* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download HIV treatment
Psychopharmacology wikipedia , lookup
Drug interaction wikipedia , lookup
CCR5 receptor antagonist wikipedia , lookup
Discovery and development of tubulin inhibitors wikipedia , lookup
Discovery and development of cyclooxygenase 2 inhibitors wikipedia , lookup
Neuropharmacology wikipedia , lookup
Discovery and development of dipeptidyl peptidase-4 inhibitors wikipedia , lookup
Discovery and development of direct thrombin inhibitors wikipedia , lookup
MTOR inhibitors wikipedia , lookup
Discovery and development of direct Xa inhibitors wikipedia , lookup
HIV vaccine wikipedia , lookup
Neuropsychopharmacology wikipedia , lookup
Discovery and development of ACE inhibitors wikipedia , lookup
Discovery and development of neuraminidase inhibitors wikipedia , lookup
Metalloprotease inhibitor wikipedia , lookup
Discovery and development of non-nucleoside reverse-transcriptase inhibitors wikipedia , lookup
Discovery and development of HIV-protease inhibitors wikipedia , lookup
Discovery and development of integrase inhibitors wikipedia , lookup
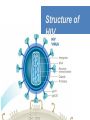
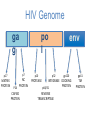
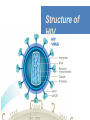
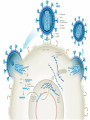
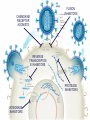
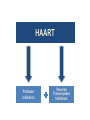
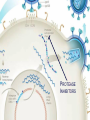
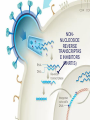
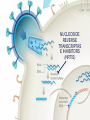
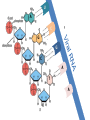
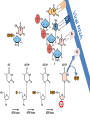
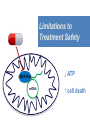
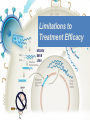
Nucleoside Analogs & HIV Treatments October 4th, 2016 PHM142 Fall 2016 Instructor: Dr. Jeffrey Henderson Meredith Ames Insun Cho Nathalie Dagenais Zhiyi Molly Yang Let’s take a look! Structure of HIV HIV Genome ga g po l p17 p7 MATRIX PROTEIN NC PROTEIN P24 CAPSID PROTEIN p10 PROTEASE env p32 gp120 INTEGRASE DOCKING PROTEIN p66/51 REVERSE TRANSCRIPTASE gp41 TM PROTEIN Structure of HIV CHEMOKINE RECEPTOR AGONISTS FUSION INHIBITORS REVERSE TRANSCRIPTAS E INHIBITORS PROTEASE INHIBITORS INTEGRASE INHIBITORS HAART Protease Inhibitors Reverse Transcriptase Inhibitors PROTEASE INHIBITORS NONNUCLEOSIDE REVERSE TRANSCRIPTAS E INHIBITORS (NNRTIS) NUCLEOSIDE REVERSE TRANSCRIPTAS E INHIBITORS (NRTIS) Quick Review: Reverse Transcription A Closer Look: Reverse Transcriptase U A C G G C A T A Zidovudine (AZT) T Lamivudine(3TC ) C T So Where do Nucleoside Analogs fit in? Abacavir Didanosine (DDI) GG & Emtricitabine, Stavudine, Tenofovir… A G C G G T C A T AA Limitations to Treatment Safety DNA Polγ mtDNA ↓ ATP ↑ cell death Limitations to Treatment Efficacy M184V K65R L74V Why Use NRTIs? Combination therapy ↓ morbidity ↓ mortality Adverse events & Resistance Summary: •HIV is a retrovirus with a genome that codes for 3 polyproteins, gp120 and gp41 are responsible for fusion into the host cell. •There exists 6 types of antiviral drugs used in the treatment of HIV, these six types of drugs include nucleoside reverse transcriptase inhibitors, non-nucleoside reverse transcriptase inhibitors, protease inhibitors, integrase inhibitors, chemokine receptor antagonists (CCR5) and lastly a relatively new class, HIV fusion inhibitors. •HAART (Highly active antiretroviral therapy) is a combination therapy often used in the treatment of HIV, which is made up of a 3 or 4 drugs, usually 2 nucleoside reverse transcriptase inhibitors with a non-nucleoside reverse transcriptase inhibitor, or alternatively one or two protease inhibitors. • Protease inhibitors: Protease inhibitors have a high affinity for the active site of HIV-1 protease and thus bind to the active site irreversibly. This binding to the active site impairs the catalytic activity of HIV-1 protease resulting in immature viral particles that don’t properly package into virions. • NNRIs specifically target HIV-1 reverse transcriptase and thus exhibit specificity. This binding has effects on the active site of HIV-1 reverse transcriptase, including conformational changes and changes in the “primer grip” in p66 (one of two subunits of the HIV-1 heterodimer) that is responsible for positioning the DNA primer within the active site. •Reverse Transcriptase: Pre-packaged Viral enzyme, heterodimer: subunits p66 & p51 of pol polyprotein; poor proof-reading capability (double edged sword) •2 Distinct Enzymatic Activities: DNA Polymerase (RNA & DNA templates) & Rnase H activity •Nucleoside Analogs: Synthetic compounds developed to mimic their physiological counterparts; inhibit reverse transcription through their incorporation into growing DNA strand (AZT: thymidine analog, 3TC: Cytidine analog) •Mechanism of action: – Active Transport into cell (some NAs may enter passively) – 3-Step Metabolic Activation: phosphorylated by host cells kinases to active form, a 5’-trisphosphate derivative – Inhibit Reverse Transcription through dual faceted approach: • Competitive Inhibition • Chain Termination (No 3’ Hydroxyl group: no phosphodiester linkage with incoming dNTP alpha-phosphate) •Pros of NRTIs: effective treatment method when used in combination with other HIV treatments in HAART to decrease resistance mutations •Cons of NRTIs: 1. Limitations in treatment safety – NRTIs inhibit DNA polymerase gamma which results in functional loss and subsequent decrease in ATP production via oxidative phosphorylation and mitochondria-regulated apoptosis; Net effect is increase cellular dysfunction and cell death 2. Limitations in treatment efficacy – NRTIs lose efficacy due to formation of mutant HIV variants, which confer resistance by two mechanisms i) discrimination between phosphorylated nucleoside analogues vs natural dNTP substrates results in decreased incorporation of drugs ii) increased reverse transcriptase phosphorlytic activity to excise drug from primer Works Cited 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. Jordheim, L. P., Durantel, D., Zoulim, F. & Dumontet, C. Advances in the development of nucleoside and nucleotide analogues for cancer and viral diseases. Nat. Rev. Drug Discov. 12, 447–464 (2013). Brinkman, K., ter Hofstede, H. J., Burger, D. M., Smeitink, J. A. & Koopmans, P. P. Adverse effects of reverse transcriptase inhibitors: mitochondrial toxicity as common pathway. AIDS Lond. Engl. 12, 1735–1744 (1998). Kohlstaedt, L. A., Wang, J., Friedman, J. M., Rice, P. A. & Steitz, T. A. Crystal structure at 3.5 A resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science 256, 1783–1790 (1992). Sarafianos, S. G., Hughes, S. H. & Arnold, E. Designing anti-AIDS drugs targeting the major mechanism of HIV-1 RT resistance to nucleoside analog drugs. Int. J. Biochem. Cell Biol. 36, 1706–1715 (2004). Brik, A. & Wong, C.-H. HIV-1 protease: mechanism and drug discovery. Org. Biomol. Chem. 1, 5–14 (2003). Fisher, R. A. et al. HIV infection is blocked in vitro by recombinant soluble CD4. Nature 331, 76–78 (1988). Robinson, W. E. et al. Human Monoclonal Antibodies to the Human Immunodeficiency Virus Type 1 (HIV-1) Transmembrane Glycoprotein gp41 Enhance HIV-1 Infection in vitro. Proc. Natl. Acad. Sci. U. S. A. 87, 3185–3189 (1990). Spence, R. A., Kati, W. M., Anderson, K. S. & Johnson, K. A. Mechanism of inhibition of HIV-1 reverse transcriptase by nonnucleoside inhibitors. Science 267, 988 (1995). Arts, E. J. & Wainberg, M. A. Mechanisms of nucleoside analog antiviral activity and resistance during human immunodeficiency virus reverse transcription. Antimicrob. Agents Chemother. 40, 527–540 (1996). Menéndez-Arias, L. Mechanisms of resistance to nucleoside analogue inhibitors of HIV-1 reverse transcriptase. Virus Res. 134, 124–146 (2008). Arnold, E. & Sarafianos, S. G. Molecular biology: An HIV secret uncovered. Nature 453, 169–170 (2008). Cherry, C. L. & Wesselingh, S. L. Nucleoside analogues and HIV: the combined cost to mitochondria. J. Antimicrob. Chemother. 51, 1091–1093 (2003). Kakuda, T. N. Pharmacology of nucleoside and nucleotide reverse transcriptase inhibitor-induced mitochondrial toxicity. Clin. Ther. 22, 685–708 (2000). Zhang, S. et al. Protease inhibitors used in the treatment of HIV+ induce β-cell apoptosis via the mitochondrial pathway and compromise insulin secretion. Am. J. Physiol. - Endocrinol. Metab. 296, E925–E935 (2009). Wang, J. et al. Structural basis of asymmetry in the human immunodeficiency virus type 1 reverse transcriptase heterodimer. Proc. Natl. Acad. Sci. 91, 7242–7246 (1994). Salie, Z. L. et al. Structural basis of HIV inhibition by translocation-defective RT inhibitor 4′-ethynyl-2-fluoro-2′deoxyadenosine (EFdA). Proc. Natl. Acad. Sci. 113, 9274–9279 (2016). Meyer, P. R., Matsuura, S. E., So, A. G. & Scott, W. A. Unblocking of chain-terminated primer by HIV-1 reverse transcriptase through a nucleotide- dependent mechanism. Proc. Natl. Acad. Sci. 95, 13471–13476 (1998).