE” (cap analysis of gene expression). In addition, we will also display the "D. mel. cDNA" track ...
module 3: transcription part ii
... using a chemical method to tag the special structure that occurs at 5’ ends of transcript, fishing out the RNA molecules using these tags, and mapping the sequence back to the genome, a method called “CAGE” (cap analysis of gene expression). In addition, we will also display the "D. mel. cDNA" track ...
... using a chemical method to tag the special structure that occurs at 5’ ends of transcript, fishing out the RNA molecules using these tags, and mapping the sequence back to the genome, a method called “CAGE” (cap analysis of gene expression). In addition, we will also display the "D. mel. cDNA" track ...
Example: search for regulatory binding sites
... Thermodynamic principle The amino acid sequence contains all the information necessary to fold a protein molecule into its native 3D state under physiological conditions: fold, denature, spontaneously refold, called Anfinsen’s thermodynamic principle Thus it should be possible to predict 3D structu ...
... Thermodynamic principle The amino acid sequence contains all the information necessary to fold a protein molecule into its native 3D state under physiological conditions: fold, denature, spontaneously refold, called Anfinsen’s thermodynamic principle Thus it should be possible to predict 3D structu ...
E. CELL SPECIALIZATION: RNA and Protein Regulation
... ___________________________________ ___________________________________ ___________________________________ ___________________________________ 2. Splicing different mRNAs from the same nRNA using different exons allows cells to choose the protein they will make – Alternative splicing occurs in ~92% ...
... ___________________________________ ___________________________________ ___________________________________ ___________________________________ 2. Splicing different mRNAs from the same nRNA using different exons allows cells to choose the protein they will make – Alternative splicing occurs in ~92% ...
Chapter 5
... Dscam complexity is essential to the establishment of the neural net by excluding self-synapses from forming ...
... Dscam complexity is essential to the establishment of the neural net by excluding self-synapses from forming ...
Masters change, slaves remain
... represses the transcription of genes required for male development and activates those required for female development. When Sxl protein is absent, tra mRNA is spliced in a nonproductive pattern, which includes exon sequences containing a stop codon. No Tra protein is expressed from the male tra mRN ...
... represses the transcription of genes required for male development and activates those required for female development. When Sxl protein is absent, tra mRNA is spliced in a nonproductive pattern, which includes exon sequences containing a stop codon. No Tra protein is expressed from the male tra mRN ...
Natural rules for Arabidopsis thaliana
... data by using different bioinformatical means and allows them to illustrate the potential mechanism of pre-mRNA splicing. Some researchers have applied large confirmed human gene samples in their studies to improve the results’ accuracy and sensitivity [14]. However, to date, there have been no repo ...
... data by using different bioinformatical means and allows them to illustrate the potential mechanism of pre-mRNA splicing. Some researchers have applied large confirmed human gene samples in their studies to improve the results’ accuracy and sensitivity [14]. However, to date, there have been no repo ...
Alternative splicing

Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. Consequently the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes. Alternative splicing is sometimes termed differential splicing.Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome; in humans, ~95% of multi-exonic genes are alternatively spliced. There are numerous modes of alternative splicing observed, of which the most common is exon skipping. In this mode, a particular exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.The production of alternatively spliced mRNAs is regulated by a system of trans-acting proteins that bind to cis-acting sites on the primary transcript itself. Such proteins include splicing activators that promote the usage of a particular splice site, and splicing repressors that reduce the usage of a particular site. Mechanisms of alternative splicing are highly variable, and new examples are constantly being found, particularly through the use of high-throughput techniques. Researchers hope to fully elucidate the regulatory systems involved in splicing, so that alternative splicing products from a given gene under particular conditions could be predicted by a ""splicing code"".Abnormal variations in splicing are also implicated in disease; a large proportion of human genetic disorders result from splicing variants. Abnormal splicing variants are also thought to contribute to the development of cancer.