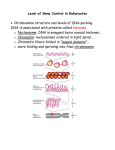
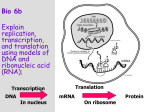
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download myoD
Survey
Document related concepts
Transcript
Section N—Regulation of transcription in eukaryotes N1 Eukaryotic transcription factors N2 Examples of transcriptional regulation N1 Eukaryotic transcription factors Transcription factor domain structure: Transcription factors other than the general transcription factors of the basal transcription complex were firt identified through their affinity for specific motifs in promoters, upstream regulatory elements of enhancer regions. These factors have two distinct activities. Firstly, they bind specifically to their DNA-binding site and, secondly, they activate transcription. These activities can be assigned to separate protein domains called activation domains and DNAbinding domains. In addition, many transcription factor occur as homo- or heterodimers, held togather by dimerization domains. A few transcription factors have ligand-binding of an accessory small molecule. Transcription factor domain structure Prokaryotes • Helix-turn-helix Eukaryotes • Zinc finger • Leucine zipper • Helix-loop-helix DNA-binding domains: The helix-turn-helix domain Characteristic: This domain contains a 60amino acid homeodomain which is encoded by a sequence called the homebox This domain consists of four alpha-helices in which helices II and III are at right angles to each other and are sparated by a beta-turn. this domain binds so that one helix, know as the recognition helix, lies partly in the major groove and interacts with the DNA. DNA-binding domains: the zinc finger domain Many, many; examples: estrogen receptor; TFIIIA Zn++ coordinated by: 2 Cys + 2 His = C2H2-type X Cys = CysX; X = 4 or 6 N-terminal end Often motif repeated 2 to 13 times Bind to DNA major grove Dimerization domains: leucine zippers Zipper: every 7th residue is a Leu Hydrophobic interface Leucine Zipper proteins contain a hydrophobic leucine residue at every sventh position in a region that is often at the C-terminal part of the DNAbinding domain. These leucines lie in an alpha-helical region and the regular repeat of these residues forms a hydrophobic surface on one side of the alpha-helix with a leucine every second turn of the helix. Dimerization domains: leucine zippers The N-terminal basic domains of each helix form a symmetrical structure in which each basic domain lies along the DNA in opposite directions, interacting with a symmetrical DNA recognition site so that the protein in effect forms a clamp around the DNA. Transcription activation domains • acidic activation domains: Comparison of the transactivation domains of yeast Gcn4 and Gal4, mammalian glucocorticoid receptor and herpes virus activator VP16 shows that they have a very high proportion of acidic amino acids. These have been called acidic activation domains or ‘acid blobs’ or ‘negative noodles’ and are characteristic of many transcription activation domains. It is still uncertain what other features are required for these regions to function as efficient transcription activation domains. • Glutamine-rich domains: were first identified in two activatio regions of the transcription factor SP1. As with acidic domains, the proportion of glutamine residues seems to be more important than overall structure. Domain swap experiments between glutamine-rich transcactivation regions from the diverse transcription factors SP1 and the Drosophila protein Antennapedia showed that these domains could substitue for each other. • Proline-rich domains: have been identified in several transcription factors. As with glutamine, a continuous run of proline residues can activate transcription. This domain is found, for example, in the c-Jun, AP2 and Oct-2 transcription factors. Repressor domains Repression of transcription may occur by indirect interference with the function of an activator. This may occur by: • Blocking the activator DNA-binding site. • Formation of a non-DNA-binding complex • Masking of the activation domain without preventing DNA binding Targets for transcriptional regulation The presence of diverse activation domains raises the question of whether the each have the same target in the basal transcription complex or different target for the activation of transcription. They are distinguishable from each other since the acidic activation domain can activate transcription from a downstream enhancer site while the proline domain only activates weakly and the glutamine domain not at all. Proposed targets of different transcriptional activators include: • Chromatin structure; • Interaction with TFIID through specific TAFIIS • Interaction with TFIIB • Interaction or modulation of the TFIIH complex activity leading to differential phosphorylation of the CTD of RNA Pol II N2 Examples of transcriptional regulation Constitutive transcription factors:SP1 • SP1 binds to a GC-rich sequence with the consensus sequence GGGCGC • It is a constitutive transcription factor whose binding site is found in the promoter of many housekeeping genes. •It contains the zinc finger motifs and has been shown to contain two glutamine-rich transactivation domains. • SP1 have been shown to interact specifically with TAFII 110, onw of thw TAFIIs which bind to the TATA binding protein to make up TFIID. Hormonal regulation: steroid hormone receptors Many transcription factors are activated by hormones which are secreted by one cell type and transmit a signal to a different cell type. One class of hormones, the steroid hormones, are lipid soluble and can diffuse through cell membranes to interact with transcripton factors called steroid hormine receptors. steroid Inibitor (HSP90) Glucocorticoid receptor In the absence of the steroid hormone, the receptors is bound to an inhibit, and located in the cytoplasm. The steroid hormone bind to the receptor and releases the receptor from the inhibitor, allowing the receptor to dimerize and translocate to the nucleus. The DNA-binding domain of the steroid hormone receptor then interacts with its specific DNA-binding sequence or response element, and this gives reise to activation of the target gene. Dissociation and dimerization Nuclear translocation Glucocorticoid response element Interferon-γ Regulation phosphorylation STAT proteins Many hormones do not diffuse into the cell. Instead, they bind to cell-surface receptors and pass a sgnal to proteins within the cell through a process called signal transduction. This process often involves protein phosphorylation. Interferon-γ induces phosphorylation of a transcription factor called STAT1α through JAK. When STAT1αbecmes phosphorylated at a specific tyrosine residue, it is able to form a homodimer which moves from the cytoplasm into the nucleus. INF- γ receptor Unphosphorylated STAT 1αmonoers Phosphorylation Dmerization Phosphorylated STAT1α dimer Nuclear translocation Response element Transcription elongation:HIV Tat Activated TFIIH phosphorylates CTD of RNA Pol II Initiation complex Cellular factors Tat Polymerase Precious transcript loops backwards to interact with the initiation compex Tat-TAR cellular factor complex activates TFIIH TAR stem-loop structure Human immunodeficiency virus (HIV) encodes an activatir protein called Tat, which is required for productive HIV gene expression. Tat binds to an RNA stem-loop structuire called TAR, which is present in the 5’-untranslated region of all HIV RNAs, just after the HIV transcription start site. The predominant effect of Tat in mammalian cells lies at the level of transcription elongation. Cell determination: myoD Nucleus myoD Other muscle-specific genes DNA OFF Embryonic precursor cell OFF 1 Determination. Signals from OFF myoD is a “master control” gene: it makes a mRNA transcription factor that can activate other The cell is now protein ireversibly muscle specific genes. MyoD (transcription factor) other cells activate a master regulatory gene, myoD, Myoblast (determined) 2 determined Differentiation. MyoD protein activates other muscle-specific transcription factors, which in turn activate genes for muscle proteins. The embryonic precursor cell is still undifferentiated mRNA MyoD Muscle cell (fully differentiated) The cell is now fully differentiated mRNA Another transcription factor mRNA mRNA Myosin, other muscle proteins, and cell-cycle blocking proteins Determination and differentiation of muscle cells Nucleus Master control gene myoD Other muscle-specific genes DNA OFF Embryonic precursor cell 1 Myoblast (determined) 成肌细胞2 Determination. Signals from other cells activate a master regulatory gene, myoD, Differentiation. MyoD protein activates other muscle-specific transcription factors, which in turn activate genes for muscle proteins. OFF OFF mRNA MyoD protein (transcription factor) mRNA MyoD Muscle cell (fully differentiated) Fig 21.10 The cell is now fully differentiated mRNA Another transcription factor The cell is now ireversibly determined to become a muscle cell. mRNA mRNA Myosin, other muscle proteins, and cell-cycle blocking proteins Determination and differentiation of muscle cells Nucleus Master control gene myoD Other muscle-specific genes DNA OFF Embryonic precursor cell OFF 1 Determination. Signals from other cells activate a master regulatory gene, myoD, OFF mRNA MyoD protein (transcription factor) Myoblast (determined) The cell is now ireversibly determined Differentiation. MyoD 2 protein activates other muscle-specific transcription factors, which in turn activate genes for muscle proteins. Muscle cell (fully differentiated) mRNA MyoD The cell is now fully differentiated mRNA Another transcription factor mRNA mRNA Myosin, other muscle proteins, and cell-cycle blocking proteins Embryonic development: homeodomain proteins The homeobox is a conserved DNA sequence which encodes the helix-turn-helix DNA binding protein structure called the homeodomain. The homeodomain was first discovered in the transcription factors encoded by homeptic genes of Drosophila. Homeotic genes Regulatory genes that control organ identity Conserved from flies to mammals