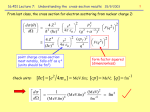
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download MIDAS2_19
Genome evolution wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene regulatory network wikipedia , lookup
Genomic imprinting wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Ridge (biology) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
MIcroarray Data Analysis System (version 2.19) Wei Liang October 2004 Microarray Data Flow Printer Scanner .tiff Image File Image Analysis Raw Gene Expression Data Gene Annotation AGED Others… Normalization / Filtering Normalized Data with Gene Annotation MAD Database Database Database Data Entry / Management Expression Analysis Interpretation of Analysis Results MIDAS is a Normalization and Filtering tool for microarray data analysis! MIDAS is a Normalization and Filtering tool for microarray data analysis! Serves as a data pre-processor for clustering analysis (MeV). Why Normalization and Filtering? .tiff Image Files Sample1 mRNA Raw Data File Cy3 intensity Cy3 RT Cy3-cDNA Cy5 RT Sample2 mRNA Systematic experimental error cDNA array Cy5-cDNA Uneven hybridization gel print-tip variations Background variations Cy5 intensity Wavelength dependent Intensity dependent Image processing algorithmdependent Why Normalization and Filtering? • The hypothesis underlying microarray analysis is that the measured intensities for each arrayed gene represent its relative expression level. • We use these intensities to identify biologically relevant patterns of expression by comparing measured levels between states on a gene-by-gene basis. • However, before the levels can be appropriately compared, one generally performs a number of transformations on the data to eliminate questionable or low quality data, to adjust the measured intensities to facilitate comparisons, and to select those genes that are significantly differentially expressed. MIDAS data analysis methods • 8 normalization/transformation methods • Total Intensity normalization Ratio Statistics normalization LOWESS (Locfit) normalization Standard deviation regularization Iterative linear regression normalization In-slide replicates analysis Iterative log mean centering normalization MA-ANOVA 10 quality control filtering methods Flip-dye consistency checking Low intensity filter Ratio Statistics confidence interval checking Spot QC flag checking Invalid-intensity checking Signal/Noise checking Cross-file-trim • 3 significant genes identification methods Slice analysis (non-statistical) Cross-slide replicates t-test (statistical) Cross-slide one-class SAM (statistical) Graphical scripting language Graphical scripting language • Read input files • Define analysis pipeline and set parameters for each analysis module • Write output files MIDAS data analysis methods • 8 normalization/transformation methods • Total Intensity normalization Ratio Statistics normalization LOWESS (Locfit) normalization Standard deviation regularization Iterative linear regression normalization In-slide replicates analysis Iterative log mean centering normalization MA-ANOVA 10 quality control filtering methods Flip-dye consistency checking Low intensity filter Ratio Statistics confidence interval checking Spot QC flag checking Invalid-intensity checking Signal/Noise checking Cross-file-trim • 3 significant genes identification methods Slice analysis (non-statistical) Cross-slide replicates t-test (statistical) Cross-slide one-class SAM (statistical) Sample data Pair # 1st file name 2nd file name 1 NFE005d0001.mev NFE005d00020.mev 2 NFE005d0002.mev NFE005d00021.mev 3 NFE005d0003.mev NFE005d00022.mev 4 NFE005d0004.mev NFE005d00023.mev 5 NFE005d0005.mev NFE005d00024.mev 6 NFE005d0006.mev NFE005d00025.mev 7 NFE005d0007.mev NFE005d00026.mev 9 NFE005d0008.mev NFE005d00027.mev 10 NFE005d0009.mev NFE005d00028.mev 11 NFE005d00010.mev NFE005d00029.mev 12 NFE005d00011.mev NFE005d00030.mev 13 NFE005d00012.mev NFE005d00031.mev 14 NFE005d00013.mev NFE005d00032.mev 15 NFE005d00014.mev NFE005d00033.mev 16 NFE005d00015.mev NFE005d00034.mev 17 NFE005d00016.mev NFE005d00035.mev 18 NFE005d00017.mev NFE005d00036.mev 19 NFE005d00018.mev NFE005d00037.mev 20 NFE005d00019.mev NFE005d00038.mev LOWESS (Locfit) normalization R-I plot: logRatio vs. logIntensityProduct A SD = 0.346 • Observations 1. Tilted tails at low intensity end and high intensity end 2. Mean not centered at 0 – intensity dependent LOWESS (Locfit) normalization Gene X A SD = 0.346 Exp factor Bio factor • If Cy3, Cy5 equally expressed, log2(Cy5/Cy3) = 0 • Two factors contributed to the up-regulated gene X: 1. Biological factors (we are interested) 2. Experimental factors, e.g. different sensitivity to red and green lasers (we are NOT interested and desire to get rid of.) LOWESS (Locfit) normalization Gene X A SD = 0.346 Exp factor Bio factor We need to find a way to extract the experimental factors Approach: Assume similar experimental factors applied to genes closer to each other in the logProd-logRatio plot Predict the Exp factor from a group of locally neighboring data --- equivalent to a curve fitting problem. LOWESS (Locfit) normalization • Local linear regression model • Tri-cube weight function • Least Squares A yi xi w( xi ) ( yi xi ) 2 w( x ) ( y x ) i ( X 'WX ) 1 X 'WY i i 2 0 Estimated values of log2(Cy5/Cy3) as function of log10(Cy3*Cy5) SD = 0.346 LOWESS (Locfit) normalization Use the estimated curve y(xi) to correct raw data Gene X A SD = 0.346 y(xi) = Exp factor Bio factor log2(Ri’/Gi’) = log2(Ri/Gi) – y(xi) log2(Ri’/Gi’) = log2(Ri/Gi) – log2 2y(xi) log2(Ri’/Gi’) = log2(Ri/Gi * 1/2y(xi)) Ri’ = Ri Gi’ = Gi * 2 y(xi) LOWESS (Locfit) normalization LOWESS-corrected RI plot B SD = 0.346 SD = 0.338 Standard deviation regularization Assumption: Within each block and each slide, spots should have the same spread for log(Cy5/Cy3, 2) values SD-Reg scales the (Cy3, Cy5) intensity pair for each spot so that the spot sets within each block or each slide will have the same standard deviation as other blocks or slides. Standard deviation regularization • Let aij be the raw log ratio for the jth spot in ith block (or slide) a’ij be the scaled log ratio for the jth spot in ith block (or slide) 2 Cy5 aij log 2 Cy3 (aij aij ) N j 1 a'ij aij 2 M M (aij aij ) N j 1 where Nj denotes the number of genes ith block or ith slide, M denotes the number of blocks or slides, aij denotes the log ratio mean of ith block (or ith slide) Standard deviation regularization Flip dye replicates consistency filter • Flip dye experiments help reduce random error • The intensities in the file pair are flipped, i.e. R1/G1 ~ G2/R2 or R1~ G2, G1 ~ R2 G1 R1 Gene1 Gene2 Gene3 Gene4 Gene5 Gene6 Gene7 Gene8 G2 R2 Flip dye replicates consistency filter • Calculate expression levels for all genes in the flip-dye pair • Filter genes with inconsistent expression levels between flip-dye replicates • For those genes passed the consistency checking, take geometric mean for the corresponding intensities from the replicated pairs How consistency is measured between replicates? Flip dye replicates consistency filter File 1 G1 R1 File 2 G2 R2 Gene 100% consistency: R1 G 2 G1 R 2 R1 G1 1 G2 R2 R1 R1R 2 log 2 G1 log 2 0 G2 G1G 2 R2 Flip dye replicates consistency Filter • SD cut vs. Threshold cut SD cut Threshold cut Regardless of datasets, always cut the same percentage for the same The percentage to cut depends on the specified log-ratio consistency range -1< log 2 1/2 < R1R 2 <1 G1G 2 R1R 2 <2 G1G 2 Flip dye replicates consistency filter • Calculate expression levels for all genes in the flip-dye pair • Filter genes with inconsistent expression levels between flip-dye replicates • For those genes passed the consistency checking, take geometric mean for the corresponding intensities from the replicated pairs Slice Analysis filter • Remove genes with z-scores beyond an interested range Slice Analysis filter • Remove genes with z-scores beyond an interested range Slice Analysis filter B SD = 0.346 SD = 0.338 • Define a slice window • Sliding the window along the log(IntensityProduct) axis • Calculate logRatioMean and logRatioSD of data points within each slice window • Calculate Z-scores of each data point Z-score = (logRatio-logRatioMean)/ logRatioSD • Trim data with Z-scores beyond interested range Slice Analysis filter 4 3 2 log2(Cy5/Cy3) 1 0 -1 -2 -3 -4 7 8 9 10 11 12 13 14 12 13 14 log(Cy3*Cy5) 8 6 4 log2(Cy5/Cy3) 2 0 -2 -4 -6 -8 7 8 9 10 11 log(Cy3*Cy5) Analysis packaging myAnalysis.prj MIDAS graphing MIDAS graphing R-I plot (.prc) Z-score Distribution plot (.his) Intensity plot (.ity, .lty) Box plot (.box) FlipDye Diagnostic plot (.rrc) SAM plot (.sam) MIDAS data viewer Statistical significant genes identification methods Two methods implemented in this release of MIDAS: • Cross-slide replicates one-class T-test • Cross-slide replicates one-class SAM SAM (Significance Analysis of Microarrays) A statistical technique for finding significant genes in a set of microarray experiments. Reference: Tusher, V.G., R. Tibshirani and G. Chu. 2001. Significance analysis of microarrays applied to the ionizing radiation response. Proceedings of the National Academy of Sciences USA 98: 51165121. Designs: • two-class unpaired • two-class paired • multi-class unpaired • censored survival • one-class (available in this release) SAM (Significance Analysis of Microarrays) One-class SAM: Identify genes whose mean expression across experiments are different from a user-specified mean. • Assign a score (d) to each gene based on its change in expression relative to the standard deviation of repeated measurements for the gene • Genes with scores > a threshold (Δ) are deemed potentially significant • For these “deemed potentially significant” genes, the proportion of them likely to have been wrongly identified by chance, or False Discovery Rate (FDR) is estimated • The goal is picking a set of differentially expressed genes with a user-satisfied FDR SAM (Significance Analysis of Microarrays) positively significant genes FDR Δ adjustment Automated report generation Automated report generation TM4 MIDAS web page http://www.tigr.org/software/tm4/midas.html http://www.tm4.org/midas.html